Abstract
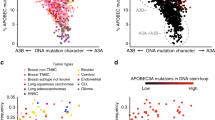
Several mutations are required for cancer development, and genome sequencing has revealed that many cancers, including breast cancer, have somatic mutation spectra dominated by C-to-T transitions1,2,3,4,5,6,7,8,9. Most of these mutations occur at hydrolytically disfavoured10 non-methylated cytosines throughout the genome, and are sometimes clustered8. Here we show that the DNA cytosine deaminase APOBEC3B is a probable source of these mutations. APOBEC3B messenger RNA is upregulated in most primary breast tumours and breast cancer cell lines. Tumours that express high levels of APOBEC3B have twice as many mutations as those that express low levels and are more likely to have mutations in TP53. Endogenous APOBEC3B protein is predominantly nuclear and the only detectable source of DNA C-to-U editing activity in breast cancer cell-line extracts. Knockdown experiments show that endogenous APOBEC3B correlates with increased levels of genomic uracil, increased mutation frequencies, and C-to-T transitions. Furthermore, induced APOBEC3B overexpression causes cell cycle deviations, cell death, DNA fragmentation, γ-H2AX accumulation and C-to-T mutations. Our data suggest a model in which APOBEC3B-catalysed deamination provides a chronic source of DNA damage in breast cancers that could select TP53 inactivation and explain how some tumours evolve rapidly and manifest heterogeneity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007)
Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, 228–231 (2010)
Sjöblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006)
Kumar, A. et al. Exome sequencing identifies a spectrum of mutation frequencies in advanced and lethal prostate cancers. Proc. Natl Acad. Sci. USA 108, 17087–17092 (2011)
Parsons, D. W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011)
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011)
Stransky, N. et al. The mutational landscape of head and neck squamous cell carcinoma. Science 333, 1157–1160 (2011)
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012)
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012)
Ehrlich, M., Norris, K. F., Wang, R. Y., Kuo, K. C. & Gehrke, C. W. DNA cytosine methylation and heat-induced deamination. Biosci. Rep. 6, 387–393 (1986)
Pavri, R. & Nussenzweig, M. C. AID targeting in antibody diversity. Adv. Immunol. 110, 1–26 (2011)
Yamanaka, S. et al. Apolipoprotein B mRNA-editing protein induces hepatocellular carcinoma and dysplasia in transgenic animals. Proc. Natl Acad. Sci. USA 92, 8483–8487 (1995)
Harris, R. S., Petersen-Mahrt, S. K. & Neuberger, M. S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002)
Refsland, E. W. et al. Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res. 38, 4274–4284 (2010)
Lackey, L. et al. APOBEC3B and AID have similar nuclear import mechanisms. J. Mol. Biol. 419, 301–314 (2012)
Albin, J. S. & Harris, R. S. Interactions of host APOBEC3 restriction factors with HIV-1 in vivo: implications for therapeutics. Expert Rev. Mol. Med. 12, e4 (2010)
Stenglein, M. D., Burns, M. B., Li, M., Lengyel, J. & Harris, R. S. APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nature Struct. Mol. Biol. 17, 222–229 (2010)
Suspène, R. et al. Somatic hypermutation of human mitochondrial and nuclear DNA by APOBEC3 cytidine deaminases, a pathway for DNA catabolism. Proc. Natl Acad. Sci. USA 108, 4858–4863 (2011)
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011)
Landry, S., Narvaiza, I., Linfesty, D. C. & Weitzman, M. D. APOBEC3A can activate the DNA damage response and cause cell-cycle arrest. EMBO Rep. 12, 444–450 (2011)
The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012)
Wei, X. et al. Exome sequencing identifies GRIN2A as frequently mutated in melanoma. Nature Genet. 43, 442–446 (2011)
Zhang, J. et al. International Cancer Genome Consortium Data Portal—a one-stop shop for cancer genomics data. Database 2011, bar026 (2011)
Di Noia, J. M. & Neuberger, M. S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007)
Loeb, L. A., Springgate, C. F. & Battula, N. Errors in DNA replication as a basis of malignant changes. Cancer Res. 34, 2311–2321 (1974)
Kidd, J. M., Newman, T. L., Tuzun, E., Kaul, R. & Eichler, E. E. Population stratification of a common APOBEC gene deletion polymorphism. PLoS Genet. 3, e63 (2007)
Carpenter, M. A. et al. Methylcytosine and normal cytosine deamination by the foreign DNA restriction enzyme APOBEC3A. J. Biol. Chem. 287, 34801–34808 (2012)
Shlyakhtenko, L. S. et al. Atomic force microscopy studies provide direct evidence for dimerization of the HIV restriction factor APOBEC3G. J. Biol. Chem. 286, 3387–3395 (2011)
Lea, D. E. & Coulson, C. A. The distribution of the numbers of mutants in bacterial populations. J. Genet. 49, 264–285 (1949)
Di Noia, J. & Neuberger, M. S. Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature 419, 43–48 (2002)
Huang, X. & Darzynkiewicz, Z. Cytometric assessment of histone H2AX phosphorylation: a reporter of DNA damage. Methods Mol. Biol. 314, 73–80 (2006)
Fairbairn, D. W., Olive, P. L. & O’Neill, K. L. The comet assay: a comprehensive review. Mutat. Res. 339, 37–59 (1995)
Acknowledgements
We thank J. Hultquist and R. Vogel for statistics, T. Hwang for bioinformatic assistance, V. Polunovsky for hTERT-HMECs, V. Simon for shRNA, S. Kaufmann, C. Lange and D. Largaespada for consultation, and the Masonic Cancer Center Breast Cancer Research Fund for purchasing the ATCC breast cancer panel. Tissues were obtained from the Masonic Cancer Center Tissue Procurement Facility, which is part of BioNet, supported by the Academic Health Center and National Institutes of Health (NIH) grants P30 CA77598 (D.Y.), P50 CA101955 (D. Buchsbaum) and KL2 RR033182 (B. Blazar). M.B.B. was supported in part by a Cancer Biology Training Grant (NIH NCI T32 CA009138) and a Department of Defense Breast Cancer Research Program Predoctoral Fellowship (BC101124). L.L. was supported in part by a National Science Foundation Predoctoral Fellowship and by a position on the Institute for Molecular Virology Training Grant NIH T32 AI083196. M.A.C. was supported by an NIH postdoctoral fellowship (F32 GM095219). A.M.L. was supported by a CIHR postdoctoral fellowship. E.W.R. was supported by a position on the Institute for Molecular Virology Training Grant NIH T32 AI083196 and subsequently by an NIH predoctoral fellowship (F31 DA033186). Computational analyses (N.A.T. and D.E.D.) were supported by federal funds from the National Cancer Institute, NIH, CBIIT/caBIG ISRCE yellow task 09-260. The Harris laboratory was supported in part by NIH R01 AI064046, NIH P01 GM091743, the Children's Cancer Research Fund, and a seed grant from the University of Minnesota Clinical and Translational Science Institute (supported by NIH 1UL1RR033183).
Author information
Authors and Affiliations
Contributions
R.S.H. conceived and managed the overall project. M.B.B. assisted R.S.H. with experimental design, project management and manuscript preparation. M.B.B., E.W.R. and B.L. generated mRNA expression profiles; L.L. and E.K.L. performed microscopy; L.L. and A.R. performed biochemical fractionations and DNA deaminase assays; M.B.B. performed uracil quantifications; A.M.L. performed thymidine kinase fluctuations; A.R. generated 3D-PCR sequences; and L.L., A.M.L., A.R. and M.A.C. determined the effect of induced A3B overexpression. M.A.C. performed deaminase assays with recombinant protein; and M.A.C. and D.K. assisted with the HPLC-ESI-MS/MS set up. N.T. was involved in HPLC-ESI-MS/MS method development. J.B.N. conducted the search and performed the bioinformatic analysis of the microarray data and developed the normalization algorithm for this analysis. N.A.T., D.E.D. and M.B.B. contributed bioinformatic analyses. All authors contributed to manuscript revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary Methods, Supplementary References, Supplementary Tables 1-11 and Supplementary Figures 1-16. (PDF 3228 kb)
Rights and permissions
About this article
Cite this article
Burns, M., Lackey, L., Carpenter, M. et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494, 366–370 (2013). https://doi.org/10.1038/nature11881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11881
This article is cited by
-
Mesoscale DNA features impact APOBEC3A and APOBEC3B deaminase activity and shape tumor mutational landscapes
Nature Communications (2024)
-
Distinguishing preferences of human APOBEC3A and APOBEC3B for cytosines in hairpin loops, and reflection of these preferences in APOBEC-signature cancer genome mutations
Nature Communications (2024)
-
The role of APOBEC3B in lung tumor evolution and targeted cancer therapy resistance
Nature Genetics (2024)
-
Progesterone from ovulatory menstrual cycles is an important cause of breast cancer
Breast Cancer Research (2023)
-
Sequence dependencies and mutation rates of localized mutational processes in cancer
Genome Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.