Abstract
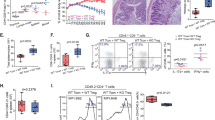
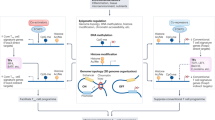
Regulatory T cells (Treg cells) have a crucial role in the immune system by preventing autoimmunity, limiting immunopathology, and maintaining immune homeostasis1. However, they also represent a major barrier to effective anti-tumour immunity and sterilizing immunity to chronic viral infections1. The transcription factor Foxp3 has a major role in the development and programming of Treg cells2,3. The relative stability of Treg cells at inflammatory disease sites has been a highly contentious subject4,5,6. There is considerable interest in identifying pathways that control the stability of Treg cells as many immune-mediated diseases are characterized by either exacerbated or limited Treg-cell function. Here we show that the immune-cell-expressed ligand semaphorin-4a (Sema4a) and the Treg-cell-expressed receptor neuropilin-1 (Nrp1) interact both in vitro, to potentiate Treg-cell function and survival, and in vivo, at inflammatory sites. Using mice with a Treg-cell-restricted deletion of Nrp1, we show that Nrp1 is dispensable for suppression of autoimmunity and maintenance of immune homeostasis, but is required by Treg cells to limit anti-tumour immune responses and to cure established inflammatory colitis. Sema4a ligation of Nrp1 restrained Akt phosphorylation cellularly and at the immunologic synapse by phosphatase and tensin homologue (PTEN), which increased nuclear localization of the transcription factor Foxo3a. The Nrp1-induced transcriptome promoted Treg-cell stability by enhancing quiescence and survival factors while inhibiting programs that promote differentiation. Importantly, this Nrp1-dependent molecular program is evident in intra-tumoral Treg cells. Our data support a model in which Treg-cell stability can be subverted in certain inflammatory sites, but is maintained by a Sema4a–Nrp1 axis, highlighting this pathway as a potential therapeutic target that could limit Treg-cell-mediated tumour-induced tolerance without inducing autoimmunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vignali, D. A., Collison, L. W. & Workman, C. J. How regulatory T cells work. Nature Rev. Immunol. 8, 523–532 (2008)
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nature Immunol. 4, 330–336 (2003)
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003)
Miyao, T. et al. Plasticity of Foxp3+ T cells reflects promiscuous Foxp3 expression in conventional T cells but not reprogramming of regulatory T cells. Immunity 36, 262–275 (2012)
Rubtsov, Y. P. et al. Stability of the regulatory T cell lineage in vivo. Science 329, 1667–1671 (2010)
Zhou, X. et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nature Immunol. 10, 1000–1007 (2009)
Qiao, M., Thornton, A. M. & Shevach, E. M. CD4+ CD25+ regulatory T cells render naive CD4+ CD25- T cells anergic and suppressive. Immunology 120, 447–455 (2007); corrigendum. 121, 146 (2007)
Collison, L. W., Pillai, M. R., Chaturvedi, V. & Vignali, D. A. Regulatory T cell suppression is potentiated by target T cells in a cell contact, IL-35- and IL-10-dependent manner. J. Immunol. 182, 6121–6128 (2009)
Kumanogoh, A. et al. Class IV semaphorin Sema4A enhances T-cell activation and interacts with Tim-2. Nature 419, 629–633 (2002)
Bruder, D. et al. Neuropilin-1: a surface marker of regulatory T cells. Eur. J. Immunol. 34, 623–630 (2004)
Kolodkin, A. L. et al. Neuropilin is a semaphorin III receptor. Cell 90, 753–762 (1997)
Weiss, J. M. et al. Neuropilin 1 is expressed on thymus-derived natural regulatory T cells, but not mucosa-generated induced Foxp3+ T reg cells. J. Exp. Med. 209, 1723–1742 (2012)
Yadav, M. et al. Neuropilin-1 distinguishes natural and inducible regulatory T cells among regulatory T cell subsets in vivo. J. Exp. Med. 209, 1713–1722 (2012)
Milpied, P. et al. Neuropilin-1 is not a marker of human Foxp3+ Treg. Eur. J. Immunol. 39, 1466–1471 (2009)
Piechnik, A. et al. The VEGF receptor neuropilin-1 NRP1) represents a promising novel target for chronic lymphocytic leukemia patients. Int. J. Cancer http://dx.doi.org/10.1002/ijc.28135 (28 February 2013)
Nishikawa, H. & Sakaguchi, S. Regulatory T cells in tumor immunity. Int. J. Cancer 127, 759–767 (2010)
Kim, J. M., Rasmussen, J. P. & Rudensky, A. Y. Regulatory T cells prevent catastrophic autoimmunity throughout the lifespan of mice. Nature Immunol. 8, 191–197 (2007)
Wilde, S. et al. Human antitumor CD8+ T cells producing Th1 polycytokines show superior antigen sensitivity and tumor recognition. J. Immunol. 189, 598–605 (2012)
Demoulin, S., Herfs, M., Delvenne, P. & Hubert, P. Tumor microenvironment converts plasmacytoid dendritic cells into immunosuppressive/tolerogenic cells: insight into the molecular mechanisms. J. Leukoc. Biol. 93, 343–352 (2013)
Castro-Rivera, E., Ran, S., Brekken, R. A. & Minna, J. D. Semaphorin 3B inhibits the phosphatidylinositol 3-kinase/Akt pathway through neuropilin-1 in lung and breast cancer cells. Cancer Res. 68, 8295–8303 (2008)
Crellin, N. K., Garcia, R. V. & Levings, M. K. Altered activation of AKT is required for the suppressive function of human CD4+CD25+ T regulatory cells. Blood 109, 2014–2022 (2007)
Haxhinasto, S., Mathis, D. & Benoist, C. The AKT-mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J. Exp. Med. 205, 565–574 (2008)
Franke, T. F. et al. The protein kinase encoded by the Akt proto-oncogene is a target of the PDGF-activated phosphatidylinositol 3-kinase. Cell 81, 727–736 (1995)
Stambolic, V. et al. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell 95, 29–39 (1998)
Pellet-Many, C., Frankel, P., Jia, H. & Zachary, I. Neuropilins: structure, function and role in disease. Biochem. J. 411, 211–226 (2008)
Merkenschlager, M. & von Boehmer, H. PI3 kinase signalling blocks Foxp3 expression by sequestering Foxo factors. J. Exp. Med. 207, 1347–1350 (2010)
Kerdiles, Y. M. et al. Foxo transcription factors control regulatory T cell development and function. Immunity 33, 890–904 (2010); erratum. 34, 135 (2011)
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nature Immunol. 11, 618–627 (2010)
Heng, T. S. & Painter, M. W. The Immunological Genome Project: networks of gene expression in immune cells. Nature Immunol. 9, 1091–1094 (2008)
Sawant, A. et al. Depletion of plasmacytoid dendritic cells inhibits tumor growth and prevents bone metastasis of breast cancer cells. J. Immunol. 189, 4258–4265 (2012)
Guy, C. S. et al. Distinct TCR signaling pathways drive proliferation and cytokine production in T cells. Nature Immunol. 14, 262–270 (2013)
Acknowledgements
The authors would like to thank E. J. Wherry and H. Chi for advice; A. Rudensky, D. Cheresh, K. Yang, T. Geiger and H. Chi for mice; D. R. Green for plasmids; and B. Triplett, M. Howard and M. McKenna at St Louis Cord Blood Bank for cord blood samples. The authors also thank K. Forbes and A. McKenna for maintenance, breeding and genotyping of mouse colonies; A. Castellaw for preparation of human cord blood samples; K. M. Vignali for assistance with cloning; A. Herrada for generating Nrp1-IgG1; A. L. Szymczak-Workman for assistance with histological analysis; S. Morgan, G. Lennon and R. Cross of the St Jude Immunology Flow Lab for cell sorting; the staff of the Shared Animal Resource Center at St Jude Children's Research Hospital for the animal husbandry; the Hartwell Center for Biotechnology and Bioinformatics at St Jude Children's Research Hospital for Affymetrix microarray analysis; the Veterinary Pathology Core for histological preparation; and the Immunology Department at St Jude Children's Research Hospital for helpful discussions. This work was supported by the National Institutes of Health (R01 AI091977 and AI039480 to D.A.A.V.; F32 AI098383 to G.M.D.), NCI Comprehensive Cancer Center Support CORE grant (CA21765, to D.A.A.V.) and ALSAC (to D.A.A.V.).
Author information
Authors and Affiliations
Contributions
G.M.D. designed and performed most of the experiments and wrote the manuscript. S.-R.W. performed critical initial experiments and identified Sema4a and Nrp1 as the ligand–receptor pair. M.E.T. conducted many of the tumour experiments. D.M.G. performed a substantial portion of the colitis experiments. C.G. performed TIRF microscopy. M.L.B. assisted with the Foxp3-deficiency rescue experiments. A.E.O. assisted with several experiments. P.V. performed histological analysis. D.F. performed computational analysis of the microarray data. J.B. provided the blocking monoclonal antibodies to Sema4a and Nrp1. C.J.W. conducted and curated the initial microarray analysis. D.A.A.V. conceived the project, directed the research and wrote the manuscript. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
J. Bonnevier is an employee of R&D Systems.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-15. (PDF 1424 kb)
Supplementary Table 1
This file contains geneset enrichment analysis for Nrp1-upregulated genesets. (XLS 44 kb)
Supplementary Table 2
This file contains geneset enrichment analysis for Nrp1-downregulated genesets. (XLS 167 kb)
Rights and permissions
About this article
Cite this article
Delgoffe, G., Woo, SR., Turnis, M. et al. Stability and function of regulatory T cells is maintained by a neuropilin-1–semaphorin-4a axis. Nature 501, 252–256 (2013). https://doi.org/10.1038/nature12428
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12428
This article is cited by
-
Maternal dendritic cells influence fetal allograft response following murine in-utero hematopoietic stem cell transplantation
Stem Cell Research & Therapy (2023)
-
LFA-1 knockout inhibited the tumor growth and is correlated with treg cells
Cell Communication and Signaling (2023)
-
IL-2 immunotherapy for targeting regulatory T cells in autoimmunity
Genes & Immunity (2023)
-
Treg plasticity and human diseases
Inflammation Research (2023)
-
Semaphorin 3G exacerbates joint inflammation through the accumulation and proliferation of macrophages in the synovium
Arthritis Research & Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.