Abstract
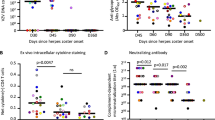
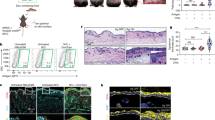
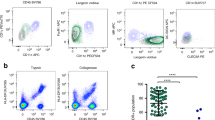
Most herpes simplex virus 2 (HSV-2) reactivations in humans are subclinical and associated with rapid expansion and containment of virus. Previous studies have shown that CD8+ T cells persist in genital skin and mucosa at the dermal–epidermal junction (DEJ)—the portal of neuronal release of reactivating virus—for prolonged time periods after herpes lesions are cleared1,2. The phenotype and function of this persistent CD8+ T-cell population remain unknown. Here, using cell-type-specific laser capture microdissection, transcriptional profiling and T-cell antigen receptor β-chain (TCRβ) genotyping on sequential genital skin biopsies, we show that CD8αα+ T cells are the dominant resident population of DEJ CD8+ T cells that persist at the site of previous HSV-2 reactivation. CD8αα+ T cells located at the DEJ lack chemokine-receptor expression required for lymphocyte egress and recirculation, express gene signatures of T-cell activation and antiviral activity, and produce cytolytic granules during clinical and virological quiescent time periods. Sequencing of the TCR β-chain repertoire reveals that the DEJ CD8αα+ T cells are oligoclonal with diverse usage of TCR variable-β genes, which differ from those commonly described for mucosa-associated invariant T cells and natural killer T cells. Dominant clonotypes are shown to overlap among multiple recurrences over a period of two-and-a-half years. Episodes of rapid asymptomatic HSV-2 containment were also associated with a high CD8 effector-to-target ratio and focal enrichment of CD8αα+ T cells. These studies indicate that DEJ CD8αα+ T cells are tissue-resident cells that seem to have a fundamental role in immune surveillance and in initial containment of HSV-2 reactivation in human peripheral tissue. Elicitation of CD8αα+ T cells may be a critical component for developing effective vaccines against skin and mucosal infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
12 June 2013
Nature 497, 494–497 (2013); doi:10.1038/nature12110 In Fig. 2a of our Letter, the label ‘CXCR7’ should be ‘CXCR6’, and the label ‘CXCR8’ should be ‘CXCR7’. This error has been corrected in the HTML and PDF versions of the paper.
References
Zhu, J. et al. Virus-specific CD8+ T cells accumulate near sensory nerve endings in genital skin during subclinical HSV-2 reactivation. J. Exp. Med. 204, 595–603 (2007)
Zhu, J. et al. Persistence of HIV-1 receptor-positive cells after HSV-2 reactivation is a potential mechanism for increased HIV-1 acquisition. Nature Med. 15, 886–892 (2009)
Wald, A., Zeh, J., Selke, S., Ashley, R. L. & Corey, L. Virologic characteristics of subclinical and symptomatic genital herpes infections. N. Engl. J. Med. 333, 770–775 (1995)
Wald, A. et al. Reactivation of genital herpes simplex virus type 2 infection in asymptomatic seropositive persons. N. Engl. J. Med. 342, 844–850 (2000)
Mark, K. E. et al. Rapidly cleared episodes of herpes simplex virus reactivation in immunocompetent adults. J. Infect. Dis. 198, 1141–1149 (2008)
Hayday, A., Theodoridis, E., Ramsburg, E. & Shires, J. Intraepithelial lymphocytes: exploring the Third Way in immunology. Nature Immunol. 2, 997–1003 (2001)
Huang, Y. et al. Mucosal memory CD8+ T cells are selected in the periphery by an MHC class I molecule. Nature Immunol. 12, 1086–1095 (2011)
Matloubian, M. et al. Lymphocyte egress from thymus and peripheral lymphoid organs is dependent on S1P receptor 1. Nature 427, 355–360 (2004)
Bromley, S. K., Thomas, S. Y. & Luster, A. D. Chemokine receptor CCR7 guides T cell exit from peripheral tissues and entry into afferent lymphatics. Nature Immunol. 6, 895–901 (2005)
Debes, G. F. et al. Chemokine receptor CCR7 required for T lymphocyte exit from peripheral tissues. Nature Immunol. 6, 889–894 (2005)
Chi, H. & Flavell, R. A. Cutting edge: regulation of T cell trafficking and primary immune responses by sphingosine 1-phosphate receptor 1. J. Immunol. 174, 2485–2488 (2005)
Peng, T. et al. An effector phenotype of CD8+ T cells at the junction epithelium during clinical quiescence of herpes simplex virus 2 infection. J. Virol. 86, 10587–10596 (2012)
Schiffer, J. T. et al. Mucosal host immune response predicts the severity and duration of herpes simplex virus-2 genital tract shedding episodes. Proc. Natl Acad. Sci. USA 107, 18973–18978 (2010)
Schiffer, J. T. et al. Frequent release of low amounts of herpes simplex virus from neurons: results of a mathematical model. Sci. Transl Med. 1, 7ra16 (2009)
Robins, H. S. et al. Comprehensive assessment of T-cell receptor β-chain diversity in αβ T cells. Blood 114, 4099–4107 (2009)
Tilloy, F. et al. An invariant T cell receptor α chain defines a novel TAP-independent major histocompatibility complex class Ib-restricted αβ T cell subpopulation in mammals. J. Exp. Med. 189, 1907–1921 (1999)
Le Bourhis, L. et al. Mucosal-associated invariant T cells: unconventional development and function. Trends Immunol. 32, 212–218 (2011)
Jing, L. et al. Cross-presentation and genome-wide screening reveal candidate T cells antigens for a herpes simplex virus type 1 vaccine. J. Clin. Invest. 122, 654–673 (2012)
Clark, R. A. et al. Skin effector memory T cells do not recirculate and provide immune protection in alemtuzumab-treated CTCL patients. Sci. Transl. Med. 4, 117ra7 (2012)
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nature Immunol. 10, 524–530 (2009)
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011)
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012)
Khanna, K. M., Bonneau, R. H., Kinchington, P. R. & Hendricks, R. L. Herpes simplex virus-specific memory CD8+ T cells are selectively activated and retained in latently infected sensory ganglia. Immunity 18, 593–603 (2003)
Purwar, R. et al. Resident memory T cells (TRM) are abundant in human lung: diversity, function, and antigen specificity. PLoS ONE 6, e16245 (2011)
Wakim, L. M., Woodward-Davis, A. & Bevan, M. J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl Acad. Sci. USA 107, 17872–17879 (2010)
Trautmann, L. et al. Selection of T cell clones expressing high-affinity public TCRs within human cytomegalovirus-specific CD8 T cell responses. J. Immunol. 175, 6123–6132 (2005)
Cunningham, A. L., Turner, R. R., Miller, A. C., Para, M. F. & Merigan, T. C. Evolution of recurrent herpes simplex lesions. An immunohistologic study. J. Clin. Invest. 75, 226–233 (1985)
Nakanishi, Y., Lu, B., Gerard, C. & Iwasaki, A. CD8+ T lymphocyte mobilization to virus-infected tissue requires CD4+ T-cell help. Nature 462, 510–513 (2009)
Magaret, A. S., Wald, A., Huang, M. L., Selke, S. & Corey, L. Optimizing PCR positivity criterion for detection of herpes simplex virus DNA on skin and mucosa. J. Clin. Microbiol. 45, 1618–1620 (2007)
Riddell, S. R. et al. Restoration of viral immunity in immunodeficient humans by the adoptive transfer of T cell clones. Science 257, 238–241 (1992)
Acknowledgements
We thank M. Huang, H. Xie, J. Vazquez and D. McDonald for technical assistance; M. Prlic and C. Desmarais for discussion and reading the manuscript and M. Miner for editing. We also thank our study participants and S. Kuntz and M. Stern for clinical assistance. This work was supported by grants from the National Institutes of Health (R37AI042528, R01AI04252815, P01AI030731 and R56AI093746) and the James B. Pendleton Charitable Trust.
Author information
Authors and Affiliations
Contributions
J.Z. and L.C. conceived the study and wrote the manuscript. J.Z. and T.P. developed the technology and analysed and interpreted the data. K.P., A.S.K., A.K., L.J. and K.D. performed the experiments. D.M.K. isolated HSV-2 reactive CD8+ T cells and peptide epitope used in the study. C.J. and A.W. directed human biopsy studies. H.R. contributed to TCR data analysis and interpretation. All authors contributed to the discussion.
Corresponding authors
Ethics declarations
Competing interests
L.C. is on the scientific advisory board for, and hold stock (<1% of company) in, Immune Design Corp. H.R. is a co-founder of Adaptive Biotechnologies, owns stock and intellectual property of this company, and consults for the company. A.W. has received research support from NIH, Genocea Biosciences, Agenus, and Gilead Sciences, and has been a consultant to Aicuris GmbH. D.M.K. is a consultant for Agenus and EISAI. C.J. is conducting research with AiCuris GmbH, which is developing treatment for HSV.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5 and Supplementary Tables 1-5. (PDF 20560 kb)
Rights and permissions
About this article
Cite this article
Zhu, J., Peng, T., Johnston, C. et al. Immune surveillance by CD8αα+ skin-resident T cells in human herpes virus infection. Nature 497, 494–497 (2013). https://doi.org/10.1038/nature12110
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12110
This article is cited by
-
Modeling human HSV infection via a vascularized immune-competent skin-on-chip platform
Nature Communications (2022)
-
Tissue-resident memory T cells in the urogenital tract
Nature Reviews Nephrology (2022)
-
The human memory T cell compartment changes across tissues of the female reproductive tract
Mucosal Immunology (2021)
-
Tissue-resident memory Th17 cells maintain stable fungal commensalism in the oral mucosa
Mucosal Immunology (2021)
-
Immovable memories: the journey to permanent residency
Nature Immunology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.