Abstract
Loss of sexual reproduction is considered an evolutionary dead end for metazoans, but bdelloid rotifers challenge this view as they appear to have persisted asexually for millions of years1. Neither male sex organs nor meiosis have ever been observed in these microscopic animals: oocytes are formed through mitotic divisions, with no reduction of chromosome number and no indication of chromosome pairing2. However, current evidence does not exclude that they may engage in sex on rare, cryptic occasions. Here we report the genome of a bdelloid rotifer, Adineta vaga (Davis, 1873)3, and show that its structure is incompatible with conventional meiosis. At gene scale, the genome of A. vaga is tetraploid and comprises both anciently duplicated segments and less divergent allelic regions. However, in contrast to sexual species, the allelic regions are rearranged and sometimes even found on the same chromosome. Such structure does not allow meiotic pairing; instead, we find abundant evidence of gene conversion, which may limit the accumulation of deleterious mutations in the absence of meiosis. Gene families involved in resistance to oxidation, carbohydrate metabolism and defence against transposons are significantly expanded, which may explain why transposable elements cover only 3% of the assembled sequence. Furthermore, 8% of the genes are likely to be of non-metazoan origin and were probably acquired horizontally. This apparent convergence between bdelloids and prokaryotes sheds new light on the evolutionary significance of sex.
Similar content being viewed by others
Main
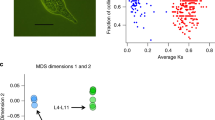
With more than 460 described species4, bdelloid rotifers (Fig. 1) represent the highest metazoan taxonomic rank in which males, hermaphrodites and meiosis are unknown. Such persistence and diversification of an ameiotic clade of animals are in contradiction with the supposed long-term disadvantages of asexuality, making bdelloids an ‘evolutionary scandal’5. Another unusual feature of bdelloid rotifers is their extreme resistance to desiccation at any stage of their life cycle6, enabling these microscopic animals to dwell in ephemeral freshwater habitats such as mosses, lichens and forest litter; this ability is presumably the source of their extreme resistance to ionizing radiation7.
Bdelloid rotifers (‘leech-like wheel-bearers’) are a clade of microscopic animals (scale bar, 100 μm) within the phylum Rotifera. Photographs of Hemichordata (Saccoglossus), Chordata (Homo) and Ecdysozoa (Drosophila) courtesy of David Remsen (MBL), John van Wyhe (http://darwin-online.org.uk) and André Karwath, respectively.
We assembled the genome of a clonal A. vaga lineage into separate haplotypes with a N50 of 260 kilobases (kb) (that is, half of the assembly was composed of fragments longer than 260 kb). Assembly size was 218 megabases (Mb) but 26 Mb of the sequence had twice the average sequencing coverage, suggesting that some nearly identical regions were not resolved during assembly (Supplementary Fig. 3); hence, the total genome size is likely to be 244 Mb, which corresponds to the estimate obtained independently using fluorometry (Supplementary Note C2). Annotation of the complete assembly (including all haplotypes) yielded 49,300 genes. Intragenomic sequence comparisons revealed numerous homologous blocks with conserved gene order (colinear regions). For each such block we computed the per-site synonymous divergence (Ks) and a colinearity metric defined as the fraction of colinear genes. Colinear blocks fell into two groups (Fig. 2a): a group characterized by high colinearity and low average synonymous divergence, and a group characterized by lower colinearity and higher synonymous divergence. The presence of two classes of colinear blocks is consistent with a tetraploid structure comprised of alleles (recent homologues) and ohnologues (ancient homologues formed by genome duplication). Allelic pairs of coding sequences are on average 96.2% identical at the nucleotide level (median = 98.6%) versus 73.6% (median = 75.1%) for ohnologous pairs. Nearly 40% (84.5 Mb) of the assembled genome sequence is organized in quartets of four homologous regions A1, A2, B1 and B2, of which A1–A2 and B1–B2 are two pairs of alleles and As are ohnologous to Bs8 (Fig. 2b).
a, Analysis of intragenomic synteny reveals two groups of colinear regions: alleles (in violet, regions characterized by a high fraction of colinear genes and low average Ks, that is, synonymous divergence) and ohnologues (in orange, with lower colinearity but higher Ks). b, Example of a genomic quartet of four scaffolds: allelic gene pairs are connected with violet curves and ohnologous gene pairs with orange curves.
We found evidence of genomic palindromes up to 705 kb in length and involving up to 148 genes. The A. vaga genome contains at least 17 such palindromic regions (Fig. 3a) reminiscent of those reported in the Y chromosomes of primates9. In all 17 cases, the arms of the palindromes present the colinearity and divergence signatures of allelic regions and do not have other allelic duplicates in the assembly, suggesting that they arose by inter-allelic rearrangements rather than by local duplications. In addition to these 17 inverted repeats, we observed three direct repeats that present the signatures of allelic blocks and involve up to 50 genes (Fig. 3a). The cumulative length of the assembly fragments (scaffolds) bearing these 20 allelic rearrangements is 7.5 Mb or 3.5% of the genome sequence. Allelic regions that are found on the same chromosome clearly cannot segregate during meiosis. Moreover, we found hundreds of colinearity breakpoints between allelic regions, and the total length of the scaffolds that have no full-length homologue in the assembly due to these breakpoints exceeds 109 Mb or 51% of the genome assembly (including 91 of the 100 largest scaffolds, Fig. 3b and Supplementary Fig. 10). As a result, it is impossible to split the assembled genome of A. vaga into haploid sets: the apparent ploidy level of A. vaga is scale-dependent, with a tetraploid structure at gene scale versus chromosome-scale haploidy. Such relaxation of constraints on genome structure is reminiscent of other mitotic lineages such as cancer cells10 and somatic tissues11.
a, In twenty cases, allelic regions are found to occur on the same chromosome. All curves shown connect allelic gene pairs. On three scaffolds both allelic regions have the same orientation (direct repeats, in pink), whereas on the seventeen other scaffolds they are inverted (palindromes, in red). b, Local colinearity between alleles does not extend to chromosome scale. Colours are arbitrary and only allelic gene pairs are represented. Asterisks highlight colinearity breakpoints between scaffold av1 and its allelic partners av44, av94, av122, av316 and av448. Further examples for other scaffolds are shown on Supplementary Fig. 10.
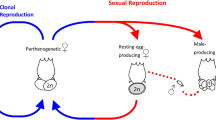
It has been proposed that, in the absence of meiosis, alleles accumulate mutations independently from one another, to the point that ancient asexuals may harbour genome-wide allele sequence divergence (ASD)12 larger than inter-individual differences (the so-called ‘Meselson effect’). However, the average inter-allelic divergence of A. vaga is only 4.4% at the nucleotide level (3% when looking at synonymous divergence), which falls in the upper range reported for sexually reproducing species13. The absence of genome-wide ASD could be explained by low mutation rates and/or by frequent mitotic recombination (such as gene conversion resulting from DNA repair)12. Although there is no evidence of reduced mutation rates in bdelloid rotifers compared with their cyclically sexual sister clade the monogononts14, we found strong signatures of recent gene conversion events in the distribution of identity track lengths, that is, distances between consecutive mismatches (Fig. 4a and Supplementary Note E1). We calculated that the probability that a given base in the genome experiences gene conversion is at least one order of magnitude greater than its probability to mutate (Supplementary Note E1), suggesting that homologous regions in the genome of A. vaga undergo concerted evolution15. Homogenization through gene conversion may either expose new mutations to selection by making them homozygous or remove them as they get overwritten with the other allelic version (Fig. 4b), thereby slowing Muller’s ratchet (that is, the irreversible accumulation of detrimental mutations in asexual populations of finite sizes, Supplementary Note E2 and Supplementary Fig. 11).
a, Evidence for gene conversion between allelic regions. If we suppose that mutations happen at random in a Poisson process of parameter 1/M (where M is the average distance between mutations), then the distance between two consecutive mismatches follows a negative exponential distribution where the proportion of identity tracks of length x equals e−x/M/M. Comparison of the observed distribution of identity track lengths with this theoretical distribution reveals a deficit of short tracks and an excess of long tracks, as expected in case of gene conversion. The same pattern was observed when gene-coding regions were excluded from the analysis (data not shown), thereby ruling out a confounding effect of selection. b, In sexual organisms, meiotic recombination can generate offspring with fewer or more deleterious mutations (hence increasing or decreasing fitness) than the previous generation. The same outcome is expected in ameiotic organisms that experience gene conversion: a deleterious allele may be overwritten by a beneficial or neutral one, resulting in an increase in fitness, or may overwrite it, resulting in decreased fitness.
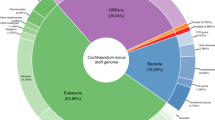
Over 8% of the genes of A. vaga are much more similar to non-metazoan sequences in GenBank than to metazoan ones (AI log score > 45 (ref. 16), Supplementary Note E4) and were therefore probably acquired through horizontal gene transfer (HGT). This class of genes has significantly fewer introns per kilobase of coding sequence compared with probable core metazoan genes (AI ≤ −45, Supplementary Table 2). More than 20% of genes with AI > 45 are found in quartets (groups of four homologous copies in conserved syntenic regions) and were therefore probably incorporated into the rotifer genome before the establishment of tetraploidy, which itself pre-dates the divergence of extant bdelloid families8. The higher the number of copies of a putative HGT gene, the higher its number of introns and the closer its guanine–cytosine (GC) content to the A. vaga genome average (Supplementary Fig. 22), which suggests that these parameters reflect the age of acquisition. We also noticed signatures of possibly very recent HGTs: 60 genes with AI > 45 are present in only one copy (with normal coverage), have no intron and have a GC content that is more than 1% above or below the genome average (the same scaffolds also bear genes of probable metazoan origin with AI < 0). In summary, there seems to be an ancient but still ongoing process of HGT at a level comparable to some bacteria17.
Some theories predict that transposable elements should be either absent from the genomes of asexuals18 or undergo unrestrained expansion after the switch to asexuality, potentially leading to species extinction unless transposable element proliferation is prevented19. We found that transposable elements cover about 3% of the A. vaga genome, which is less than the percentage reported in most other metazoans (including the genome of the obligate parthenogenetic nematode Meloidogyne incognita, 36% of which is made up of repetitive elements20). Another surprising feature is the high diversity of transposable-element families and the extremely low copy numbers observed for each of them (Supplementary Table 3). Out of 255 families, the overwhelming majority (209) are represented by only one or two full-length copies (for 24 families, no full-length copies could be identified), and for each full-length copy there are, on average, only about ten times as many transposable-element fragments. This relatively low abundance of decayed copies and the fact that long-terminal-repeat (LTR) retrotransposons have identical or nearly identical LTRs (Supplementary Table 4) suggest that most low-copy-number families represent recent arrivals. This is consistent with an ongoing process of acquisition of transposable elements by HGT.
This hypothesis is further supported by the significantly higher density of transposable elements observed around HGTs and vice-versa (Supplementary Note E5). If A. vaga has been acquiring transposable elements by HGT, a question that arises is what keeps their number lower than in most other metazoans. Many fragmented copies have apparently been formed through microhomology-mediated deletions. Excision of LTR retrotransposons has also been occurring through LTR–LTR recombination, leaving behind numerous solo LTRs: for example, two Juno1 insertions, Juno1.1 and Juno1.2, which were present as full-length copies in the 2006 A. vaga fosmid library21, exist in the current assembly only as solo LTRs (in the same genomic environments and with the same target site duplications). Finally, there is evidence for expansion and diversification of the RNA-mediated silencing machinery. In addition to Dicer1 proteins, which are shared by all metazoans, A. vaga possesses a deep-branching Dicer-like clade with uncertain taxonomic placement (Supplementary Fig. 20). The Argonaute/Piwi and RNA-directed RNA polymerase (RdRP) families are also expanded (Supplementary Figs 18 and 19). It is plausible that these proteins participate in epigenetic silencing of transposable elements (as was recently observed for single-copy transgenes in Caenorhabditis elegans22), thereby preventing horizontally transferred transposable elements from multiplying upon arrival.
Overall, the genome of A. vaga comprises more genes than usually reported for metazoans (Supplementary Note F2), as its haplotypes were assembled separately. Even taking this into account, the gene repertoire of A. vaga features expansion of several gene families. For example, the genome of A. vaga comprises 284 homeobox superclass genes, mostly found in four copies (quartets) but not organized in clusters; very few ohnologues have been lost, resulting in more homeobox genes than in any other metazoan genome sequenced (Supplementary Note F5). Genes putatively related to oxido-reduction processes are substantially more abundant in A. vaga than in other metazoan species, and most of the corresponding genes appear to be constitutively expressed (Supplementary Table 9). This is consistent with the recent report of an effective antioxidant protection system in bdelloid rotifers23. Carbohydrate-active enzymes (CAZymes) in the genome of A. vaga are also notably diverse and abundant, with 1,075 genes falling into 202 characterized families. With 623 glycoside hydrolases (involved in the hydrolysis of sugar bonds) and 412 glycosyltransferases (responsible for building sugar bonds), the CAZyme richness of A. vaga ranks highest among metazoans and is only comparable to some plants such as poplars24. A. vaga has the richest repertoire of glycoside hydrolases of any organism sequenced so far, hinting at a diversity of feeding habits; 52% of the CAZymes have an AI > 45 and were therefore probably acquired through horizontal gene transfer.
A. vaga has lost 1,250 genes compared with the inferred last common ancestor of Protostomia, the genome of which comprised at least 7,844 unique protein-coding genes (Supplementary Note E6). A total of 137 PFAM domains typically present in metazoans could not be detected in the assembled genome sequence (Supplementary Data 10). Of particular interest are missing domains involved in reproductive processes (Supplementary Note F1); for example, the Zona pellucida-like domain (notably found in sperm-binding proteins25) is present in an average of 36 copies in metazoan genomes but is absent in A. vaga. In contrast, we found multiple copies of most metazoan genes involved in DNA repair and homologous recombination, including a considerably divergent Spo11 but no Rad52 and Msh3.
To conclude, our analysis of a lineage of the bdelloid rotifer Adineta vaga reveals positive evidence for asexual evolution: its genome structure does not allow pairing of homologous chromosomes and therefore seems incompatible with conventional meiosis (Fig. 5). However, we cannot rule out that other forms of recombination occur in bdelloid populations in ways that do not require homologous pairing, such as parasexuality26. The high number of horizontally acquired genes, including some seemingly recent ones, suggests that HGTs may also be occurring from rotifer to rotifer. It is plausible that the repeated cycles of desiccation and rehydration experienced by A. vaga in its natural habitats have had a major role in shaping its genome: desiccation presumably causes DNA double-strand breaks, and these breaks that allow integration of horizontally transferred genetic material also promote gene conversion when they are repaired. Hence, the homogenizing and diversifying roles of sex may have been replaced in bdelloids by gene conversion and horizontal gene transfer, in an unexpected convergence of evolutionary strategy with prokaryotes.
Genes are represented with letters, and dashed lines connect allelic gene pairs. A meiotic genome (left) alternates between a haploid phase (in which a single allele of each gene is present) and a diploid phase (in which the genes are present in two allelic versions arranged colinearly on homologous chromosomes). In the ameiotic genome of A. vaga (right), alleles are distributed in blocks that are shuffled across chromosomes, resulting notably in intrachromosomal repeats (direct or inverted). As a consequence, chromosomes have no homologues and cannot be paired.
Methods Summary
Genomic DNA was extracted from laboratory cultures of a clonal A. vaga lineage and shotgun-sequenced using 454 and Illumina platforms at respective coverage of 25 and 440 times (using both single reads and mate reads from inserts up to 20 kb). The 454 reads were assembled into contigs using MIRA27; the contigs obtained were corrected using single Illumina reads and linked into scaffolds using paired Illumina reads28 (Supplementary Table 1). We annotated protein-coding genes by integrating evidence from RNA sequencing, ab initio predictions and comparison with UniProt. Most synteny and Ka/Ks (non-synonymous divergence/synonymous divergence) analyses were performed using the package MCScanX29 and synteny plots were drawn using Circos30.
Accession codes
Accessions
European Nucleotide Archive
Sequence Read Archive
Data deposits
The sequencing reads and assembly are available at the Sequence Read Archive (accessions ERP002115 and SRP020364 for DNA, ERP002474 and SRP020358 for cDNA) and at the European Nucleotide Archive (accession CAWI1000000000), respectively. The assembly and annotation can be browsed and downloaded at http://www.genoscope.cns.fr/adineta, whereas the result of the orthology analysis is accessible at http://ioda.univ-provence.fr/.
Change history
21 August 2013
The European Nucleotide Archive accession number was corrected.
References
Danchin, E. G. J., Flot, J.-F., Perfus-Barbeoch, L. & Van Doninck, K. In Evolutionary Biology — Concepts, Biodiversity, Macroevolution and Genome Evolution (ed. Pontarotti, P. ) 223–242 (Springer, 2011)
Hsu, W. S. Oogenesis in the Bdelloidea rotifer Philodina roseola Ehrenberg. Cellule 57, 283–296 (1956)
Davis, H. A new Callidina: with the result of experiments on the desiccation of rotifers. Month. Microscopical J. 9, 201–209 (1873)
Segers, H. Annotated checklist of the rotifers (Phylum Rotifera), with notes on nomenclature, taxonomy and distribution. Zootaxa 1564, 1–104 (2007)
Maynard Smith, J. Contemplating life without sex. Nature 324, 300–301 (1986)
Ricci, C. Anhydrobiotic capabilities of bdelloid rotifers. Hydrobiologia 387–388, 321–326 (1998)
Gladyshev, E. & Meselson, M. Extreme resistance of bdelloid rotifers to ionizing radiation. Proc. Natl Acad. Sci. USA 105, 5139–5144 (2008)
Hur, J. H., Van Doninck, K., Mandigo, M. L. & Meselson, M. Degenerate tetraploidy was established before bdelloid rotifer families diverged. Mol. Biol. Evol. 26, 375–383 (2009)
Rozen, S. et al. Abundant gene conversion between arms of palindromes in human and ape Y chromosomes. Nature 423, 873–876 (2003)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Vijg, J. & Dollé, M. E. T. Large genome rearrangements as a primary cause of aging. Mech. Ageing Dev. 123, 907–915 (2002)
Birky, C. W., Jr Heterozygosity, heteromorphy, and phylogenetic trees in asexual eukaryotes. Genetics 144, 427–437 (1996)
Leffler, E. M. et al. Revisiting an old riddle: what determines genetic diversity levels within species? PLoS Biol. 10, e1001388 (2012)
Welch, D. B. M. & Meselson, M. S. Rates of nucleotide substitution in sexual and anciently asexual rotifers. Proc. Natl Acad. Sci. USA 98, 6720–6724 (2001)
Teshima, K. M. & Innan, H. The effect of gene conversion on the divergence between duplicated genes. Genetics 166, 1553–1560 (2004)
Gladyshev, E. A., Meselson, M. & Arkhipova, I. R. Massive horizontal gene transfer in bdelloid rotifers. Science 320, 1210–1213 (2008)
Syvanen, M. Evolutionary implications of horizontal gene transfer. Annu. Rev. Genet. 46, 341–358 (2012)
Hickey, D. A. Selfish DNA: a sexually-transmitted nuclear parasite. Genetics 101, 519–531 (1982)
Arkhipova, I. & Meselson, M. Deleterious transposable elements and the extinction of asexuals. Bioessays 27, 76–85 (2005)
Abad, P. et al. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nature Biotechnol. 26, 909–915 (2008)
Gladyshev, E. A., Meselson, M. & Arkhipova, I. R. A deep-branching clade of retrovirus-like retrotransposons in bdelloid rotifers. Gene 390, 136–145 (2007)
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012)
Krisko, A., Leroy, M., Radman, M. & Meselson, M. Extreme anti-oxidant protection against ionizing radiation in bdelloid rotifers. Proc. Natl Acad. Sci. USA 109, 2354–2357 (2012)
Geisler-Lee, J. et al. Poplar carbohydrate-active enzymes. Gene identification and expression analyses. Plant Physiol. 140, 946–962 (2006)
Bork, P. & Sander, C. A large domain common to sperm receptors (Zp2 and Zp3) and TGF-β type III receptor. FEBS Lett. 300, 237–240 (1992)
Forche, A. et al. The parasexual cycle in Candida albicans provides an alternative pathway to meiosis for the formation of recombinant strains. PLoS Biol. 6, e110 (2008)
Chevreux, B., Wetter, T. & Suhai, S. Genome sequence assembly using trace signals and additional sequence information. Proc. German Conf. Bioinf. 99, 45–56 (1999)
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011)
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012)
Krzywinski, M. et al. Circos: An information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009)
Acknowledgements
The authors would like to thank M. Meselson for his support during the initiation phase of this project and for inspiring us with his seminal works on bdelloid genetics. The authors are also grateful to M. Radman for useful discussions, M. Knapen and N. Debortoli for participating in laboratory work, M. Lliros for helping with Fig. 1, S. Henrissat for participating in CAZyme analyses, and S. Oztas, B. Vacherie, P. Lenoble and S. Mangenot for performing PCR validations of the assembly. This work was supported by Genoscope-CES (where most of the sequencing was performed), by US National Science Foundation grants MCB-0821956 and MCB-1121334 to I.A., by German Research Foundation grant HA 5163/2-1 to O.H., by grant 11.G34.31.0008 from the Ministry of Education and Science of the Russian Federation to A.S.K., by grant NSF CAREER number 0644282 to M.K., by US National Science Foundation grant MCB-0923676 to D.B.M.W., by FRFC grant 2.4.655.09.F from the Belgian Fonds National de la Recherche Scientifique (FNRS) and a start-up grant from the University of Namur to K.V.D.; J.F.F. and K.V.D. thank also J.-P. Descy (University of Namur) for funding support.
Author information
Authors and Affiliations
Contributions
Bo.H., X.L., and B.N. are joint second authors; O.J. and K.V.D. are joint last authors. Bo.H., X.L., F.R. and B.H.L. maintained the rotifer cultures; Bo.H., X.L., F.R. and B.H.L. prepared the genomic DNA; X.L., D.B.M.W. and B.H.L. carried out gene expression experiments; Bo.H., X.L. and B.H.L. prepared complementary DNAs; K.L., J.P. and B.H.L. carried out the sequencing; J.F.F., A.C., V.B., O.J., B.N., J.M.A. and C.D.S. assembled the genome, validated the assembly and built the gene set; J.F.F., J.M.A., V.B., G.A.B., M.D.R., E.G.J.D., O.A.V., M.K., P.W., O.J. and K.V.D. analysed the genome structure; Bo.H., E.G.J.D., M.D.R., J.F.F., A.H., Be.H., B.H.L., R.K., B.L., J.F.R., F.R., A.S.K., E.W., D.B.M.W. and K.V.D. analysed the gene families; I.A., J.B., O.P. and I.Y. annotated and analysed the transposable elements; O.C., P.G., B.W., R.B., P.P. and K.V.D. carried out orthology analysis; I.A., E.G., E.G.J.D., P.G., B.W., F.R., D.B.M.W., P.P., J.F.F. and O.J. analysed the horizontal gene transfers; O.A.V., J.F.F., G.A.B., A.S.K. and D.B.M.W. analysed the signatures of gene conversion; O.H. modelled the effect of gene conversion on Muller’s ratchet; J.F.F., O.J. and K.V.D. wrote the core of the manuscript, with contributions from I.A., E.G.J.D., A.H., B.N., O.H., Be.H., Bo.H., R.K., J.M.A., J.F.R., O.A.V., M.K., A.S.K., D.B.M.W., P.P. and P.W.; and P.W., J.W., R.B., D.B.M.W., P.P., O.J. and K.V.D. designed the project and acquired funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Notes A-F, Supplementary Figures 1-22, Supplementary Tables 1-9, a list of the Supplementary Data files (see separate zipped file), and Supplementary References. For more details see contents list on pages 2-3. (PDF 7692 kb)
Supplementary Data
This zipped file contains Supplementary Data files 1-10 (see Supplementary Information file page 56 for details). (ZIP 5124 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-Non-Commercial-ShareAlike 3.0 Unported licence. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Flot, JF., Hespeels, B., Li, X. et al. Genomic evidence for ameiotic evolution in the bdelloid rotifer Adineta vaga . Nature 500, 453–457 (2013). https://doi.org/10.1038/nature12326
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12326
This article is cited by
-
Ionizing radiation responses appear incidental to desiccation responses in the bdelloid rotifer Adineta vaga
BMC Biology (2024)
-
Back to the roots, desiccation and radiation resistances are ancestral characters in bdelloid rotifers
BMC Biology (2023)
-
Horizontal acquisition of a DNA ligase improves DNA damage tolerance in eukaryotes
Nature Communications (2023)
-
Common origin of sterol biosynthesis points to a feeding strategy shift in Neoproterozoic animals
Nature Communications (2023)
-
Nuclear genome annotation of wheel animals and thorny-headed worms: inferences about the last common ancestor of Syndermata (Rotifera s.l.)
Hydrobiologia (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.