Abstract
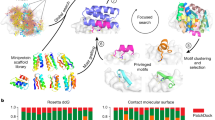
The ability to design proteins with high affinity and selectivity for any given small molecule is a rigorous test of our understanding of the physiochemical principles that govern molecular recognition. Attempts to rationally design ligand-binding proteins have met with little success, however, and the computational design of protein–small-molecule interfaces remains an unsolved problem1. Current approaches for designing ligand-binding proteins for medical2 and biotechnological uses rely on raising antibodies against a target antigen in immunized animals3,4 and/or performing laboratory-directed evolution of proteins with an existing low affinity for the desired ligand5,6,7, neither of which allows complete control over the interactions involved in binding. Here we describe a general computational method for designing pre-organized and shape complementary small-molecule-binding sites, and use it to generate protein binders to the steroid digoxigenin (DIG). Of seventeen experimentally characterized designs, two bind DIG; the model of the higher affinity binder has the most energetically favourable and pre-organized interface in the design set. A comprehensive binding-fitness landscape of this design, generated by library selections and deep sequencing, was used to optimize its binding affinity to a picomolar level, and X-ray co-crystal structures of two variants show atomic-level agreement with the corresponding computational models. The optimized binder is selective for DIG over the related steroids digitoxigenin, progesterone and β-oestradiol, and this steroid binding preference can be reprogrammed by manipulation of explicitly designed hydrogen-bonding interactions. The computational design method presented here should enable the development of a new generation of biosensors, therapeutics and diagnostics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schreier, B., Stumpp, C., Wiesner, S. & Höcker, B. Computational design of ligand binding is not a solved problem. Proc. Natl Acad. Sci. USA 106, 18491–18496 (2009)
de Wolf, F. A. & Brett, G. M. Ligand-binding proteins: their potential for application in systems for controlled delivery and uptake of ligands. Pharmacol. Rev. 52, 207–236 (2000)
Hunter, M. M., Margolies, M. N., Ju, A. & Haber, E. High-affinity monoclonal antibodies to the cardiac glycoside, digoxin. J. Immunol. 129, 1165–1172 (1982)
Shen, X. Y., Orson, F. M. & Kosten, T. R. Vaccines against drug abuse. Clin. Pharmacol. Ther. 91, 60–70 (2012)
Bradbury, A. R. M., Sidhu, S., Dübel, S. & McCafferty, J. Beyond natural antibodies: the power of in vitro display technologies. Nature Biotechnol. 29, 245–254 (2011)
Brustad, E. M. & Arnold, F. H. Optimizing non-natural protein function with directed evolution. Curr. Opin. Chem. Biol. 15, 201–210 (2011)
Chen, G. et al. Isolation of high-affinity ligand-binding proteins by periplasmic expression with cytometric screening (PECS). Nature Biotechnol. 19, 537–542 (2001)
Telmer, P. G. & Shilton, B. H. Structural studies of an engineered zinc biosensor reveal an unanticipated mode of zinc binding. J. Mol. Biol. 354, 829–840 (2005)
Baker, D. An exciting but challenging road ahead for computational enzyme design. Protein Sci. 19, 1817–1819 (2010)
Jiang, L. et al. De novo computational design of retro-Aldol enzymes. Science 319, 1387–1391 (2008)
Khare, S. D. & Fleishman, S. J. Emerging themes in the computational design of novel enzymes and protein–protein interfaces. FEBS Lett. 587, 1147–1154 (2013)
Khersonsky, O. et al. Bridging the gaps in design methodologies by evolutionary optimization of the stability and proficiency of designed Kemp eliminase KE59. Proc. Natl Acad. Sci. USA 109, 10358–10363 (2012)
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008)
Wang, L. et al. Structural analyses of covalent enzyme–substrate analog complexes reveal strengths and limitations of de novo enzyme design. J. Mol. Biol. 415, 615–625 (2012)
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nature Chem. Biol. 5, 789–796 (2009)
Fleishman, S. J., Khare, S. D., Koga, N. & Baker, D. Restricted sidechain plasticity in the structures of native proteins and complexes. Protein Sci. 20, 753–757 (2011)
Zanghellini, A. et al. New algorithms and an in silico benchmark for computational enzyme design. Protein Sci. 15, 2785–2794 (2006)
Lawrence, M. C. & Colman, P. M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993)
The Digitalis Investigation Group. The effect of digoxin on mortality and morbidity in patients with heart failure. N. Engl. J. Med. 336, 525–533 (1997)
Eisel, D., Seth, O., Grünewald-Janho, S. & Kruchen, B. DIG Application Manual for Nonradioactive in situ Hybridization 4th edn (Roche Diagnostics, 2008)
Flanagan, R. J. & Jones, A. L. Fab antibody fragments: some applications in clinical toxicology. Drug Saf. 27, 1115–1133 (2004)
Chao, G. et al. Isolating and engineering human antibodies using yeast surface display. Nature Protocols 1, 755–768 (2006)
Fowler, D. M. et al. High-resolution mapping of protein sequence-function relationships. Nature Methods 7, 741–746 (2010)
McLaughlin, R. N., Jr, Poelwijk, F. J., Raman, A., Gosal, W. S. & Ranganathan, R. The spatial architecture of protein function and adaptation. Nature 491, 138–142 (2012)
Whitehead, T. A. et al. Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing. Nature Biotechnol. 30, 543–548 (2012)
Fersht, A. R. et al. Hydrogen bonding and biological specificity analysed by protein engineering. Nature 314, 235–238 (1985)
Frederick, K. K., Marlow, M. S., Valentine, K. G. & Wand, A. J. Conformational entropy in molecular recognition by proteins. Nature 448, 325–329 (2007)
Fleishman, S. J. & Baker, D. Role of the biomolecular energy gap in protein design, structure, and evolution. Cell 149, 262–273 (2012)
Kuhlman, B. & Baker, D. Native protein sequences are close to optimal for their structures. Proc. Natl Acad. Sci. USA 97, 10383–10388 (2000)
Rossi, A. M. & Taylor, C. W. Analysis of protein-ligand interactions by fluorescence polarization. Nature Protocols 6, 365–387 (2011)
Fleishman, S. J. et al. RosettaScripts: A scripting language interface to the Rosetta macromolecular modeling suite. PLoS ONE 6, e20161 (2011)
Siegel, J. B. et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels–Alder reaction. Science 329, 309–313 (2010)
Richter, F., Leaver-Fay, A., Khare, S. D., Bjelic, S. & Baker, D. De novo enzyme design using Rosetta3. PLoS ONE 6, e19230 (2011)
Kellogg, E. H., Leaver-Fay, A. & Baker, D. Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins 79, 830–838 (2011)
Cooper, S. et al. Predicting protein structures with a multiplayer online game. Nature 466, 756–760 (2010)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Benatuil, L., Perez, J. M., Belk, J. & Hsieh, C.-M. An improved yeast transformation method for the generation of very large human antibody libraries. Protein Eng. Des. Sel. 23, 155–159 (2010)
Fowler, D. M., Araya, C. L., Gerard, W. & Fields, S. Enrich: software for analysis of protein function by enrichment and depletion of variants. Bioinformatics 27, 3430–3431 (2011)
Acknowledgements
We thank E.-M. Strauch and P.-S. Huang for providing the ZZ/pETCON and S2/pETCON plasmids, respectively, and B. Shen for assistance with data processing, modelling and refinement of the X-ray crystal structures. We thank J. P. Sumida for assistance with analytical ultracentrifugation data collection, processing, and analysis. Analytical ultracentrifugation experiments were performed at the University of Washington Analytical Biopharmacy Core, which is supported by the Washington State Life Sciences Discovery Fund and the Center for the Intracellular Delivery of Biologics. We thank S. Fleishman, O. Khersonsky and P.-S. Huang for comments on the manuscript. J.W.N. acknowledges a pre-doctoral fellowship from the National Human Genome Research Institute under the Interdisciplinary Training in Genome Sciences program (T32 HG00035). This work was supported by grants from DTRA and DARPA to D.B., a grant from the Swiss National Science Foundation to K.J., and National Science Foundation grant MCB1121896 to C.G.K.
Author information
Authors and Affiliations
Contributions
C.E.T., S.D.K. and D.B. designed the research. S.D.K., C.E.T. and J.D. performed computations. S.D.K. wrote program code. C.E.T. experimentally characterized the designs, constructed libraries, performed affinity maturation and deep sequencing selections, and conducted binding and biochemical studies of DIG10. J.D. characterized DIG5. J.W.N. analysed deep sequencing data. C.E.T. and J.D. prepared protein samples for crystallographic analysis. L.D. and B.S. crystallized the protein samples and analysed crystallographic data. A.S. and K.J. synthesized DIG-PEG3-biotin and DIG-PEG3-Alexa488. C.E.T. prepared protein samples for ITC studies, and W.J. and C.G.K. performed ITC experiments and analysed ITC data. C.E.T., S.D.K. and D.B. analysed data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figure legends 1-2, Supplementary Methods, Supplementary Tables 1-22, Supplementary Figures 1-21 with legends, and Supplementary Data. Supplementary Methods contains additional computational and experimental methods. Supplementary Tables contain information about design metrics, experimental observations and statistics, and primer sequences. Supplementary Figures provide additional experimental results. Supplementary Data contains command lines and protocols for running design calculations. (PDF 12410 kb)
Rights and permissions
About this article
Cite this article
Tinberg, C., Khare, S., Dou, J. et al. Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501, 212–216 (2013). https://doi.org/10.1038/nature12443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12443
This article is cited by
-
Drug design on quantum computers
Nature Physics (2024)
-
Design of allosteric sites into rotary motor V1-ATPase by restoring lost function of pseudo-active sites
Nature Chemistry (2023)
-
De novo prediction of explicit water molecule positions by a novel algorithm within the protein design software MUMBO
Scientific Reports (2023)
-
Protein sequence design with a learned potential
Nature Communications (2022)
-
Rapid biosensor development using plant hormone receptors as reprogrammable scaffolds
Nature Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.