Abstract
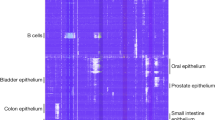
DNA methylation is a defining feature of mammalian cellular identity and is essential for normal development1,2. Most cell types, except germ cells and pre-implantation embryos3,4,5, display relatively stable DNA methylation patterns, with 70–80% of all CpGs being methylated6. Despite recent advances, we still have a limited understanding of when, where and how many CpGs participate in genomic regulation. Here we report the in-depth analysis of 42 whole-genome bisulphite sequencing data sets across 30 diverse human cell and tissue types. We observe dynamic regulation for only 21.8% of autosomal CpGs within a normal developmental context, most of which are distal to transcription start sites. These dynamic CpGs co-localize with gene regulatory elements, particularly enhancers and transcription-factor-binding sites, which allow identification of key lineage-specific regulators. In addition, differentially methylated regions (DMRs) often contain single nucleotide polymorphisms associated with cell-type-related diseases as determined by genome-wide association studies. The results also highlight the general inefficiency of whole-genome bisulphite sequencing, as 70–80% of the sequencing reads across these data sets provided little or no relevant information about CpG methylation. To demonstrate further the utility of our DMR set, we use it to classify unknown samples and identify representative signature regions that recapitulate major DNA methylation dynamics. In summary, although in theory every CpG can change its methylation state, our results suggest that only a fraction does so as part of coordinated regulatory programs. Therefore, our selected DMRs can serve as a starting point to guide new, more effective reduced representation approaches to capture the most informative fraction of CpGs, as well as further pinpoint putative regulatory elements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
Gene Expression Omnibus
Data deposits
WGBS data are deposited at the Gene Expression Omnibus (see Supplementary Table 1 for the specific accession numbers). Supplementary Table 2 is available under GEO accession number GSE46644.
References
Bestor, T. H. The DNA methyltransferases of mammals. Hum. Mol. Genet. 9, 2395–2402 (2000)
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–432 (2007)
Seisenberger, S. et al. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 48, 849–862 (2012)
Smith, Z. D. et al. A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 (2012)
Hackett, J. A. & Surani, M. A. DNA methylation dynamics during the mammalian life cycle. Phil. Trans. R. Soc. B 368, 20110328 (2013)
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002)
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nature Rev. Genet. 14, 204–220 (2013)
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nature Struct. Mol. Biol. 20, 274–281 (2013)
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009)
Nazor, K. L. et al. Recurrent variations in DNA methylation in human pluripotent stem cells and their differentiated derivatives. Cell Stem Cell 10, 620–634 (2012)
Weber, M. et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nature Genet. 37, 853–862 (2005)
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008)
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010)
Varley, K. E. et al. Dynamic DNA methylation across diverse human cell lines and tissues. Genome Res. 23, 555–567 (2013)
Irizarry, R. A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nature Genet. 41, 178–186 (2009)
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nature Genet. 44, 40–46 (2012)
Cohen, N. M., Kenigsberg, E. & Tanay, A. Primate CpG islands are maintained by heterogeneous evolutionary regimes involving minimal selection. Cell 145, 773–786 (2011)
Lienert, F. et al. Identification of genetic elements that autonomously determine DNA methylation states. Nature Genet. 43, 1091–1097 (2011)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012)
Zhu, J. et al. Genome-wide chromatin state transitions associated with developmental and environmental cues. Cell 152, 642–654 (2013)
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012)
Ravasi, T. et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140, 744–752 (2010)
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011)
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012)
Bell, C. G. et al. Human-specific CpG “beacons” identify loci associated with human-specific traits and disease. Epigenetics 7, 1188–1199 (2012)
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009)
Ehrlich, M. DNA hypomethylation in cancer cells. Epigenomics 1, 239–259 (2009)
Dedeurwaerder, S. et al. Evaluation of the Infinium Methylation 450K technology. Epigenomics 3, 771–784 (2011)
Gnirke, A. et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nature Biotechnol. 27, 182–189 (2009)
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008)
Acknowledgements
We would like to thank K. Clement, P. Samavarchi-Tehrani, Z. Smith, M. Chan and R. Karnik for discussions and feedback. We would also like to thank F. Kelley, T. Durham, C. Epstein, N. Shoresh, G. Lauwers and the Massachusetts General Hospital tissue repository for assisting in sample and data management. E.D.R. is supported by the National Institutes of Health (NIH) Roadmap Epigenomics Project (ES017690). D.A.B. is supported by NIH grants P30AG10161, R01AG17917, R01AG15819 and R01AG36042. A.M. is supported by the Pew Charitable Trusts and is a New York Stem Cell Foundation, Robertson Investigator. This work was funded by NIH grants (U01ES017155 and P01GM099117) and The New York Stem Cell Foundation.
Author information
Authors and Affiliations
Contributions
M.J.Z. and A.M. conceived the study and interpreted the results. M.J.Z. designed the statistical framework, analysis strategy and analysed the data. H.G. performed in-house WGBS library production, F.M. contributed bioinformatics tools and J.D. performed cell culture experiments. L.T.-Y.T. and E.D.R. provided adipocyte nuclei for WGBS profiling, and P.L.D. and D.A.B. made adult brain and Alzheimer’s disease samples available. O.K. provided support on analysis strategy and statistical methods. B.E.B. and A.M. organized samples as part of the NIH Roadmap Epigenomics Project. H.G., A.G. and A.M. established the WGBS at the Broad Institute. A.M. supervised the project. M.Z. and A.M. wrote the paper with assistance from the other authors.
Corresponding author
Ethics declarations
Competing interests
M.J.Z. and A.M. declare competing financial interests owing to the filing of a patent application on the selected regions.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5, legends for Supplementary Tables 1-3 (see separate files for Supplementary Tables 1 and 3 and for Supplementary Table 2, see link in main paper), Supplementary Methods and additional references. (PDF 486 kb)
Supplementary Table 1
This file contains a summary of data sets, accession numbers and quality measures for all WGBS libraries use in this study. (XLSX 13 kb)
Supplementary Table 3
This file contains Motif enrichment results for cell type specific hypomethylated regions. (XLSX 240 kb)
Supplementary Data
This file contains the data associated with this paper. (ZIP 103 kb)
Rights and permissions
About this article
Cite this article
Ziller, M., Gu, H., Müller, F. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013). https://doi.org/10.1038/nature12433
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12433
This article is cited by
-
Increased CpG methylation at the CDH1 locus in inflamed ileal mucosa of patients with Crohn disease
Clinical Epigenetics (2024)
-
DNA methylation haplotype block signatures responding to Staphylococcus aureus subclinical mastitis and association with production and health traits
BMC Biology (2024)
-
The regulation of plasma gelsolin by DNA methylation in ovarian cancer chemo-resistance
Journal of Ovarian Research (2024)
-
Tumor microenvironment deconvolution identifies cell-type-independent aberrant DNA methylation and gene expression in prostate cancer
Clinical Epigenetics (2024)
-
Differential methylation of linoleic acid pathway genes is associated with PTSD symptoms – a longitudinal study with Burundian soldiers returning from a war zone
Translational Psychiatry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.