Abstract
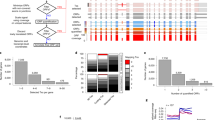
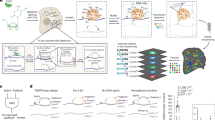
Transcript function is determined by sequence elements arranged on an individual RNA molecule. Variation in transcripts can affect messenger RNA stability, localization and translation1, or produce truncated proteins that differ in localization2 or function3. Given the existence of overlapping, variable transcript isoforms, determining the functional impact of the transcriptome requires identification of full-length transcripts, rather than just the genomic regions that are transcribed4,5. Here, by jointly determining both transcript ends for millions of RNA molecules, we reveal an extensive layer of isoform diversity previously hidden among overlapping RNA molecules. Variation in transcript boundaries seems to be the rule rather than the exception, even within a single population of yeast cells. Over 26 major transcript isoforms per protein-coding gene were expressed in yeast. Hundreds of short coding RNAs and truncated versions of proteins are concomitantly encoded by alternative transcript isoforms, increasing protein diversity. In addition, approximately 70% of genes express alternative isoforms that vary in post-transcriptional regulatory elements, and tandem genes frequently produce overlapping or even bicistronic transcripts. This extensive transcript diversity is generated by a relatively simple eukaryotic genome with limited splicing, and within a genetically homogeneous population of cells. Our findings have implications for genome compaction, evolution and phenotypic diversity between single cells. These data also indicate that isoform diversity as well as RNA abundance should be considered when assessing the functional repertoire of genomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
The data reported in this paper have been deposited in GEO under accession number GSE39128 and are also accessible at http://steinmetzlab.embl.de/TIFSeq.
References
Di Giammartino, D. C., Nishida, K. & Manley, J. L. Mechanisms and consequences of alternative polyadenylation. Mol. Cell 43, 853–866 (2011)
Carlson, M. & Botstein, D. Two differentially regulated mRNAs with different 5′ ends encode secreted with intracellular forms of yeast invertase. Cell 28, 145–154 (1982)
Ungewitter, E. & Scrable, H. Δ40p53 controls the switch from pluripotency to differentiation by regulating IGF signaling in ESCs. Genes Dev. 24, 2408–2419 (2010)
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009)
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008)
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nature Genet. 38, 626–635 (2006)
Ozsolak, F. et al. Comprehensive polyadenylation site maps in yeast and human reveal pervasive alternative polyadenylation. Cell 143, 1018–1029 (2010)
Zhang, Z. & Dietrich, F. S. Mapping of transcription start sites in Saccharomyces cerevisiae using 5′ SAGE. Nucleic Acids Res. 33, 2838–2851 (2005)
Cherry, J. M. et al. Saccharomyces Genome Database: the genomics resource of budding yeast. Nucleic Acids Res. 40, D700–D705 (2012)
Rhee, H. S. & Pugh, B. F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature 483, 295–301 (2012)
Tan-Wong, S. M. et al. Gene loops enhance transcriptional directionality. Science 338, 671–675 (2012)
Riordan, D. P., Herschlag, D. & Brown, P. O. Identification of RNA recognition elements in the Saccharomyces cerevisiae transcriptome. Nucleic Acids Res. 39, 1501–1509 (2011)
Hood, H. M., Neafsey, D. E., Galagan, J. & Sachs, M. S. Evolutionary roles of upstream open reading frames in mediating gene regulation in fungi. Annu. Rev. Microbiol. 63, 385–409 (2009)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Gerashchenko, M. V., Lobanov, A. V. & Gladyshev, V. N. Genome-wide ribosome profiling reveals complex translational regulation in response to oxidative stress. Proc. Natl Acad. Sci. USA 109, 17394–17399 (2012)
Kondo, T. et al. Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329, 336–339 (2010)
Chatton, B., Walter, P., Ebel, J. P., Lacroute, F. & Fasiolo, F. The yeast VAS1 gene encodes both mitochondrial and cytoplasmic valyl-tRNA synthetases. J. Biol. Chem. 263, 52–57 (1988)
Fournier, C. T. et al. Amino termini of many yeast proteins map to downstream start codons. J. Proteome Res. 11, 5712–5719 (2012)
Yao, P. et al. Coding region polyadenylation generates a truncated tRNA synthetase that counters translation repression. Cell 149, 88–100 (2012)
Majumdar, S., Ghatak, J., Mukherji, S., Bhattacharjee, H. & Bhaduri, A. UDPgalactose 4-epimerase from Saccharomyces cerevisiae. A bifunctional enzyme with aldose 1-epimerase activity. Eur. J. Biochem. 271, 753–759 (2004)
Mayer, A. et al. CTD tyrosine phosphorylation impairs termination factor recruitment to RNA polymerase II. Science 336, 1723–1725 (2012)
Xu, Z. et al. Antisense expression increases gene expression variability and locus interdependency. Mol. Syst. Biol. 7, 468 (2011)
Miura, F. et al. Absolute quantification of the budding yeast transcriptome by means of competitive PCR between genomic and complementary DNAs. BMC Genomics 9, 574 (2008)
Ruan, X. & Ruan, Y. Genome wide full-length transcript analysis using 5′ and 3′ paired-end-tag next generation sequencing (RNA-PET). Methods Mol. Biol. 809, 535–562 (2012)
Wilkening, S. et al. An efficient method for genome-wide polyadenylation site mapping and RNA quantification. Nucleic Acids Res. 41, e65 (2013)
Venters, B. J. & Pugh, B. F. A canonical promoter organization of the transcription machinery and its regulators in the Saccharomyces genome. Genome Res. 19, 360–371 (2009)
Kaplan, N. et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458, 362–366 (2009)
Acknowledgements
We thank R. Aiyar for help in editing and refining the manuscript. We thank W. Huber, C. Zhu, A. I. Järvelin, S. Clauder-Münster, J. Zaugg, S. Adjalley, G. Lin and the members of the Steinmetz laboratory for helpful discussions and critical comments on the manuscript. We thank V. N. Gladyshev and C. Pineau for sharing published data. This study was technically supported by the EMBL Genomics Core Facility. This study was financially supported by the National Institutes of Health (to L.M.S.). V.P. was supported by an EMBO fellowship.
Author information
Authors and Affiliations
Contributions
W.W., V.P. and L.M.S. conceived the project. V.P. developed the TIF-Seq method and performed experiments. W.W. and V.P. performed the analysis. V.P., W.W. and L.M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-24, Supplementary Methods, a Supplementary Discussion, Supplementary Tables 1-5, legends for the Supplementary Data (see separate file) and Supplementary References. (PDF 2848 kb)
Supplementary Data
This zipped file contains Supplementary Data files 1-10 – see Supplementary Information for full legends. Supplementary Data 10 was updated on 4 July 2013 to correct a misannotation. (ZIP 15418 kb)
Rights and permissions
About this article
Cite this article
Pelechano, V., Wei, W. & Steinmetz, L. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature 497, 127–131 (2013). https://doi.org/10.1038/nature12121
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12121
This article is cited by
-
Characterization and optimization of 5´ untranslated region containing poly-adenine tracts in Kluyveromyces marxianus using machine-learning model
Microbial Cell Factories (2024)
-
Quantitative analysis of transcription start site selection reveals control by DNA sequence, RNA polymerase II activity and NTP levels
Nature Structural & Molecular Biology (2024)
-
Monitoring the 5′UTR landscape reveals isoform switches to drive translational efficiencies in cancer
Oncogene (2023)
-
Ceg1 depletion reveals mechanisms governing degradation of non-capped RNAs in Saccharomyces cerevisiae
Communications Biology (2023)
-
New insights into genome annotation in Podospora anserina through re-exploiting multiple RNA-seq data
BMC Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.