Abstract
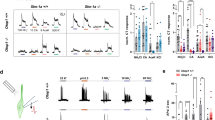
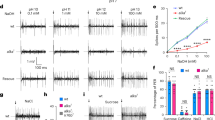
In the tongue, distinct classes of taste receptor cells detect the five basic tastes; sweet, sour, bitter, sodium salt and umami1,2. Among these qualities, bitter and sour stimuli are innately aversive, whereas sweet and umami are appetitive and generally attractive to animals. By contrast, salty taste is unique in that increasing salt concentration fundamentally transforms an innately appetitive stimulus into a powerfully aversive one3,4,5,6,7. This appetitive–aversive balance helps to maintain appropriate salt consumption3,4,6,8, and represents an important part of fluid and electrolyte homeostasis. We have shown previously that the appetitive responses to NaCl are mediated by taste receptor cells expressing the epithelial sodium channel, ENaC8, but the cellular substrate for salt aversion was unknown. Here we examine the cellular and molecular basis for the rejection of high concentrations of salts. We show that high salt recruits the two primary aversive taste pathways by activating the sour- and bitter-taste-sensing cells. We also demonstrate that genetic silencing of these pathways abolishes behavioural aversion to concentrated salt, without impairing salt attraction. Notably, mice devoid of salt-aversion pathways show unimpeded, continuous attraction even to very high concentrations of NaCl. We propose that the ‘co-opting’ of sour and bitter neural pathways evolved as a means to ensure that high levels of salt reliably trigger robust behavioural rejection, thus preventing its potentially detrimental effects on health.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chandrashekar, J., Hoon, M. A., Ryba, N. J. P. & Zuker, C. S. The receptors and cells for mammalian taste. Nature 444, 288–294 (2006)
Yarmolinsky, D. A., Zuker, C. S. & Ryba, N. J. P. Common sense about taste: from mammals to insects. Cell 139, 234–244 (2009)
Beauchamp, G. K., Bertino, M., Burke, D. & Engelman, K. Experimental sodium depletion and salt taste in normal human volunteers. Am. J. Clin. Nutr. 51, 881–889 (1990)
Duncan, C. J. Salt preferences of birds and mammals. Physiologic. Zool. 35, 120–132 (1962)
Eylam, S. & Spector, A. C. Taste discrimination between NaCl and KCl is disrupted by amiloride in inbred mice with amiloride-insensitive chorda tympani nerves. Am. J. Physiol. Regul. Integr. Comp. Physiol. 288, R1361–R1368 (2005)
Contreras, R. J. in Neural Mechanisms in Taste (ed. Cagan, R. H. ) 119–145 (CRC, 1989)
Lindemann, B. Receptors and transduction in taste. Nature 413, 219–225 (2001)
Chandrashekar, J. et al. The cells and peripheral representation of sodium taste in mice. Nature 464, 297–301 (2010)
Doolin, R. E. & Gilbertson, T. A. Distribution and characterization of functional amiloride-sensitive sodium channels in rat tongue. J. Gen. Physiol. 107, 545–554 (1996)
Halpern, B. P. Amiloride and vertebrate gustatory responses to NaCl. Neurosci. Biobehav. Rev. 23, 5–47 (1998)
Heck, G. L., Mierson, S. & DeSimone, J. A. Salt taste transduction occurs through an amiloride-sensitive sodium transport pathway. Science 223, 403–405 (1984)
Hettinger, T. P. & Frank, M. E. Specificity of amiloride inhibition of hamster taste responses. Brain Res. 513, 24–34 (1990)
Spector, A. C., Guagliardo, N. A. & St John, S. J. Amiloride disrupts NaCl versus KCl discrimination performance: implications for salt taste coding in rats. J. Neurosci. 16, 8115–8122 (1996)
Mueller, K. L. et al. The receptors and coding logic for bitter taste. Nature 434, 225–229 (2005)
Zhang, Y. et al. Coding of sweet, bitter, and umami tastes: different receptor cells sharing similar signaling pathways. Cell 112, 293–301 (2003)
Huang, A. L. et al. The cells and logic for mammalian sour taste detection. Nature 442, 934–938 (2006)
Chandrashekar, J. et al. The taste of carbonation. Science 326, 443–445 (2009)
Yamamoto, M. et al. Reversible suppression of glutamatergic neurotransmission of cerebellar granule cells in vivo by genetically manipulated expression of tetanus neurotoxin light chain. J. Neurosci. 23, 6759–6767 (2003)
Gao, Z. G., Jiang, Q., Jacobson, K. A. & Ijzerman, A. P. Site-directed mutagenesis studies of human A2A adenosine receptors: involvement of glu13 and his278 in ligand binding and sodium modulation. Biochem. Pharmacol. 60, 661–668 (2000)
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012)
Zhu, X. L. & Sly, W. S. Carbonic anhydrase IV from human lung. Purification, characterization, and comparison with membrane carbonic anhydrase from human kidney. J. Biol. Chem. 265, 8795–8801 (1990)
Baird, T. T., Jr, Waheed, A., Okuyama, T., Sly, W. S. & Fierke, C. A. Catalysis and inhibition of human carbonic anhydrase IV. Biochemistry 36, 2669–2678 (1997)
Hellekant, G., Ninomiya, Y. & Danilova, V. Taste in chimpanzees II: single chorda tympani fibers. Physiol. Behav. 61, 829–841 (1997)
Breslin, P. A. & Beauchamp, G. K. Suppression of bitterness by sodium: variation among bitter taste stimuli. Chem. Senses 20, 609–623 (1995)
Hallock, R. M., Tatangelo, M., Barrows, J. & Finger, T. E. Residual chemosensory capabilities in double P2X2/P2X3 purinergic receptor null mice: intraoral or postingestive detection? Chem. Senses 34, 799–808 (2009)
Ohkuri, T., Horio, N., Stratford, J. M., Finger, T. E. & Ninomiya, Y. Residual chemoresponsiveness to acids in the superior laryngeal nerve in “taste-blind” (P2X2/P2X3 double-KO) mice. Chem. Senses 37, 523–532 (2012)
Ugawa, S., Ueda, T., Yamamura, H., Nagao, M. & Shimada, S. Coexpression of vanilloid receptor subtype-1 and acid-sensing ion channel genes in the human trigeminal ganglion neurons. Chem. Senses 30 (suppl. 1). i195 (2005)
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998)
Nelson, G. et al. An amino-acid taste receptor. Nature 416, 199–202 (2002)
Oka, Y. et al. Odorant receptor map in the mouse olfactory bulb: in vivo sensitivity and specificity of receptor-defined glomeruli. Neuron 52, 857–869 (2006)
Acknowledgements
We thank E. Vitalis and N. Propp for the generation and maintenance of mouse lines, and K. Mueller for the construction of T2R-Sapphire lines. We also thank J. Chandrashekar, W. Sly and A. Waheed for advice and discussions, and K. Scott and members of our laboratories for comments. Y.O. was supported by the Japan Society for Promotion of Science. This research was supported in part by the intramural research program of the National Institutes of Health and National Institute of Dental and Craniofacial Research (to N.J.P.R.). C.S.Z. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.O. designed the study, carried out electrophysiological, biochemical, pharmacological and behavioural experiments, analysed data and wrote the paper; M.B. carried out nerve recordings and behavioural studies; L.v.B. carried out nerve recordings and localization studies. N.J.P.R. and C.S.Z. designed the study, analysed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
C.S.Z. is a scientific founder and scientific advisory board member of Senomyx.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-9 and Supplementary References. (PDF 836 kb)
Rights and permissions
About this article
Cite this article
Oka, Y., Butnaru, M., von Buchholtz, L. et al. High salt recruits aversive taste pathways. Nature 494, 472–475 (2013). https://doi.org/10.1038/nature11905
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11905
This article is cited by
-
The proton channel OTOP1 is a sensor for the taste of ammonium chloride
Nature Communications (2023)
-
Contribution of TRPC3-mediated Ca2+ entry to taste transduction
Pflügers Archiv - European Journal of Physiology (2023)
-
Taste–taste associations in chemotherapy-induced subjective taste alterations: findings from a questionnaire survey in an outpatient clinic
Supportive Care in Cancer (2023)
-
Molecular logic of salt taste reception in special reference to transmembrane channel-like 4 (TMC4)
The Journal of Physiological Sciences (2022)
-
The impact of excessive salt intake on human health
Nature Reviews Nephrology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.