Abstract
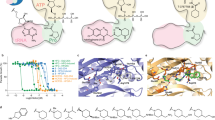
Febrifugine is the active component of the Chinese herb Chang Shan (Dichroa febrifuga Lour.)1,2, which has been used for treating malaria-induced fever for about 2,000 years. Halofuginone (HF), the halogenated derivative of febrifugine, has been tested in clinical trials for potential therapeutic applications in cancer and fibrotic disease3,4,5,6. Recently, HF was reported to inhibit TH17 cell differentiation by activating the amino acid response pathway7, through inhibiting human prolyl-transfer RNA synthetase (ProRS) to cause intracellular accumulation of uncharged tRNA8,9. Curiously, inhibition requires the presence of unhydrolysed ATP. Here we report an unusual 2.0 Å structure showing that ATP directly locks onto and orients two parts of HF onto human ProRS, so that one part of HF mimics bound proline and the other mimics the 3′ end of bound tRNA. Thus, HF is a new type of ATP-dependent inhibitor that simultaneously occupies two different substrate binding sites on ProRS. Moreover, our structure indicates a possible similar mechanism of action for febrifugine in malaria treatment. Finally, the elucidation here of a two-site modular targeting activity of HF raises the possibility that substrate-directed capture of similar inhibitors might be a general mechanism that could be applied to other synthetases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Koepfli, J. B., Mead, J. F. & Brockman, J. A., Jr An alkaloid with high antimalarial activity from Dichroa febrifuga . J. Am. Chem. Soc. 69, 1837 (1947)
Coatney, G. R., Cooper, W. C., Culwell, W. B., White, W. C. & Imboden, C. A., Jr Studies in human malaria. XXV. Trial of febrifugine, an alkaloid obtained from Dichroa febrifuga Lour., against the Chesson strain of Plasmodium vivax . J. Natl. Malar. Soc. 9, 183–186 (1950)
Pines, M., Snyder, D., Yarkoni, S. & Nagler, A. Halofuginone to treat fibrosis in chronic graft-versus-host disease and scleroderma. Biol. Blood Marrow Transplant. 9, 417–425 (2003)
Pines, M. & Nagler, A. Halofuginone: a novel antifibrotic therapy. Gen. Pharmacol. 30, 445–450 (1998)
de Jonge, M. J. et al. Phase I and pharmacokinetic study of halofuginone, an oral quinazolinone derivative in patients with advanced solid tumours. Eur. J. Cancer 42, 1768–1774 (2006)
Koon, H. B. et al. Phase II AIDS Malignancy Consortium trial of topical halofuginone in AIDS-related Kaposi sarcoma. J. Acquir. Immune Defic. Syndr. 56, 64–68 (2011)
Sundrud, M. S. et al. Halofuginone inhibits TH17 cell differentiation by activating the amino acid starvation response. Science 324, 1334–1338 (2009)
Keller, T. L. et al. Halofuginone and other febrifugine derivatives inhibit prolyl-tRNA synthetase. Nature Chem. Biol. 8, 311–317 (2012)
Kilberg, M. S., Pan, Y. X., Chen, H. & Leung-Pineda, V. Nutritional control of gene expression: how mammalian cells respond to amino acid limitation. Annu. Rev. Nutr. 25, 59–85 (2005)
Schimmel, P., Tao, J. & Hill, J. Aminoacyl tRNA synthetases as targets for new anti-infectives. FASEB J. 12, 1599–1609 (1998)
Hill, J. Aminoacyl sulfamides for the treatment of hyperproliferative disorders. US Patent 5,824. 657 (1998)
Carter, C. W., Jr Cognition, mechanism, and evolutionary relationships in aminoacyl-tRNA synthetases. Annu. Rev. Biochem. 62, 715–748 (1993)
Yaremchuk, A., Tukalo, M., Grotli, M. & Cusack, S. A succession of substrate induced conformational changes ensures the amino acid specificity of Thermus thermophilus prolyl-tRNA synthetase: comparison with histidyl-tRNA synthetase. J. Mol. Biol. 309, 989–1002 (2001)
Zhu, S. et al. Synthesis and biological evaluation of febrifugine analogues as potential antimalarial agents. Bioorg. Med. Chem. 17, 4496–4502 (2009)
Sankaranarayanan, R. et al. The structure of threonyl-tRNA synthetase-tRNAThr complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell 97, 371–381 (1999)
Heacock, D., Forsyth, C. J., Shiba, K. & Musier-Forsyth, K. Synthesis and aminoacyl-tRNA synthetase inhibitory activity of prolyl adenylate analogs. Bioorg. Chem. 24, 273–289 (1996)
Nakama, T., Nureki, O. & Yokoyama, S. Structural basis for the recognition of isoleucyl-adenylate and an antibiotic, mupirocin, by isoleucyl-tRNA synthetase. J. Biol. Chem. 276, 47387–47393 (2001)
Rock, F. L. et al. An antifungal agent inhibits an aminoacyl-tRNA synthetase by trapping tRNA in the editing site. Science 316, 1759–1761 (2007)
Pantoliano, M. W. et al. High-density miniaturized thermal shift assays as general strategy for drug discovery. J. Biomol. Screen. 6, 429–440 (2001)
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modeling. Bioinformatics 22, 195–201 (2006)
Beebe, K. et al. A universal plate format for increased throughput of assays that monitor multiple aminoacyl transfer RNA synthetase activities. Anal. Biochem. 368, 111–121 (2007)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16, 10881–10890 (1988)
Gouet, P., Courcelle, E., Stuart, D. I. & Metoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999)
Shi, J. P. & Schimmel, P. Aminoacylation of alanine minihelices. ‘Discriminator’ base modulates transition state of single turnover reaction. J. Biol. Chem. 266, 2705–2708 (1991)
Acknowledgements
We thank K. Musier-Forsyth for providing the ProRS gene plasmid and Pro-SA, staff at beamline 7-1 of Stanford Synchrotron Radiation Lightsource for assistance in X-ray diffraction data collection, and M. Guo for comments. This work was supported by National Institutes of Health grants GM15539, GM23562 and GM88278 and by a fellowship from the National Foundation for Cancer Research.
Author information
Authors and Affiliations
Contributions
H.Z., L.S., X.-L.Y. and P.S. designed the experiments. H.Z. and L.S. performed the experiments and all authors analysed the data. All authors discussed the results and H.Z. and P.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5 and Supplementary Table 1. (PDF 1223 kb)
Rights and permissions
About this article
Cite this article
Zhou, H., Sun, L., Yang, XL. et al. ATP-directed capture of bioactive herbal-based medicine on human tRNA synthetase. Nature 494, 121–124 (2013). https://doi.org/10.1038/nature11774
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11774
This article is cited by
-
Tyrosine-targeted covalent inhibition of a tRNA synthetase aided by zinc ion
Communications Biology (2023)
-
Elucidating the path to Plasmodium prolyl-tRNA synthetase inhibitors that overcome halofuginone resistance
Nature Communications (2022)
-
Structural basis of malaria parasite phenylalanine tRNA-synthetase inhibition by bicyclic azetidines
Nature Communications (2021)
-
Inhibitory mechanism of reveromycin A at the tRNA binding site of a class I synthetase
Nature Communications (2021)
-
Reprogramming of mitochondrial proline metabolism promotes liver tumorigenesis
Amino Acids (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.