Abstract
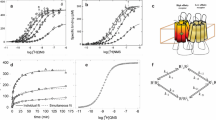
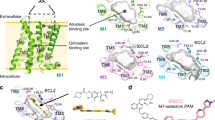
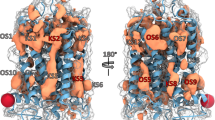
The design of G-protein-coupled receptor (GPCR) allosteric modulators, an active area of modern pharmaceutical research, has proved challenging because neither the binding modes nor the molecular mechanisms of such drugs are known1,2. Here we determine binding sites, bound conformations and specific drug–receptor interactions for several allosteric modulators of the M2 muscarinic acetylcholine receptor (M2 receptor), a prototypical family A GPCR, using atomic-level simulations in which the modulators spontaneously associate with the receptor. Despite substantial structural diversity, all modulators form cation–π interactions with clusters of aromatic residues in the receptor extracellular vestibule, approximately 15 Å from the classical, ‘orthosteric’ ligand-binding site. We validate the observed modulator binding modes through radioligand binding experiments on receptor mutants designed, on the basis of our simulations, either to increase or to decrease modulator affinity. Simulations also revealed mechanisms that contribute to positive and negative allosteric modulation of classical ligand binding, including coupled conformational changes of the two binding sites and electrostatic interactions between ligands in these sites. These observations enabled the design of chemical modifications that substantially alter a modulator’s allosteric effects. Our findings thus provide a structural basis for the rational design of allosteric modulators targeting muscarinic and possibly other GPCRs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Conn, P. J., Jones, C. K. & Lindsley, C. W. Subtype selective allosteric modulators of muscarinic receptors for the treatment of CNS disorders. Trends Pharmacol. Sci. 30, 148–155 (2009)
Keov, P., Sexton, P. M. & Christopoulos, A. Allosteric modulation of G protein-coupled receptors: a pharmacological perspective. Neuropharmacology 60, 24–35 (2011)
Filmore, D. It’s a GPCR world. Modern Drug Discov. 7, 24–28 (2004)
Jakubik, J. & El-Fakahany, E. E. Allosteric modulation of muscarinic acetylcholine receptors. Pharmaceuticals 3, 2838–2860 (2010)
Haga, K. et al. Structure of human M2 muscarinic acetylcholine receptor bound to an antagonist. Nature 482, 547–551 (2012)
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 6091, 232–236 (2012)
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011)
Lazareno, S. & Birdsall, N. J. Detection, quantitation, and verification of allosteric interactions of agents with labeled and unlabeled ligands at G protein-coupled receptors: interactions of strychnine and acetylcholine at muscarinic receptors. Mol. Pharmacol. 48, 362–378 (1995)
Dror, R. O. et al. Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc. Natl Acad. Sci. USA 108, 13118–13123 (2011)
Shan, Y. et al. How does a drug molecule find its target binding site? J. Am. Chem. Soc. 133, 9181–9183 (2011)
Buch, I., Giorgino, T. & De Fabritiis, G. Complete reconstruction of an enzyme-inhibitor binding process by molecular dynamics simulations. Proc. Natl Acad. Sci. USA 108, 10184–10189 (2011)
Prilla, S., Schrobang, J., Ellis, J., Höltje, H. D. & Mohr, K. Allosteric interactions with muscarinic acetylcholine receptors: complex role of the conserved tryptophan M2422Trp in a cryptical cluster of amino acids for baseline affinity, subtype selectivity, and cooperativity. Mol. Pharmacol. 70, 181–193 (2006)
Huang, X.-P., Prilla, S., Mohr, K. & Ellis, J. Critical amino acid residues of the common allosteric site on the M2 muscarinic acetylcholine receptor. Mol. Pharmacol. 68, 769–778 (2005)
May, L. T. et al. Structure-function studies of allosteric agonism at M2 muscarinic acetylcholine receptors. Mol. Pharmacol. 72, 463–476 (2007)
Trankle, C. et al. Interactions of orthosteric and allosteric ligands with [3H]dimethyl-W84 at the common allosteric site of muscarinic M2 receptors. Mol. Pharmacol. 64, 180–190 (2003)
Ballesteros, J. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G-protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Matsui, H., Lazareno, S. & Birdsall, N. J. Probing of the location of the allosteric site on m1 muscarinic receptors by site-directed mutagenesis. Mol. Pharmacol. 47, 88–98 (1995)
Ma, L. et al. Selective activation of the M1 muscarinic acetylcholine receptor achieved by allosteric potentiation. Proc. Natl Acad. Sci. USA 106, 15950–15955 (2009)
Daiss, J. O. et al. N+/Si replacement as a tool for probing the pharmacophore of allosteric modulators of muscarinic M2 receptors: synthesis, allosteric potency, and positive cooperativity of silicon-based W84 derivatives. Organometallics 21, 803–811 (2002)
Choe, H. W. et al. Crystal structure of metarhodopsin II. Nature 471, 651–655 (2011)
Bock, A. et al. The allosteric vestibule of a seven transmembrane helical receptor controls G-protein coupling. Nat. Commun. 3, 1044 (2012)
Shoichet, B. & Kobilka, B. Structure-based drug screening for G-protein-coupled receptors. Trends Pharmacol. Sci. 33, 268–272 (2012)
Totrov, M. & Abagyan, R. Flexible ligand docking to multiple receptor conformations: a practical alternative. Curr. Opin. Struct. Biol. 18, 178–184 (2008)
Avlani, V., May, L. T., Sexton, P. M. & Christopoulos, A. Application of a kinetic model to the apparently complex behavior of negative and positive allosteric modulators of muscarinic acetylcholine receptors. J. Pharmacol. Exp. Ther. 308, 1062–1072 (2004)
Gao, Z.-G. et al. Identification of essential residues involved in the allosteric modulation of the human A3 adenosine receptor. Mol. Pharmacol. 63, 1021–1031 (2003)
Silvano, E. et al. The tetrahydroisoquinoline derivative SB269,652 is an allosteric antagonist at dopamine D3 and D2 receptors. Mol. Pharmacol. 78, 925–934 (2010)
Lazareno, S., Popham, A. & Birdsall, N. J. Analogs of WIN 62,577 define a second allosteric site on muscarinic receptors. Mol. Pharmacol. 62, 1492–1505 (2002)
Yanamala, N. & Klein-Seetharaman, J. Allosteric modulation of G protein coupled receptors by cytoplasmic, transmembrane, and extracellular ligands. Pharmaceuticals 3, 3324–3342 (2010)
Shaw, D. E. et al. Millisecond-scale molecular dynamics simulation on Anton. In Proceedings of the Conference on High Performance Computing, Networking, Storage, and Analysis (ACM Press, 2009); available at http://dl.acm.org/citation.cfm?id=1654099 (2009)
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998)
Dror, R. O. et al. Identification of two distinct inactive conformations of the β2-adrenergic receptor reconciles structural and biochemical observations. Proc. Natl Acad. Sci. USA 106, 4689–4694 (2009)
Fahmy, K. et al. Protonation states of membrane-embedded carboxylic acid groups in rhodopsin and metarhodopsin II: a Fourier-transform infrared spectroscopy study of site-directed mutants. Proc. Natl Acad. Sci. USA 90, 10206–10210 (1993)
Everett, A. J., Openshaw, H. T. & Smith, G. F. The constitution of aspidospermine. Part III. Reactivity at the nitrogen atoms, and biogenetic considerations. J. Chem. Soc. 1120–1123. (1957)
Rosenbaum, D. M. et al. Structure and function of an irreversible agonist–β2 adrenoceptor complex. Nature 469, 236–240 (2011)
Kruse, A. et al. Structure and dynamics of the M3 muscarinic acetylcholine receptor. Nature 482, 552–556 (2012)
Shaw, D. E. et al. Atomic-level characterization of the structural dynamics of proteins. Science 330, 341–346 (2010)
Kräutler, V., van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001)
Tuckerman, M., Berne, B. J. & Martyna, G. J. Reversible multiple time scale molecular dynamics. J. Chem. Phys. 97, 1990–2001 (1992)
Shan, Y., Klepeis, J. L., Eastwood, M. P., Dror, R. O. & Shaw, D. E. Gaussian split Ewald: a fast Ewald mesh method for molecular simulation. J. Chem. Phys. 122, 54101 (2005)
Bourne, P. E., Ginell, S. L., Low, B. W. & Lessinger, L. Structure of a potent neuromuscular blocking agent: caracurine-II dimethochloride octahydrate, [C40H44N4O2]2+·2Cl−·8H2O. J. Cryst. Spectroscop. Res. 15, 453–471 (1985)
DeLano, W. L. The PyMOL Molecular Graphics System v. 1.5.0.3-01 (Schrödinger, LLC, New York, New York, 2012)
Mackerell, A. D., Jr, Feig, M. & Brooks, C. L., III Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004)
Piana, S., Lindorff-Larsen, K. & Shaw, D. E. How robust are protein folding simulations with respect to force field parameterization? Biophys. J. 100, L47–L49 (2011)
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010)
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010)
Caldwell, J. & Kollman, P. Cation–π interactions: nonadditive effects are critical in their accurate representation. J. Am. Chem. Soc. 117, 4177–4178 (1995)
Schneider, H. et al. Host-guest supramolecular chemistry. 34. The incremental approach to noncovalent interactions: Coulomb and van der Waals effects in organic ion pairs. J. Am. Chem. Soc. 20, 7698–7703 (1991)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Gnagey, A. L., Seidenberg, M. & Ellis, J. Site-directed mutagenesis reveals two epitopes involved in the subtype selectivity of the allosteric interactions of gallamine at muscarinic acetylcholine receptors. Mol. Pharmacol. 56, 1245–1253 (1999)
Voigtländer, U. et al. Allosteric site on muscarinic acetylcholine receptors: identification of two amino acids in the muscarinic M2 receptor that account entirely for the M2/M5 subtype selectivities of some structurally diverse allosteric ligands in N-methylscopolamine-occupied receptors. Mol. Pharmacol. 64, 21–31 (2003)
Avlani, V. A. et al. Critical role for the second extracellular loop in the binding of both orthosteric and allosteric G protein-coupled receptor ligands. J. Biol. Chem. 282, 25677–25686 (2007)
Acknowledgements
We thank T. Mildorf, A. Kruse and B. Kobilka for comments; J. Klepeis, B. Gregersen, J.-L. Li, K. Palmo, A. Donchev and particularly A. Taube for advice and support related to force fields and quantum mechanical calculations; Z. Fan for assistance with statistical analysis; A. Lerer and T. O’Donnell for assistance with simulation and analysis software; A. Philippsen for creating the video; A. Stewart for assistance with mutagenesis and cell line generation; K. Ban and J. Harjani for assistance with chemical synthesis; J. Swarbrick for recording and analysing the two-dimensional NMR data; B. Sleebs and S. Marcuccio for provision of synthetic reagents; J. Dang for advice on analytical chemistry; and M. Kirk and R. Kastleman for editorial assistance. Portions of this work were financed by Program Grant no. 519461 from the National Health and Medical Research Council (NHMRC) of Australia, with synthetic chemistry infrastructure support from the Australian Federal Education Investment Fund Super Science Initiative and Victoria’s Science Agenda Investment Fund. A.C. and P.M.S. are Principal Research Fellows of the NHMRC; J.B.B. is a Senior Research Fellow of the NHMRC; J.R.L. is a Career Development Awardee of the NHMRC.
Author information
Authors and Affiliations
Contributions
R.O.D. conceived this study and, with D.E.S., oversaw molecular dynamics simulations and analysis. R.O.D., H.F.G., D.W.B., J.R.V., A.C.P. and D.H.A. designed and analysed molecular dynamics simulations. H.F.G., J.R.V., A.C.P. and D.H.A. performed molecular dynamics simulations. R.O.D., H.F.G., D.W.B. and J.R.V. performed computational design of receptor mutants and of the modulator 4P-C7/3-phth. C.V. performed all biological assays and, with J.R.V. and A.C., analysed experimental data. M.C. and J.R.L. performed mutagenesis and generated the stable cell lines. J.B.B. and R.R. designed, planned and executed the synthesis of 4P-C7/3-phth, with active input from D.W.B. P.M.S. and A.C. supervised the cell-based biological studies. R.O.D., H.F.G., D.W.B., A.C. and D.E.S. wrote the manuscript. R.O.D., A.C. and D.E.S. supervised the overall research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-25, Supplementary Tables 1-8, Supplementary Methods and additional references. (PDF 5767 kb)
The allosteric modulator C7/3-phth binds spontaneously to the M2 receptor in an unbiased molecular dynamics simulation
For clarity, the lipid bilayer, ions, and water molecules are not shown. The video speeds up 44-fold after the modulator binds at 120 ns; the amount of simulated time between successive video frames is 1.08 ns before this point and 47.5 ns afterwards. The Cartesian components of the protein Cα positions were smoothed using Fourier-based Gaussian smoothing (σ = 7.2 ns). Ligand coordinates were not smoothed before the ligand bound, but once it bound, both the Cartesian components of its atom positions and its internal angles were smoothed using Gaussian filters (σ = 7.2 ns). The video was created using OpenStructure17. This is simulation 1 under condition A (Supplementary Table 2). (MP4 2760 kb)
Rights and permissions
About this article
Cite this article
Dror, R., Green, H., Valant, C. et al. Structural basis for modulation of a G-protein-coupled receptor by allosteric drugs. Nature 503, 295–299 (2013). https://doi.org/10.1038/nature12595
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12595
This article is cited by
-
Fly casting with ligand sliding and orientational selection supporting complex formation of a GPCR and a middle sized flexible molecule
Scientific Reports (2022)
-
Allostery of atypical modulators at oligomeric G protein-coupled receptors
Scientific Reports (2021)
-
A lysine–cysteine redox switch with an NOS bridge regulates enzyme function
Nature (2021)
-
Cryptic pocket formation underlies allosteric modulator selectivity at muscarinic GPCRs
Nature Communications (2019)
-
A mechanism for the activation of the mechanosensitive Piezo1 channel by the small molecule Yoda1
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.