Abstract
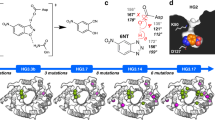
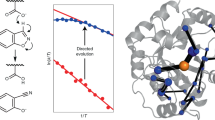
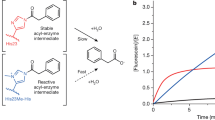
Linus Pauling established the conceptual framework for understanding and mimicking enzymes more than six decades ago1. The notion that enzymes selectively stabilize the rate-limiting transition state of the catalysed reaction relative to the bound ground state reduces the problem of design to one of molecular recognition. Nevertheless, past attempts to capitalize on this idea, for example by using transition state analogues to elicit antibodies with catalytic activities2, have generally failed to deliver true enzymatic rates. The advent of computational design approaches, combined with directed evolution, has provided an opportunity to revisit this problem. Starting from a computationally designed catalyst for the Kemp elimination3—a well-studied model system for proton transfer from carbon—we show that an artificial enzyme can be evolved that accelerates an elementary chemical reaction 6 × 108-fold, approaching the exceptional efficiency of highly optimized natural enzymes such as triosephosphate isomerase. A 1.09 Å resolution crystal structure of the evolved enzyme indicates that familiar catalytic strategies such as shape complementarity and precisely placed catalytic groups can be successfully harnessed to afford such high rate accelerations, making us optimistic about the prospects of designing more sophisticated catalysts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pauling, L. Molecular architecture and biological reactions. Chem. Eng. News 24, 1375–1377 (1946)
Hilvert, D. Critical analysis of antibody catalysis. Annu. Rev. Biochem. 69, 751–793 (2000)
Privett, H. K. et al. Iterative approach to computational enzyme design. Proc. Natl Acad. Sci. USA 109, 3790–3795 (2012)
Albery, W. J. & Knowles, J. R. Free-energy profile for the reaction catalyzed by triosephosphate isomerase. Biochemistry 15, 5627–5631 (1976)
Blacklow, S. C., Raines, R. T., Lim, W. A., Zamore, P. D. & Knowles, J. R. Triosephosphate isomerase catalysis is diffusion controlled. Biochemistry 27, 1158–1165 (1988)
Knowles, J. R. Enzyme catalysis: not different, just better. Nature 350, 121–124 (1991)
Wierenga, R. K., Kapetaniou, E. G. & Venkatesan, R. Triosephosphate isomerase: A highly evolved biocatalyst. Cell. Mol. Life Sci. 67, 3961–3982 (2010)
Richard, J. P. A paradigm for enzyme-catalyzed proton transfer at carbon: Triosephosphate isomerase. Biochemistry 51, 2652–2661 (2012)
Casey, M. L., Kemp, D. S., Paul, K. G. & Cox, D. D. Physical organic-chemistry of benzisoxazoles. 1. Mechanism of base-catalyzed decomposition of benzisoxazoles. J. Org. Chem. 38, 2294–2301 (1973)
Richard, J. P. Acid-base catalysis of the elimination and isomerization-reactions of triose phosphates. J. Am. Chem. Soc. 106, 4926–4936 (1984)
Thorn, S. N., Daniels, R. G., Auditor, M. T. M. & Hilvert, D. Large rate accelerations in antibody catalysis by strategic use of haptenic charge. Nature 373, 228–230 (1995)
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008)
Korendovych, I. V. et al. Design of a switchable eliminase. Proc. Natl Acad. Sci. USA 108, 6823–6827 (2011)
Khersonsky, O. et al. Bridging the gaps in design methodologies by evolutionary optimization of the stability and proficiency of designed Kemp eliminase KE59. Proc. Natl Acad. Sci. USA 109, 10358–10363 (2012)
Putman, S. J., Coulson, A. F. W., Farley, I. R. T., Riddleston, B. & Knowles, J. R. Specificity and kinetics of triose phosphate isomerase from chicken muscle. Biochem. J. 129, 301–310 (1972)
Wolfenden, R. Enzyme catalysis: Conflicting requirements of substrate access and transition-state affinity. Mol. Cell. Biochem. 3, 207–211 (1974)
Herschlag, D. The role of induced fit and conformational changes of enzymes in specificity and catalysis. Bioorg. Chem. 16, 62–96 (1988)
Hu, Y., Houk, K. N., Kikuchi, K., Hotta, K. & Hilvert, D. Nonspecific medium effects versus specific group positioning in the antibody and albumin catalysis of the base-promoted ring-opening reactions of benzisoxazoles. J. Am. Chem. Soc. 126, 8197–8205 (2004)
Debler, E. W. et al. Structural origins of efficient proton abstraction from carbon by a catalytic antibody. Proc. Natl Acad. Sci. USA 102, 4984–4989 (2005)
Gandour, R. D. On the importance of orientation in general base catalysis by carboxylate. Bioorg. Chem. 10, 169–176 (1981)
Bryan, P., Pantoliano, M. W., Quill, S. G., Hsiao, H. Y. & Poulos, T. Site-directed mutagenesis and the role of the oxyanionhole in subtilisin. Proc. Natl Acad. Sci. USA 83, 3743–3745 (1986)
Na, J., Houk, K. N. & Hilvert, D. Transition state of the base-promoted ring-opening of isoxazoles. Theoretical prediction of catalytic functionalities and design of haptens for antibody production. J. Am. Chem. Soc. 118, 6462–6471 (1996)
Frushicheva, M. P., Cao, J., Chu, Z. T. & Warshel, A. Exploring challenges in rational enzyme design by simulating the catalysis in artificial Kemp eliminase. Proc. Natl Acad. Sci. USA 107, 16869–16874 (2010)
Albery, W. J. & Knowles, J. R. Evolution of enzyme function and the development of catalytic efficiency. Biochemistry 15, 5631–5640 (1976)
Voigt, C. A., Gordon, D. B. & Mayo, S. L. Trading accuracy for speed: A quantitative comparison of search algorithms in protein sequence design. J. Mol. Biol. 299, 789–803 (2000)
Warshel, A. et al. Electrostatic basis for enzyme catalysis. Chem. Rev. 106, 3210–3235 (2006)
Allen, B. D., Nisthal, A. & Mayo, S. L. Experimental library screening demonstrates the successful application of computational protein design to large structural ensembles. Proc. Natl Acad. Sci. USA 107, 19838–19843 (2010)
Kiss, G., Röthlisberger, D., Baker, D. & Houk, K. N. Evaluation and ranking of enzyme designs. Protein Sci. 19, 1760–1773 (2010)
Kiss, G., Çelebi-Ölçüm, N., Moretti, R., Baker, D. & Houk, K. N. Computational enzyme design. Angew. Chem. Int. Ed. Engl. 52, 5700–5725 (2013)
Kries, H., Blomberg, R. & Hilvert, D. De novo enzymes by computational design. Curr. Opin. Chem. Biol. 17, 221–228 (2013)
Lassila, J. K., Privett, H. K., Allen, B. D. & Mayo, S. L. Combinatorial methods for small-molecule placement in computational enzyme design. Proc. Natl Acad. Sci. USA 103, 16710–16715 (2006)
Leung, D. W., Chen, E. & Goeddel, D. V. A method for random mutagenesis of a defined DNA segment using a modified polymerase chain reaction. Technique 1, 11–15 (1989)
Stemmer, W. P. C. DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc. Natl Acad. Sci. USA 91, 10747–10751 (1994)
Reetz, M. T., Kahakeaw, D. & Lohmer, R. Addressing the numbers problem in directed evolution. ChemBioChem 9, 1797–1804 (2008)
Reetz, M. T. & Carballeira, J. D. Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nature Protocols 2, 891–903 (2007)
Kibbe, W. A. OligoCalc: An online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007)
Kille, S. et al. Reducing codon redundancy and screening effort of combinatorial protein libraries created by saturation mutagenesis. ACS Synth. Biol. 2, 83–92 (2013)
Gamper, M., Hilvert, D. & Kast, P. Probing the role of the C-terminus of Bacillus subtilis chorismate mutase by a novel random protein-termination strategy. Biochemistry 39, 14087–14094 (2000)
Iranpoor, N., Firouzabadi, H. & Nowrouzi, N. A novel method for the highly efficient synthesis of 1,2-benzisoxazoles under neutral conditions using the Ph3P/DDQ system. Tetrahedr. Lett. 47, 8247–8250 (2006)
Kikuchi, K., Hannak, R. B., Guo, M. J., Kirby, A. J. & Hilvert, D. Toward bifunctional antibody catalysis. Bioorg. Med. Chem. 14, 6189–6196 (2006)
Cheng, Y.-C. & Prusoff, W. H. Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Chovancova, E. et al. CAVER 3.0: A tool for the analysis of transport pathways in dynamic protein structures. PLOS Comput. Biol. 8, e1002708 (2012)
Morozov, A. V., Kortemme, T., Tsemekhman, K. & Baker, D. Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations. Proc. Natl Acad. Sci. USA 101, 6946–6951 (2004)
Acknowledgements
The authors are grateful to A. Aires-Trapote and C. Mayer for experimental assistance. We also thank C. Stutz and B. Blattmann for help in protein crystallization, and the beamline staff at the Swiss Light Source for support during data collection. This work was supported by the Swiss National Science Foundation (SNSF), the National Center of Excellence in Research (NCCR) Structural Biology program of the SNSF, the ETH Zurich, and the Defense Advanced Research Projects Agency (DARPA). Fellowships from the Fonds des Verbandes der chemischen Industrie (to R.B.), the Stipendienfonds der Schweizer Chemischen Industrie (to H.K.), the Studienstiftung des deutschen Volkes (to R.B. and H.K.), and the National Security Science and Engineering Faculty Fellowship (to S.L.M.) are gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
D.H., M.G.G., S.L.M., H.K.P., P.R.E.M., D.M.P., H.K. and R.B. designed the experiments. R.B. and H.K. evolved and biochemically characterized the variants; D.M.P. and P.R.E.M. crystallized the proteins and solved their structures. The manuscript and figures were prepared by R.B., H.K., D.M.P. and D.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Figures 1-4. (PDF 1427 kb)
Supplementary Data
This file contains source data for Supplementary Figure 3. (XLSX 44 kb)
Source data
Rights and permissions
About this article
Cite this article
Blomberg, R., Kries, H., Pinkas, D. et al. Precision is essential for efficient catalysis in an evolved Kemp eliminase. Nature 503, 418–421 (2013). https://doi.org/10.1038/nature12623
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12623
This article is cited by
-
A non-canonical nucleophile unlocks a new mechanistic pathway in a designed enzyme
Nature Communications (2024)
-
Designed active-site library reveals thousands of functional GFP variants
Nature Communications (2023)
-
Directed evolution of a wax ester synthase for production of fatty acid ethyl esters in Saccharomyces cerevisiae
Applied Microbiology and Biotechnology (2023)
-
Building enzymes from scratch
Nature Chemistry (2022)
-
NMR-guided directed evolution
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.