Abstract
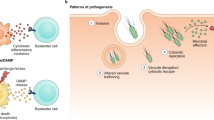
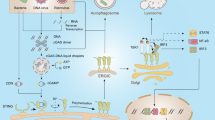
Our innate immune system distinguishes microbes from self by detecting conserved pathogen-associated molecular patterns1. However, these are produced by all microbes, regardless of their pathogenic potential. To distinguish virulent microbes from those with lower disease-causing potential the innate immune system detects conserved pathogen-induced processes2, such as the presence of microbial products in the host cytosol, by mechanisms that are not fully resolved. Here we show that NOD1 senses cytosolic microbial products by monitoring the activation state of small Rho GTPases. Activation of RAC1 and CDC42 by bacterial delivery or ectopic expression of SopE, a virulence factor of the enteric pathogen Salmonella, triggered the NOD1 signalling pathway, with consequent RIP2 (also known as RIPK2)-mediated induction of NF-κB-dependent inflammatory responses. Similarly, activation of the NOD1 signalling pathway by peptidoglycan required RAC1 activity. Furthermore, constitutively active forms of RAC1, CDC42 and RHOA activated the NOD1 signalling pathway. Our data identify the activation of small Rho GTPases as a pathogen-induced process sensed through the NOD1 signalling pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Janeway, C. A., Jr Approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb. Symp. Quant. Biol. 54, 1–13 (1989)
Vance, R. E., Isberg, R. R. & Portnoy, D. A. Patterns of pathogenesis: discrimination of pathogenic and nonpathogenic microbes by the innate immune system. Cell Host Microbe 6, 10–21 (2009)
Fu, Y. & Galan, J. E. The Salmonella typhimurium tyrosine phosphatase SptP is translocated into host cells and disrupts the actin cytoskeleton. Mol. Microbiol. 27, 359–368 (1998)
Zhou, D., Mooseker, M. S. & Galan, J. E. An invasion-associated Salmonella protein modulates the actin-bundling activity of plastin. Proc. Natl Acad. Sci. USA 96, 10176–10181 (1999)
Hardt, W. D., Chen, L. M., Schuebel, K. E., Bustelo, X. R. & Galan, J. E. S. typhimurium encodes an activator of Rho GTPases that induces membrane ruffling and nuclear responses in host cells. Cell 93, 815–826 (1998)
Hapfelmeier, S. et al. Role of the Salmonella pathogenicity island 1 effector proteins SipA, SopB, SopE, and SopE2 in Salmonella enterica subspecies 1 serovar Typhimurium colitis in streptomycin-pretreated mice. Infect. Immun. 72, 795–809 (2004)
Bruno, V. M. et al. Salmonella Typhimurium type III secretion effectors stimulate innate immune responses in cultured epithelial cells. PLoS Pathog. 5, e1000538 (2009)
Figueiredo, J. F. et al. Salmonella enterica Typhimurium SipA induces CXC-chemokine expression through p38MAPK and JUN pathways. Microbes Infect. 11, 302–310 (2009)
Hobbie, S., Chen, L. M., Davis, R. J. & Galan, J. E. Involvement of mitogen-activated protein kinase pathways in the nuclear responses and cytokine production induced by Salmonella typhimurium in cultured intestinal epithelial cells. J. Immunol. 159, 5550–5559 (1997)
Smith, M. F., Jr et al. Toll-like receptor (TLR) 2 and TLR5, but not TLR4, are required for Helicobacter pylori-induced NF-κB activation and chemokine expression by epithelial cells. J. Biol. Chem. 278, 32552–32560 (2003)
Lu, W. et al. Cutting edge: enhanced pulmonary clearance of Pseudomonas aeruginosa by Muc1 knockout mice. J. Immunol. 176, 3890–3894 (2006)
Keestra, A. M. et al. A Salmonella virulence factor activates the NOD1/NOD2 signaling pathway. mBio 2, e00266–00211 (2011)
Coso, O. A. et al. The small GTP-binding proteins Rac1 and Cdc42 regulate the activity of the JNK/SAPK signaling pathway. Cell 81, 1137–1146 (1995)
Schlumberger, M. C. et al. Amino acids of the bacterial toxin SopE involved in G nucleotide exchange on Cdc42. J. Biol. Chem. 278, 27149–27159 (2003)
Müller, A. J., Hoffmann, C. & Hardt, W. D. Caspase-1 activation via Rho GTPases: a common theme in mucosal infections? PLoS Pathog. 6, e1000795 (2010)
Hapfelmeier, S. et al. The Salmonella pathogenicity island (SPI)-2 and SPI-1 type III secretion systems allow Salmonella serovar typhimurium to trigger colitis via MyD88-dependent and MyD88-independent mechanisms. J. Immunol. 174, 1675–1685 (2005)
Le Bourhis, L. et al. Role of Nod1 in mucosal dendritic cells during Salmonella pathogenicity island 1-independent Salmonella enterica serovar Typhimurium infection. Infect. Immun. 77, 4480–4486 (2009)
Geddes, K. et al. Nod1 and Nod2 regulation of inflammation in the Salmonella colitis model. Infect. Immun. 78, 5107–5115 (2010)
Inohara, N. et al. An induced proximity model for NF-κB activation in the Nod1/RICK and RIP signaling pathways. J. Biol. Chem. 275, 27823–27831 (2000)
Girardin, S. E. et al. CARD4/Nod1 mediates NF-κB and JNK activation by invasive Shigella flexneri. EMBO Rep. 2, 736–742 (2001)
Argast, G. M., Fausto, N. & Campbell, J. S. Inhibition of RIP2/RIck/CARDIAK activity by pyridinyl imidazole inhibitors of p38 MAPK. Mol. Cell. Biochem. 268, 129–140 (2005)
Barthel, M. et al. Pretreatment of mice with streptomycin provides a Salmonella enterica serovar Typhimurium colitis model that allows analysis of both pathogen and host. Infect. Immun. 71, 2839–2858 (2003)
Wong, K. W., Mohammadi, S. & Isberg, R. R. The polybasic region of Rac1 modulates bacterial uptake independently of self-association and membrane targeting. J. Biol. Chem. 283, 35954–35965 (2008)
Aktories, K., Schmidt, G. & Just, I. Rho GTPases as targets of bacterial protein toxins. Biol. Chem. 381, 421–426 (2000)
Lopez, C. A. et al. Phage-mediated acquisition of a type III secreted effector protein boosts growth of Salmonella by nitrate respiration. mBio 3, e00143–12 (2012)
Boyer, L. et al. Pathogen-derived effectors trigger protective immunity via activation of the Rac2 enzyme and the IMD or Rip kinase signaling pathway. Immunity 35, 536–549 (2011)
Meylan, E. & Tschopp, J. The RIP kinases: crucial integrators of cellular stress. Trends Biochem. Sci. 30, 151–159 (2005)
Tattoli, I., Travassos, L. H., Carneiro, L. A., Magalhaes, J. G. & Girardin, S. E. The Nodosome: Nod1 and Nod2 control bacterial infections and inflammation. Semin. Immunopathol. 29, 289–301 (2007)
Fukazawa, A. et al. GEF-H1 mediated control of NOD1 dependent NF-κB activation by Shigella effectors. PLoS Pathogens 4, e1000228 (2008)
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: a Laboratory Manual (Cold Spring Harb. Lab. Press, 1989)
Schmieger, H. Phage P22-mutants with increased or decreased transduction abilities. Mol. Gen. Genet. 119, 75–88 (1972)
Lawes, M. & Maloy, S. MudSacI, a transposon with strong selectable and counterselectable markers: use for rapid mapping of chromosomal mutations in Salmonella typhimurium. J. Bacteriol. 177, 1383–1387 (1995)
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002)
Acknowledgements
We would like to thank S.-P. Nuccio for providing PCR primers for the construction of the bacterial strains. This work was supported by Public Health Service Grants AI044170 and AI076246. A.M.K. is supported by the American Heart Association Grant 12SDG12220022.
Author information
Authors and Affiliations
Contributions
A.M.K. contributed to the experimental design, performed experiments and contributed to Figs 1b–f, 2a–e, g, 3a–c, f, g and Supplementary Figs 1, 2d, 3a, b, d, 4a–c, 7 and 8. M.G.W. performed experiments, constructed bacterial strains and contributed to Figs 1a, e, f, 3d, g and Supplementary Figs 2a, b, e, h, g, e, 4c, 5b and 8. J.J.A. constructed expression plasmids and contributed to Figs 2f and 3c and Supplementary Fig. 7. S.P.F. constructed expression plasmids and contributed to Fig. 2g and Supplementary Figs 2f, 3c and 6a, b. M.N.X. contributed to Fig. 1e, f and Supplementary Fig. 5a. S.E.W. constructed bacterial strains, contributed to Supplementary Fig. 5b and critically read the manuscript. A.K. constructed expression plasmids and contributed to Supplementary Figs 2c and 7. V.P. constructed bacterial strains. M.M.R. contributed to Supplementary Fig. 2d. J.F.T.W. constructed expression plasmids. R.A.E. performed mass spectrometry. A.M.K., A.J.B. and R.M.T. provided financial support for the study and contributed to the experimental design. A.M.K. and A.J.B. were responsible for the overall study design and for writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3, Supplementary References, and Supplementary Figures 1-8. (PDF 1254 kb)
Rights and permissions
About this article
Cite this article
Keestra, A., Winter, M., Auburger, J. et al. Manipulation of small Rho GTPases is a pathogen-induced process detected by NOD1. Nature 496, 233–237 (2013). https://doi.org/10.1038/nature12025
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12025
This article is cited by
-
A two-step activation mechanism enables mast cells to differentiate their response between extracellular and invasive enterobacterial infection
Nature Communications (2024)
-
Nod1-dependent NF-kB activation initiates hematopoietic stem cell specification in response to small Rho GTPases
Nature Communications (2023)
-
Role of Rho GTPases in inflammatory bowel disease
Cell Death Discovery (2023)
-
Optineurin links Hace1-dependent Rac ubiquitylation to integrin-mediated mechanotransduction to control bacterial invasion and cell division
Nature Communications (2022)
-
Organellar homeostasis and innate immune sensing
Nature Reviews Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.