Abstract
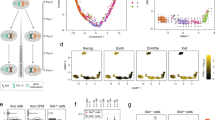
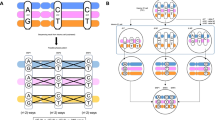
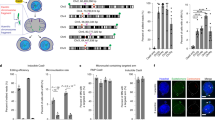
Down’s syndrome is a common disorder with enormous medical and social costs, caused by trisomy for chromosome 21. We tested the concept that gene imbalance across an extra chromosome can be de facto corrected by manipulating a single gene, XIST (the X-inactivation gene). Using genome editing with zinc finger nucleases, we inserted a large, inducible XIST transgene into the DYRK1A locus on chromosome 21, in Down’s syndrome pluripotent stem cells. The XIST non-coding RNA coats chromosome 21 and triggers stable heterochromatin modifications, chromosome-wide transcriptional silencing and DNA methylation to form a ‘chromosome 21 Barr body’. This provides a model to study human chromosome inactivation and creates a system to investigate genomic expression changes and cellular pathologies of trisomy 21, free from genetic and epigenetic noise. Notably, deficits in proliferation and neural rosette formation are rapidly reversed upon silencing one chromosome 21. Successful trisomy silencing in vitro also surmounts the major first step towards potential development of ‘chromosome therapy’.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mégarbané, A. et al. The 50th anniversary of the discovery of trisomy 21: the past, present, and future of research and treatment of Down syndrome. Genet. Med. 11, 611–616 (2009)
Gardiner, K. J. Molecular basis of pharmacotherapies for cognition in Down syndrome. Trends Pharmacol. Sci. 31, 66–73 (2010)
Prandini, P. et al. Natural gene-expression variation in Down syndrome modulates the outcome of gene-dosage imbalance. Am. J. Hum. Genet. 81, 252–263 (2007)
Haydar, T. F. & Reeves, R. H. Trisomy 21 and early brain development. Trends Neurosci. 35, 81–91 (2012)
O’Doherty, A. et al. An aneuploid mouse strain carrying human chromosome 21 with Down syndrome phenotypes. Science 309, 2033–2037 (2005)
Lee, B. & Davidson, B. L. Gene therapy grows into young adulthood: special review issue. Hum. Mol. Genet. 20, R1 (2011)
Hall, L. L. et al. X-inactivation reveals epigenetic anomalies in most hESC but identifies sublines that initiate as expected. J. Cell. Physiol. 216, 445–452 (2008)
Nazor, K. L. et al. Recurrent variations in DNA methylation in human pluripotent stem cells and their differentiated derivatives. Cell Stem Cell 10, 620–634 (2012)
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527–542 (1992)
Clemson, C. M., McNeil, J. A., Willard, H. F. & Lawrence, J. B. XIST RNA paints the inactive X chromosome at interphase: evidence for a novel RNA involved in nuclear/chromosome structure. J. Cell Biol. 132, 259–275 (1996)
Heard, E. Delving into the diversity of facultative heterochromatin: the epigenetics of the inactive X chromosome. Curr. Opin. Genet. Dev. 15, 482–489 (2005)
Hall, L. L. & Lawrence, J. B. XIST RNA and architecture of the inactive X chromosome: implications for the repeat genome. Cold Spring Harb. Symp. Quant. Biol. 75, 345–356 (2010)
Carrel, L. & Willard, H. F. X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434, 400–404 (2005)
Lee, J. T., Strauss, W. M., Dausman, J. A. & Jaenisch, R. A 450 kb transgene displays properties of the mammalian X-inactivation center. Cell 86, 83–94 (1996)
Hall, L. L., Clemson, C. M., Byron, M., Wydner, K. & Lawrence, J. B. Unbalanced X;autosome translocations provide evidence for sequence specificity in the association of XIST RNA with chromatin. Hum. Mol. Genet. 11, 3157–3165 (2002)
Hall, L. L. et al. An ectopic human XIST gene can induce chromosome inactivation in postdifferentiation human HT-1080 cells. Proc. Natl Acad. Sci. USA 99, 8677–8682 (2002)
Moehle, E. A. et al. Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc. Natl Acad. Sci. USA 104, 3055–3060 (2007)
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nature Rev. Genet. 11, 636–646 (2010)
DeKelver, R. C. et al. Functional genomics, proteomics, and regulatory DNA analysis in isogenic settings using zinc finger nuclease-driven transgenesis into a safe harbor locus in the human genome. Genome Res. 20, 1133–1142 (2010)
Park, I. H. et al. Disease-specific induced pluripotent stem cells. Cell 134, 877–886 (2008)
Aït Yahya-Graison, E. et al. Classification of human chromosome 21 gene-expression variations in Down syndrome: impact on disease phenotypes. Am. J. Hum. Genet. 81, 475–491 (2007)
Biancotti, J. C. et al. Human embryonic stem cells as models for aneuploid chromosomal syndromes. Stem Cells 28, 1530–1540 (2010)
Csankovszki, G., Nagy, A. & Jaenisch, R. Synergism of Xist RNA, DNA methylation, and histone hypoacetylation in maintaining X chromosome inactivation. J. Cell Biol. 153, 773–784 (2001)
Cotton, A. M. et al. Chromosome-wide DNA methylation analysis predicts human tissue-specific X inactivation. Hum. Genet. 130, 187–201 (2011)
Guidi, S., Ciani, E., Bonasoni, P., Santini, D. & Bartesaghi, R. Widespread proliferation impairment and hypocellularity in the cerebellum of fetuses with down syndrome. Brain Pathol. 21, 361–373 (2011)
Shi, Y. et al. A human stem cell model of early Alzheimer’s disease pathology in Down syndrome. Sci. Transl. Med. 4, 124ra29 (2012)
Lavon, N. et al. Derivation of euploid human embryonic stem cells from aneuploid embryos. Stem Cells 26, 1874–1882 (2008)
Li, L. B. et al. Trisomy correction in down syndrome induced pluripotent stem cells. Cell Stem Cell 11, 615–619 (2012)
Doyon, J. B. et al. Rapid and efficient clathrin-mediated endocytosis revealed in genome-edited mammalian cells. Nature Cell Biol. 13, 331–337 (2011)
Miller, J. C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nature Biotechnol. 25, 778–785 (2007)
Guschin, D. Y. et al. A rapid and general assay for monitoring endogenous gene modification. Methods Mol. Biol. 649, 247–256 (2010)
Urnov, F. D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005)
Byron, M., Hall, L. L. & Lawrence, J. B. A multifaceted FISH approach to study endogenous RNAs and DNAs in native nuclear and cell structures. Curr. Protoc. Hum. Gen. Chapter 4, Unit 4 15. (2013)
Irizarry, R. A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003)
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nature Genet. 39, 457–466 (2007)
Acknowledgements
We appreciate recent initiatives by administrators of NIGMS and NIH to support more high-risk, high-impact research. Research began with support from GM053234 to J.B.L. for basic X chromosome research, and was made fully possible by GM085548 and GM096400 RC4 to J.B.L. C.J.B. and A.M.C. were supported by CIHR (MOP-13680) to C.J.B. We thank T. Flotte for encouragement and advice regarding genome editing strategies, and similarly appreciate the support of S. Jones and P. Newburger. We thank T. Collingwood for initial discussions regarding this project, and the George Daley laboratory (Harvard) for the Down’s syndrome iPS cell line. L. Lizotte, Z. Matijasevic, K. Smith and E. Swanson provided various assistance. M. S. Kobor and L. Lam (Kobor laboratory) assisted with methylation analysis. D.M.C. is supported by an NIH fellowship 1F32CA154086 and B.R.C. (O. Rando laboratory) is supported by NIH training grant 2T32HD007439 (G. Witman, PI).
Author information
Authors and Affiliations
Contributions
J.J., with the assistance of Y.J., designed and produced all constructs, edited all cell lines, and designed and performed most experiments. J.B.L., J.J. and L.L.H. were the main contributors to designing experiments and interpreting results. J.B.L., J.J., L.L.H. and F.D.U. wrote the manuscript. F.D.U., P.D.G. and G.J.C. engineered and validated ZFNs. J.R.P. performed Cel1 and Southern analysis. J.-C.C. performed SNP analysis, characterized three sub-clones, and helped with proliferation experiments. J.J. and Y.J., with help from J.-C.C., M.B., H.J.K. and L.L.H., carried out initial screening of targeted iPS cell sub-clones. H.J.K. edited and characterized primary Down’s syndrome fibroblast line. A.M.C. and C.J.B. carried out DNA methylation analysis and provided XIST cDNA. J.J. and F.D.U. prepared the microarray library. D.M.C. and B.R.C. analysed microarray data with help from D.A.S., D.Y.G. and E.J.R.
Corresponding authors
Ethics declarations
Competing interests
J.B.L. and L.L.H. are the inventors on an issued patent describing the concept of epigenetic chromosome therapy by targeted addition of non-coding RNA. G.J.C., D.A.S., D.Y.G., J.R.P., E.J.R., P.D.G. and F.D.U. are full-time employees of Sangamo BioSciences.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-9 and Supplementary Tables 1-4. (PDF 5079 kb)
Rights and permissions
About this article
Cite this article
Jiang, J., Jing, Y., Cost, G. et al. Translating dosage compensation to trisomy 21. Nature 500, 296–300 (2013). https://doi.org/10.1038/nature12394
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12394
This article is cited by
-
Understanding the genetic mechanisms and cognitive impairments in Down syndrome: towards a holistic approach
Journal of Neurology (2024)
-
Modeling specific aneuploidies: from karyotype manipulations to biological insights
Chromosome Research (2023)
-
Consequences of gaining an extra chromosome
Chromosome Research (2023)
-
Modeling human neurodevelopmental diseases with brain organoids
Cell Regeneration (2022)
-
APP and DYRK1A regulate axonal and synaptic vesicle protein networks and mediate Alzheimer’s pathology in trisomy 21 neurons
Molecular Psychiatry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.