Abstract
Cryptophyte and chlorarachniophyte algae are transitional forms in the widespread secondary endosymbiotic acquisition of photosynthesis by engulfment of eukaryotic algae. Unlike most secondary plastid-bearing algae, miniaturized versions of the endosymbiont nuclei (nucleomorphs) persist in cryptophytes and chlorarachniophytes. To determine why, and to address other fundamental questions about eukaryote–eukaryote endosymbiosis, we sequenced the nuclear genomes of the cryptophyte Guillardia theta and the chlorarachniophyte Bigelowiella natans. Both genomes have >21,000 protein genes and are intron rich, and B. natans exhibits unprecedented alternative splicing for a single-celled organism. Phylogenomic analyses and subcellular targeting predictions reveal extensive genetic and biochemical mosaicism, with both host- and endosymbiont-derived genes servicing the mitochondrion, the host cell cytosol, the plastid and the remnant endosymbiont cytosol of both algae. Mitochondrion-to-nucleus gene transfer still occurs in both organisms but plastid-to-nucleus and nucleomorph-to-nucleus transfers do not, which explains why a small residue of essential genes remains locked in each nucleomorph.
Similar content being viewed by others
Main
The photosynthetic organelles (plastids) of algae evolved from cyanobacteria by endosymbiosis1,2. The ‘primary’ plastids of red algae, glaucophyte algae and green algae, and their land-plant descendants, probably arose just once, more than a billion years ago3,4. Subsequent to this key event, the primary plastids of red and green algae were laterally transferred to other eukaryotes by secondary and tertiary endosymbioses, spawning some of the most abundant and ecologically important aquatic photosynthesizers on Earth such as diatoms, giant kelp, bloom-forming haptophytes and toxic dinoflagellates, as well as parasites such as the malaria pathogen Plasmodium3.
We have sequenced the nuclear genomes of two unicellular algae that are remarkable in their genetic and cellular complexity: the cryptophyte Guillardia theta and the chlorarachniophyte Bigelowiella natans. The secondary plastids of these independently evolved algae are unique in retaining a relict endosymbiont nucleus (the nucleomorph). Cryptophyte and chlorarachniophyte cells thus have four genomes and require complex subcellular protein-targeting machinery and inter-compartment coordination (Fig. 1). The B. natans nuclear genome is the first to be sequenced from a rhizarian protist, and the G. theta nuclear genome sequence is the first from a cryptophyte. They fill critical gaps on the tree of eukaryotic life, shed light on the pattern and process of host–endosymbiont integration, and reveal why nucleomorphs persist in cryptophytes and chlorarachniophytes but have been lost in other algae and parasites with secondary plastids.
The cryptophyte alga G. theta and the chlorarachniophyte alga B. natans have plastids bound by four membranes. In cryptophytes, the outermost plastid membrane is continuous with the nuclear envelope and its surface is studded with ribosomes, which co-translationally insert nucleus-encoded, organelle-targeted proteins. Between the inner and outer membrane pairs is the periplastidial compartment (PPC), which contains the nucleomorph (NM), the relict nucleus of the eukaryotic endosymbiont. The predicted numbers of protein-coding genes in the plastid, mitochondrial (MT), nucleomorph and nuclear genomes of G. theta and B. natans are shown. Additional abbreviations: C, carbohydrate; PY, pyrenoid.
Genomic and transcriptomic complexity
The nuclear genomes of B. natans and G. theta are approximately 95 and 87 megabase pairs (Mb) in size, respectively (Table 1 and Supplementary Tables 1.4.1 and 1.4.2; see Supplementary Information for sequencing and assembly details). Compared to the genomes of other secondary plastid-bearing algae, such as the diatoms Phaeodactylum tricornutum5 and Thalassiosira pseudonana6, and the filamentous brown alga Ectocarpus siliculosus7, the B. natans and G. theta genomes are gene rich (>21,000 predicted protein genes each, >85% of which are supported by RNA-seq data). Of the inferred proteins, 51% in G. theta and 47% in B. natans are unique, that is, have no detectable homologues in any other organism. Both genomes contain a large number of paralogues, constituting 2,636 multi-gene families in B. natans and 3,284 in G. theta (Supplementary Table 1.6.2).
As inferred from functional classifications based on the euKaryotic Orthologous Groups (KOG) database8, and protein family analyses (Supplementary Information 2.6), the G. theta and B. natans genomes are essentially ‘complete’ with respect to the major hallmarks of eukaryotic cellular complexity (>97% of a set of ‘core eukaryotic genes’9 are present in both organisms). These include components of the endomembrane system (Supplementary Information 2.6.3), transcription, RNA processing and modification, post-translational modification and protein turnover, and cytoskeleton. Examples of particularly large gene families include RNA processing and modification proteins, ankyrin repeat-containing proteins in B. natans (Supplementary Figs 1.6.3 and 1.6.5) and putative tyrosine kinases in G. theta (Supplementary Figs 1.6.4 and 1.6.6).
B. natans and G. theta protein genes are rich in spliceosomal introns. Examination of B. natans RNA-seq data revealed an unexpected finding: unlike all characterized unicellular species—indeed, unlike all characterized non-metazoans—B. natans shows complex and ubiquitous alternative splicing (Supplementary Information 2.2). Heavy use of various major alternative-splicing mechanisms was observed, including intron retention or inclusion (22% of B. natans introns were retained in >20% of the gene transcripts; Supplementary Fig. 2.2.1a) and exon skipping, which was found at levels higher than those observed in all characterized unicellular and multicellular species, and human tissues, being comparable only to the level observed in the human cerebral cortex (Supplementary Fig. 2.2.1b; exon skipping was confirmed by RNA-seq and expressed-sequence-tag (EST) data as well as polymerase chain reaction with reverse transcription (RT–PCR)). Hundreds of cases of alternative 5′ and 3′ splice-site usage were also identified, many involving alternative splicing at 3′ AG dinucleotides spaced three nucleotides apart (NAGNAG boundaries, Supplementary Fig. 2.2.5c), and whose role in expanding mammalian proteomes has been reported recently10.
We next examined the possible biological significance of the observed transcriptional complexity in B. natans. Two features of the B. natans alternative exons suggest that much of the exon skipping reflects spliceosomal ‘noise’ (that is, splicing errors). First, most skipped exons are nearly constitutively spliced (that is, skipped only occasionally), perhaps suggesting that exon skipping is not regulated (Supplementary Fig. 2.2.4b). Second, the proportion of exons that maintain reading frame (that is, are a multiple of three nucleotides) is close to random expectation (and similar to constitutive exons) (Supplementary Fig. 2.2.4c). This proportion is lower than that observed for cassette exons in human and fly, in which maintenance of the reading frame is associated with functional (and evolutionarily conserved) alternative splicing (for example, refs 11, 12). Nevertheless, even if most of the splicing complexity seen in B. natans simply reflects mis-splicing, many of these alternative transcripts might have important functions. A systematic survey of RNA-seq data identified 246 cases of genes whose alternative isoforms differentially include or exclude amino-terminal signal-peptide-encoding regions (three of which were verified by RT–PCR), suggesting that alternative splicing has a role in the generation of proteins targeted to different subcellular compartments (below). Alternative splicing has recently been shown to mediate dual targeting of glycolytic enzymes to the cytosol and peroxisome in fungi13.
Subcellular proteomes
Cryptophyte and chlorarachniophyte nucleomorphs are residual, endosymbiotic nuclei with tiny genomes <1 Mb in size14,15,16,17. The G. theta and B. natans nucleomorph genomes have only 487 (ref. 17) and 331 (ref. 15) protein genes, respectively, comprised of a limited set of ‘housekeeping’ genes, 31 or fewer genes for plastid-targeted proteins, and an abundance of ‘ORFan’ genes that typically show no detectable sequence similarity to known proteins14.
Like plastids and mitochondria, nucleomorphs and their genomes are reduced beyond self-sufficiency; they depend on nucleus-encoded proteins that are targeted to the periplastidial compartment (PPC), the residual endosymbiont cytoplasm in which the nucleomorph resides (Fig. 1). However, only a handful of PPC-targeted proteins are known (for example, see refs 18, 19, 20, 21) and the true extent of this dependence is unclear. Indeed, why nucleomorph genomes have been retained at all is a long-standing mystery of plastid evolution. Bearing in mind our knowledge of the G. theta and B. natans plastid22,23, nucleomorph15,17 and mitochondrial (this study) genome sequences, we carried out a comprehensive examination of nucleus-encoded proteins predicted to be targeted to each subcellular compartment (Supplementary Information 1.9), with emphasis on the PPC.
Our in silico-predicted mitochondrial, plastid, and PPC and nucleomorph proteomes for G. theta and B. natans are summarized in Supplementary Table 1.9.1 and Supplementary Fig. 1.9.4.1.2, together with a predicted set of >600 evolutionarily conserved endoplasmic reticulum and Golgi proteins (Supplementary Information 1.9). The limited overlap in proteins predicted to be targeted to different compartments suggests that the search strategies successfully differentiated among plastid-, PPC- and nucleomorph-, and host endoplasmic-reticulum- and Golgi-targeted proteins, which is important because in both cryptophytes and chlorarachniophytes the signal peptide secretion system is the first step in trafficking proteins to each of these compartments1. We analysed these proteomes in order to compare and contrast the biology of the independently evolved plastid and periplastidial compartments in G. theta and B. natans.
G. theta is predicted to have twice as many PPC- and nucleomorph-targeted proteins as B. natans (2,401 versus 1,002, after removal of ambiguously assigned proteins). A KOG-based breakdown of the unique (that is, non-overlapping) proteins in the PPC or nucleomorph proteomes revealed three KOG categories that are particularly ‘enriched’ in G. theta relative to B. natans: post-translational modification, protein turnover and chaperones; signal transduction; and carbohydrate transport and metabolism (Fig. 2a and Supplementary Table 1.9.4.1.1). The biological significance of these observations was further revealed through the mapping of nucleomorph- and nucleus-encoded, PPC-targeted proteins to KEGG (Kyoto Encyclopedia of Genes and Genomes) metabolic pathways (Supplementary Information 2.4). The G. theta PPC possesses canonical components of the protein-degrading proteasome, which B. natans seems to have lost entirely (KEGG pathway map 03050, Supplementary Fig. 2.4.1). G. theta also seems to have more PPC-localized proteins dedicated to protein folding (molecular chaperones) and RNA degradation, and a much greater diversity of metabolic enzymes, including those involved in amino acid biosynthesis (Supplementary Information 2.6.1 and Supplementary Fig. 2.4.1). In contrast, B. natans has a larger number of predicted nucleomorph-localized, spliceosome-associated proteins than does G. theta (Supplementary Fig. 2.4.1), which correlates with the marked difference in intron abundance: 852 in the B. natans nucleomorph genome15 versus just 17 in G. theta17).
a, Histogram showing the number of proteins predicted to be targeted to the PPC of G. theta and B. natans broken down by KOG functional category. For each KOG category, nucleomorph (NM)- and nucleus (NU)-encoded proteins are shown (PPC proteins predicted to be targeted to more than one subcellular compartment were removed; see Supplementary Fig. 1.9.4.1.2). b, Histogram showing the diversity of protein functions in the G. theta and B. natans PPC relative to free-living organisms (colour-coding as in a). Numbers of distinct KOG identifiers (IDs) in the PPC proteomes are plotted as a percentage of the average number of KOG IDs across 25 KOG categories for 6 organisms: Chlamydomonas reinhardtii, Ostreococcus tauri, Arabidopsis lyrata, Emiliania huxleyi, Dictyostelium purpureum and Phaeodactylum tricornutum (see Supplementary Information 1.9.4.3). Plastid and mitochondrial proteins were removed before calculating the averages (see Supplementary Information). KOG categories are as follows: A, RNA processing and modification; B, chromatin structure and dynamics; C, energy production and conversion; D, cell cycle control, cell division and chromosome partitioning; E, amino acid transport and metabolism; F, nucleotide transport and metabolism; G, carbohydrate transport and metabolism; H, coenzyme transport and metabolism; I, lipid transport and metabolism; J, translation, ribosomal structure and biogenesis; K, transcription; L, replication, recombination and repair; M, cell wall, membrane or envelope biogenesis; N, cell motility; O, post-translational modification, protein turnover, chaperones; P, inorganic ion transport and metabolism; Q, secondary metabolites biosynthesis, transport and catabolism; R, general function prediction only; S, function unknown; T, signal transduction; U, intracellular trafficking, secretion and vesicular transport; V, defence mechanisms; W, extracellular structures; Y, nuclear structure; Z, cytoskeleton. Higher KOG categories are as follows: CP, cellular processing and signalling; Hyp, poorly characterized; Inf, information storage and processing; Met, metabolism.
Host nuclear control over organelle biology is apparent in both G. theta and B. natans in the form of nucleus-encoded transcription-associated proteins (presumably regulating nucleomorph gene expression; Supplementary Information 2.6.1), putative DNA replication machinery, and ‘cell cycle-related’ proteins (for example, protein kinases) (Supplementary Fig. 2.4.1). Other processes in the PPC and nucleomorph are driven mainly by nucleomorph-encoded proteins, translation being a prominent example (Fig. 2a; KOG category J (translation, ribosomal structure and biogenesis)). Near-complete repertoires of small and large PPC ribosomal subunits could be inferred for G. theta, somewhat less so for B. natans, and in both cases the bulk of the constituent proteins are nucleomorph-encoded (Supplementary Fig. 2.4.2). This mirrors the pattern seen in plastid genomes24, in which core processes such as transcription and translation primarily involve proteins synthesized ‘on-site’.
Carbon metabolism differs substantially in G. theta and B. natans as inferred from the identification of putative carbohydrate-active enzymes (Supplementary Information 1.11 and Tables 2.3.1 and 2.3.2). Subcellular mapping of putative glycolysis-associated proteins in G. theta (Supplementary Fig. 2.4.5) reveals many PPC-localized reactions catalysed by key enzymes such as glucan, water dikinase, alpha-amylase, hexokinase, 6-phosphofructokinase and phosphoglucomutase. These enzymes form a link to the synthesis and degradation of starch, which occurs in the PPC in G. theta25. Thirty-six candidate proteins for PPC, plastid or cytosol metabolite shuttling in G. theta were identified from a set of 757 putative membrane-transport-associated proteins (Supplementary Information 1.10 and Supplementary Table 1.10.2). The distribution of glycolytic enzymes in B. natans is very different from that of G. theta, with a more heterogeneous mix of PPC-, plastid-, mitochondrion- and host cytosol-localized proteins (Supplementary Figs 2.4.3 and 2.4.4). In chlorarachniophytes the main carbohydrate storage product is a β-1,3-glucan located in the host cytoplasm26 (Fig. 1), and we identified numerous enzymes that are likely to have roles in β-glucan metabolism (Supplementary Information 2.3.2.1 and Supplementary Table 2.3.4).
We next examined the reduction in the PPC and nucleomorph proteomes of cryptophytes and chlorarachniophytes relative to the free-living organisms from which they evolved. We used the number of different KOG identifiers present in each of the 25 KOG functional categories as a measure of the diversity of biochemical processes taking place in the B. natans and G. theta PPC (taking into account nucleomorph-encoded proteins) (Supplementary Information 1.9.4.3). A total of 237 and 452 unique KOG identifiers were assigned to the B. natans and G. theta PPC proteome data sets, respectively (Supplementary Table 1.9.4.3.1). For most KOG categories the number of KOG identifiers in G. theta and B. natans is <25% of the average calculated from a set of 6 free-living organisms (algae with primary and secondary plastids plus a heterotrophic amoeba; Fig. 2b and Supplementary Information 1.9.4.3). Some functional categories are, predictably, completely absent in both organisms (for example, n = cell motility), whereas in G. theta the number of KOG identifiers in three different KOG categories exceeds 40% of the ‘free-living’ average (category J (translation, ribosomal structure and biogenesis), G (carbohydrate transport and metabolism) and P (inorganic ion transport and metabolism); Fig. 2b). On balance, the PPC of cryptophytes and chlorarachniophytes is highly reduced, but has retained an unexpectedly broad range of biochemical processes. These data provide the basis for addressing many fundamental questions about algal cell biology, including how many homologues of G. theta and B. natans PPC proteins are retained in the nucleomorph-lacking PPC of algae such as diatoms and haptophytes21, and what exactly are the biochemical determinants of the protein trafficking pathways in cryptophytes, chlorarachniophytes and other secondary plastid-bearing algae1,27. Making sense of the hundreds of predicted PPC and nucleomorph proteins in G. theta and B. natans with unknown functions (Supplementary Table 1.9.4.1.1) will be a substantial challenge.
Endosymbiotic gene transfer and replacement
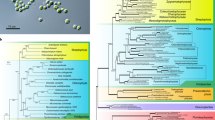
Endosymbiotic gene transfer (EGT)—the movement of DNA from endosymbiont to host before, during and after the evolution of an organelle—has had a notable role in the evolution of algae and their nuclear genomes28,29. The genomes of eukaryotes that are known or proposed to have undergone secondary endosymbioses involving red or green algal endosymbionts are now regularly queried for the presence or absence of so-called ‘red’ genes or ‘green’ genes (for example, see refs 30, 31). Organisms with (or thought to have once had) red algal secondary plastids are predicted to have ‘red’ genes in their nuclear genomes and ‘green’ genes should be found in the nuclei of organisms with green algal secondary plastids. Quantification of these algal signatures has the potential to answer fundamental questions about the spread and secondary loss of plastids across the eukaryotic tree but has led to conflicting results. For example, a large and unexpected number of ‘green’ genes were found in the genomes of diatoms32 and were interpreted to be evidence of a cryptic secondary endosymbiosis involving a green alga before the establishment of the red alga-derived plastid that diatoms currently harbour. However, these results were re-evaluated and found to be unconvincing33. The ability to detect genes of algal origin in nuclear genomes and to accurately distinguish between ‘red’ and ‘green’ has been shown to be complicated by a number of factors including taxonomic sampling bias, phylogenetic artefacts and large data sets consisting of thousands of complex trees that are invariably processed in an automated fashion31,33,34. We carried out a comprehensive phylogenomic investigation of EGT in the nuclear genomes of B. natans and G. theta, whose plastids and nucleomorphs are of green and red algal ancestry, respectively, using protocols and programs designed to address, to the extent possible, those issues mentioned above and other potential problems (Supplementary Information 1.12 and Supplementary Fig. 1.12.3).
From a set of 6,181 B. natans genes for which protein-based phylogenetic trees could be generated, automated tree sorting and manual curation resulted in the identification of 353 genes (5.7%) for which an algal origin could be confidently inferred (Fig. 3 and Supplementary Fig. 1.12.3). As expected, a large proportion (207; 59%) of these were green algal in nature, although 45 (22%) were classified as being derived from red algae. This pattern resembles that seen in an early EST-based analysis of B. natans proteins and was attributed to the mixotrophic lifestyle of chlorarachniophyte algae35. For G. theta, 508 of 7,451 genes (6.8%) were deemed to be algal in origin (Fig. 3 and Supplementary Fig. 1.12.3). Interestingly, more than twice as many G. theta genes in the manually curated set were classified as green (252) than red (100), and 9 examples of apparently glaucophyte-derived genes were identified. These results should be interpreted with caution, however, because although our analyses included all available red algal protein data sets (Supplementary Table 1.12.1), taxon sampling is still biased towards ‘green’ lineages. In fact, the majority (143 out of 252) of the protein trees for ‘green’ genes in G. theta contained no red algal homologues (Fig. 3) and 147 out of 508 were considered ‘algal’ but ambiguous with respect to which type. Thus, increased taxon sampling from red algae will presumably affect several predictions and perhaps enable more meaningful interpretation of others, as underscored by recent authors investigating ‘red’ and ‘green’ signals in other organisms with red secondary plastids31,33,34. The same can be said for interpretation of the red algal genes in B. natans, which has a green alga-derived secondary plastid. These uncertainties are exacerbated further by the still-unresolved phylogenetic position of the host component of cryptophytes relative to primary and secondary plastid-bearing algae36. Consequently, testing hypotheses about possible biological explanations for the diversity of algal nuclear genes seen in G. theta and B. natans, such as the relative contributions of endosymbiotic versus horizontal gene transfer, cannot currently be carried out without careful consideration of taxon sampling and methodological artefacts.
a, Histogram showing the proportion of ‘algal’ genes or proteins and their inferred origin by automated tree sorting and manual curation; bar height is relative to the total number of trees built for each organism and the raw counts are indicated on the bars (Supplementary Fig. 1.12.3). Exclusive affiliations are those in which the B. natans or G. theta homologue forms a clade solely with the group in question (for example, red algae), whereas inclusive affiliations enable sequences from other secondary and/or tertiary plastid-bearing algae within the clade to be present. ‘Green’ is defined as chlorophyte and/or streptophyte algae (including land plants). ‘Only plantae’ means trees containing only sequences from green algae and/or red algae and/or glaucophytes; algal origin therefore cannot be inferred with confidence. Only trees in the ‘red’, ‘green’ and ‘glaucophyte’ categories provide unambiguous information on the specific evolutionary origin of the B. natans or G. theta proteins. b, Pie charts showing the predicted locations of the algal proteins presented in a. Endoplasmic reticulum and Golgi proteins are those identified at the level of 75% confidence (see Supplementary Information 1.9.3). The ‘cytosolic’ category includes all proteins with no positive prediction for any of the four proteomes investigated.
Nevertheless, phylogenomic data taken together with subcellular targeting predictions show that the B. natans and G. theta nuclear genomes possess a complex mosaic of genes whose evolutionary histories do not reliably predict where their protein products function within the cell. A large portion of the alga-derived proteins identified in both organisms seem to function in their host cytosolic compartments, and clear examples of algal proteins targeted to the mitochondrion, endoplasmic reticulum or Golgi apparatus, plastid, and PPC or nucleomorph were also found (Fig. 3b). These results show that during the course of host–endosymbiont integration proteins often acquire new functions and/or new locations in which to function.
Gene duplication has also played a part in the ‘re-purposing’ of G. theta and B. natans nuclear genes of both host and endosymbiont ancestry (Supplementary Information 2.5.2 and Supplementary Fig. 2.5.2.2). Of the 508 ‘algal’ genes in the G. theta nuclear genome, 71 were found to belong to paralogous gene clusters (that is, genes that have duplicated subsequent to EGT); in ∼25% of these cases, paralogues encode proteins predicted to be targeted to multiple compartments, most often the PPC and host cytosol (Supplementary Fig. 2.5.2.1). A similar picture is seen in B. natans. In other cases the opposite pattern is observed, that is, duplication of apparent host-derived genes followed by organelle targeting of one or more of the protein products, sometimes as compensation for the loss of a nucleomorph gene (below). The present-day composition of the G. theta and B. natans subcellular proteomes is the product of extensive mixing and matching of proteins derived from their hosts and from endosymbionts that have become organelles. No clear pattern in the fates of individual endosymbiont-derived genes (loss, retention, duplication or re-purposing) is apparent.
Why do nucleomorphs persist?
Nuclear mitochondrial DNAs (NUMTs) and/or nuclear plastid DNAs (NUPTs) have been found in most eukaryotes studied so far; rates of EGT seem to vary substantially from lineage to lineage37,38. Although most such transfers involve small, apparently random fragments of organellar DNA that have no notable impact on the nuclear genome, entire genes can be transferred and expressed in their new environment (for example, see refs 39, 40). Instances in which NUMTs have altered existing genes by introducing new introns or truncating the gene through frameshifts have also been observed (for example, see refs 41, 42, 43).
Nuclear genome sequences from a rhizarian and a cryptophyte provide the first opportunity to test if NUMTs, NUPTs and, most importantly, nucleomorph-derived DNAs (NUNMs) reside in these genomes. Given that these types of ‘recent’ EGT recapitulate an important process by which endosymbiont and organellar genomes are initially reduced and can ultimately be lost, the presence or absence of NUPTs and NUNMs has the potential to provide insight into the fate of plastid and nucleomorph genomes of secondary endosymbiotic origin. A bioinformatic screen (Supplementary Information 2.5) revealed seven NUMTs in the B. natans nuclear genome (Supplementary Table 2.5.3.1) and 13 in G. theta (Supplementary Table 2.5.3.2). All of the fragments were small (<320 nucleotides) and none contained entire genes. Their point of origin in their respective mitochondrial genomes seems random both in terms of content and position, and most were integrated into non-coding regions.
In marked contrast, no recent transfers of NUPTs or NUNMs were observed in B. natans or G. theta. The presence of NUMTs in both nuclear genomes demonstrates that EGT happens, so there is no obvious impediment to the incorporation of organelle-donated DNA. One explanation for the apparent absence of NUNMs and NUPTs in the B. natans and G. theta genomes is the ‘limited transfer window’ hypothesis, which posits that cells with multiple copies of an organelle are more likely to have EGTs than those with single organelles because lysis of a single organelle to release DNA into the host nucleocytoplasm would be catastrophic44,45. Consistent with our observations, G. theta and B. natans cells have a single plastid–nucleomorph complex per cell, the lysis of which would presumably be fatal. In contrast, cryptophytes can have large, reticulate mitochondria that undergo fission and fusion46, and in chlorarachniophytes each cell generally has multiple mitochondria that reside in both the cytoplasm and (when present) filopodia47. The ability of the limited transfer window hypothesis to explain the absence of NUPTs and NUNMs in G. theta and B. natans could be tested further by searching the nuclear genomes of cryptophytes and chlorarachniophytes that contain multiple plastid–nucleomorph complexes per cell48,49.
The consequences of the lack of NUNMs and NUPTs in the B. natans and G. theta nuclear genomes are considerable. In the absence of EGT, inactivation and loss of essential plastid and nucleomorph genes cannot be compensated for by the classical gene transfer–protein re-targeting scenario, as occurs in other systems39,50. Our results show that indirect ‘solutions’ have evolved, most notably the duplication and functional reassignment of host-derived nuclear genes. For example, a nucleus-encoded cyclin-dependent kinase regulatory subunit protein (also known as kin(cdc)) predicted to function in the G. theta PPC or nucleomorph is not specifically related to kin(cdc) homologues encoded in the nucleomorph genomes of two other cryptophytes14,16, but instead is a recent duplicate of an apparently host-derived homologue (Supplementary Fig. 2.5.2.2a). A similar pattern is seen in a variety of other nucleus-encoded, PPC-targeted proteins in G. theta (Supplementary Information 2.5.5 and Supplementary Table 2.5.5.1). In B. natans, alternative splicing may serve as an additional mechanism for increasing proteome complexity and compensating for the loss of organellar genes. Over recent evolutionary time scales, nucleomorph genome reduction seems to have slowed markedly for lack of an easy solution to the problem of nucleomorph gene loss.
Extensive EGT has nevertheless occurred in the ancestors of B. natans and G. theta. Some of the protein products of these transferred genes are targeted to the plastid and PPC but most are not (Fig. 3b). Genetic and biochemical mosaicism is thus rampant in both organisms, with host-, endosymbiont- and foreign alga-derived proteins contributing to processes taking place in their various subcellular compartments. The extent to which such mosaicism exists in other cryptophytes and chlorarachniophytes remains unknown. Nevertheless, it seems likely that close inspection of the genomes of all algae that evolved by eukaryote–eukaryote endosymbiosis will reveal a level of mosaicism beyond that which is typically assumed. This conclusion has important implications for the use of genomic data to infer a robust tree of eukaryotes that includes secondary and tertiary plastid-bearing phototrophs, and more generally, for our understanding of the evolution of the eukaryotic cell.
Methods Summary
DNA was extracted from axenic cultures established from single-cell isolates of Guillardia theta and Bigelowiella natans (Bigelow Laboratory for Ocean Sciences). Three different-sized libraries, 3-kb, 8-kb and 34-kb fosmids, were generated and sequenced at the Joint Genome Institute (JGI) using Sanger sequencing. Additional 454 sequencing was used to fill gaps, and sequence reads were assembled using a modified version of Arachne. Gene models were generated and annotated for the resulting genomic scaffolds using JGI's gene modelling pipeline with additional manual curation. Gene modelling, annotation and alternative splicing analyses were assisted by three messenger RNA data sets: ESTs generated before the genome projects, JGI-generated ESTs and RNA-seq data. Proteome predictions for the plastid, mitochondrion, endoplasmic reticulum or Golgi, and periplastidial compartment were generated using independent bioinformatic pipelines. Maximum likelihood phylogenetic trees were generated from protein sequences retrieved from a local database and the positions of the B. natans and G. theta proteins were assessed using a combination of automated filtering and manual curation. Complete materials and methods are described in the Supplementary Information.
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The G. theta and B. natans genome sequences and annotations are available through the JGI Genome Portal at http://jgi.doe.gov/Gtheta and http://jgi.doe.gov/Bnatans and have been deposited in GenBank under the accession numbers AEIE00000000 and ADNK00000000, respectively.
References
Gould, S. B., Waller, R. F. & McFadden, G. I. Plastid evolution. Annu. Rev. Plant Biol. 59, 491–517 (2008)
Gray, M. W. The endosymbiont hypothesis revisited. Int. Rev. Cytol. 141, 233–357 (1992)
Palmer, J. D. The symbiotic birth and spread of plastids: how many times and whodunnit? J. Phycol. 39, 4–11 (2003)
Yoon, H. S., Hackett, J. D., Ciniglia, C., Pinto, G. & Bhattacharya, D. A molecular timeline for the origin of photosynthetic eukaryotes. Mol. Biol. Evol. 21, 809–818 (2004)
Bowler, C. et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456, 239–244 (2008)
Armbrust, E. V. et al. The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306, 79–86 (2004)
Cock, J. M. et al. The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature 465, 617–621 (2010)
Koonin, E. V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5, R7 (2004)
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007)
Bradley, R. K., Merkin, J., Lambert, N. J. & Burge, C. B. Alternative splicing of RNA triplets is often regulated and accelerates proteome evolution. PLoS Biol. 10, e1001229 (2012)
Irimia, M. et al. Widespread evolutionary conservation of alternatively spliced exons in Caenorhabditis. Mol. Biol. Evol. 25, 375–382 (2008)
Sorek, R., Shamir, R. & Ast, G. How prevalent is functional alternative splicing in the human genome? Trends Genet. 20, 68–71 (2004)
Freitag, J., Ast, J. & Bolker, M. Cryptic peroxisomal targeting via alternative splicing and stop codon read-through in fungi. Nature 485, 522–525 (2012)
Tanifuji, G. et al. Complete nucleomorph genome sequence of the nonphotosynthetic alga Cryptomonas paramecium reveals a core nucleomorph gene set. Genome Biol. Evol. 3, 44–54 (2011)
Gilson, P. R. et al. Complete nucleotide sequence of the chlorarachniophyte nucleomorph: nature's smallest nucleus. Proc. Natl Acad. Sci. USA 103, 9566–9571 (2006)
Lane, C. E. et al. Nucleomorph genome of Hemiselmis andersenii reveals complete intron loss and compaction as a driver of protein structure and function. Proc. Natl Acad. Sci. USA 104, 19908–19913 (2007)
Douglas, S. E. et al. The highly reduced genome of an enslaved algal nucleus. Nature 410, 1091–1096 (2001)
Gould, S. B. et al. Nucleus-to-nucleus gene transfer and protein retargeting into a remnant cytoplasm of cryptophytes and diatoms. Mol. Biol. Evol. 23, 2413–2422 (2006)
Gile, G. H. & Keeling, P. J. Nucleus-encoded periplastid-targeted EFL in chlorarachniophytes. Mol. Biol. Evol. 25, 1967–1977 (2008)
Hirakawa, Y., Burki, F. & Keeling, P. J. Nucleus- and nucleomorph-targeted histone proteins in a chlorarachniophyte alga. Mol. Microbiol. 80, 1439–1449 (2011)
Moog, D., Stork, S., Zauner, S. & Maier, U. G. In silico and in vivo investigations of the proteins of a minimized eukaryotic cytoplasm. Genome Biol. Evol. 3, 375–382 (2011)
Douglas, S. E. & Penny, S. L. The plastid genome of the cryptophyte alga, Guillardia theta: complete sequence and conserved synteny groups confirm its common ancestry with red algae. J. Mol. Evol. 48, 236–244 (1999)
Rogers, M. B., Gilson, P. R., Su, V., McFadden, G. I. & Keeling, P. J. The complete chloroplast genome of the chlorarachniophyte Bigelowiella natans: evidence for independent origins of chlorarachniophyte and euglenid secondary endosymbionts. Mol. Biol. Evol. 24, 54–62 (2007)
Martin, W. & Herrmann, R. G. Gene transfer from organelles to the nucleus: how much, what happens, and why? Plant Physiol. 118, 9–17 (1998)
Deschamps, P. et al. Nature of the periplastidial pathway of starch synthesis in the cryptophyte Guillardia theta. Eukaryot. Cell 5, 954–963 (2006)
McFadden, G. I., Gilson, P. R. & Sims, I. M. Preliminary characterization of carbohydrate stores from chlorarachniophytes (Division: Chlorarachniophyta). Phycol. Res. 45, 145–151 (1997)
Hirakawa, Y., Gile, G. H., Ota, S., Keeling, P. J. & Ishida, K. Characterization of periplastidal compartment-targeting signals in chlorarachniophytes. Mol. Biol. Evol. 27, 1538–1545 (2010)
Martin, W., Brinkmann, H., Savonna, C. & Cerff, R. Evidence for a chimeric nature of nuclear genomes: eubacterial origin of eukaryotic glyceraldehyde-3-phosphate dehydrogenase genes. Proc. Natl Acad. Sci. USA 90, 8692–8696 (1993)
Timmis, J. N., Ayliffe, M. A., Huang, C. Y. & Martin, W. Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nature Rev. Genet. 5, 123–135 (2004)
Stiller, J. W., Huang, J., Ding, Q., Tian, J. & Goodwillie, C. Are algal genes in nonphotosynthetic protists evidence of historical plastid endosymbioses? BMC Genomics 10, 484 (2009)
Woehle, C., Dagan, T., Martin, W. F. & Gould, S. B. Red and problematic green phylogenetic signals among thousands of nuclear genes from the photosynthetic and apicomplexa-related Chromera velia. Genome Biol. Evol. 3, 1220–1230 (2011)
Moustafa, A. et al. Genomic footprints of a cryptic plastid endosymbiosis in diatoms. Science 324, 1724–1726 (2009)
Deschamps, P. & Moreira, D. Reevaluating the green contribution to diatom genomes. Genome Biol. Evol. 4, 683–688 (2012)
Burki, F. et al. Re-evaluating the green versus red signal in eukaryotes with secondary plastid of red algal origin. Genome Biol. Evol. 4, 738–747 (2012)
Archibald, J. M., Rogers, M. B., Toop, M., Ishida, K. & Keeling, P. J. Lateral gene transfer and the evolution of plastid-targeted proteins in the secondary plastid-containing alga Bigelowiella natans. Proc. Natl Acad. Sci. USA 100, 7678–7683 (2003)
Burki, F., Okamoto, N., Pombert, J. F. & Keeling, P. J. The evolutionary history of haptophytes and cryptophytes: phylogenomic evidence for separate origins. Proc. R. Soc. B 279, 2246–2254 (2012)
Richly, E. & Leister, D. NUMTs in sequenced eukaryotic genomes. Mol. Biol. Evol. 21, 1081–1084 (2004)
Richly, E. & Leister, D. NUPTs in sequenced eukaryotes and their genomic organization in relation to NUMTs. Mol. Biol. Evol. 21, 1972–1980 (2004)
Adams, K. L. & Palmer, J. D. Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Mol. Phylogenet. Evol. 29, 380–395 (2003)
Stegemann, S. & Bock, R. Experimental reconstruction of functional gene transfer from the tobacco plastid genome to the nucleus. Plant Cell 18, 2869–2878 (2006)
Ricchetti, M., Tekaia, F. & Dujon, B. Continued colonization of the human genome by mitochondrial DNA. PLoS Biol. 2, e273 (2004)
Noutsos, C., Kleine, T., Armbruster, U., DalCorso, G. & Leister, D. Nuclear insertions of organellar DNA can create novel patches of functional exon sequences. Trends Genet. 23, 597–601 (2007)
Curtis, B. A. & Archibald, J. M. A spliceosomal intron of mitochondrial DNA origin. Curr. Biol. 20, R919–R920 (2010)
Barbrook, A. C., Howe, C. J. & Purton, S. Why are plastid genomes retained in non-photosynthetic organisms? Trends Plant Sci. 11, 101–108 (2006)
Smith, D. R., Crosby, K. & Lee, R. W. Correlation between nuclear plastid DNA abundance and plastid number supports the limited transfer window hypothesis. Genome Biol. Evol. 3, 365–371 (2011)
Hill, D. R. A. & Wetherbee, R. Proteomonas sulcata gen. et sp. nov. (Cryptophyceae), a cryptomonad with two morphologically distinct and alternating forms. Phycologia 25, 521–543 (1986)
Hibberd, D. J. & Norris, R. E. Cytology and ultrastructure of Chlorarachnion reptans (Chlorarachniophyta divisio nova, Chlorarachniophyceae classis nova). J. Phycol. 20, 310–330 (1984)
Kugrens, P. & Clay, B. L. in Freshwater Algae of North America 715–755 (Elsevier Science, 2003)
Ota, S., Kudo, A. & Ishida, K.-I. Gymnochlora dimorpha sp. nov., a new chlorarachniophyte with unique daughter cell behavior. Phycologia 50, 317–326 (2011)
Martin, W. et al. Gene transfer to the nucleus and the evolution of chloroplasts. Nature 393, 162–165 (1998)
Acknowledgements
The work conducted by the US Department of Energy Joint Genome Institute is supported by the Office of Science of the US Department of Energy under Contract no. DE-AC02-05CH11231. RNA-seq data used in the paper were generated by the National Center for Genome Resources as part of the Gordon and Betty Moore Foundation’s Marine Microbial Eukaryote Transcriptome Project. B.A.C. and J.F.H. were supported by a Special Research Opportunities Grant from the Natural Sciences and Engineering Research Council of Canada awarded to J.M.A. and M.W.G. J.M.A., P.J.K., M.W.G. and C.H.S. are members of the Canadian Institute for Advanced Research, Program in Integrated Microbial Biodiversity. G.I.M. is an Australian Research Council Federation Fellow and a Howard Hughes International Scholar. We thank R. A. Andersen (Bigelow Laboratories) for assistance with single-cell isolations, C. X. Chan for a tree-sorting PERL script, H. Gutierrez for help with SM protein family annotation, and B. Read for permission to analyse the Emiliania huxleyi genome sequenced by the JGI.
Author information
Authors and Affiliations
Contributions
Nucleic acid sample preparation: C.E.L., D.F.S. and J.F.H. Genome and transcriptome sequencing and assembly: J.S., J.G., C.B., A.K.B., J.A.C., R.K., E.L. and S.L. Genome annotation and/or analysis: B.A.C., G.T., F.B., A.G., M.I., S.M., M.C.A., S.G.B., G.H.G., Y.H., J.F.H., A.K., S.A.R., J.S., A. Symeonidi, M.E., R.J.M.E., E.K.H., M.J.K., T.N., M.O., A.R.-P., E.V.A., S.J.A., R.G.B., P.C., J.B.D., D.G.D., N.M.F., B.R.G., C.J.G., F.H., B.H., M.P.H., K.-I.I., E.K., L.K., P.G.K., Y.L., S.-B.M., U.G.M., D.M., T.M., J.A.D.N., N.T.O., A.M.P., E.J.P., T.A.R., G.R., S.W.R., C.S., S. Schaack., S. Shirato, C.H.S., S. Suzuki, A.Z.W., S.Z., J.G., A. Salamov, C.E.L., M.W.G. and J.M.A. Project management: K.B., I.V.G. and J.S. Project coordination: J.M.A., M.W.G., P.J.K., C.E.L. and G.I.M. Writing: J.M.A., B.A.C., M.W.G., G.I.M., P.J.K., C.E.L., G.T., F.B., A.G., M.I., S.M., M.C.A., S.G.B., G.H.G., J.F.H., A.K., S.A.R., J.S., A. Symeonidi, R.J.M.E., E.K.H., M.J.K., T.N., A.R.-P., J.B.D., E.K., P.G.K., E.J.P., S.W.R., S.S., A.K.B. and I.V.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Data and Analyses for the Guillardia theta and Bigelowiella natans nuclear genome sequences. Supplementary References, 35 Supplementary Figures and 24 Supplementary Tables are also included (see Table of Contents for more details). (PDF 10546 kb)
Supplementary Table S1.9.4.2.1
This file contains datasets used for support vector machine prediction of PPC proteins in Bigelowiella natans and Guillardia theta. (XLS 79 kb)
Supplementary Table S1.9.4.2.2
This file contains dataset sizes used for support vector machine prediction of PPC proteins in Bigelowiella natans and Guillardia theta. (XLS 22 kb)
Supplementary Table S1.12.1
This file contains a list of taxa used in phylogenomic analyses of the nuclear genomes of Bigelowiella natans and Guillardia theta. (XLS 107 kb)
Supplementary Table S2.3.3
This file contains proteins identified in the search for putative carbohydrate-active enzymes in Guillardia theta. (XLS 56 kb)
Supplementary Table S2.3.4
This file contains proteins identified in the search for putative carbohydrate-active enzymes in Bigelowiella natans. (XLS 50 kb)
Supplementary Table S2.3.4.1
This file contains a list of proteins possibly involved in the ER-localized synthesis of N-glycanes in Guillardia theta and Bigelowiella natans. (XLS 27 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Curtis, B., Tanifuji, G., Burki, F. et al. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature 492, 59–65 (2012). https://doi.org/10.1038/nature11681
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11681
This article is cited by
-
Horizontal gene transfer in eukaryotes: aligning theory with data
Nature Reviews Genetics (2024)
-
Contrasting outcomes of genome reduction in mikrocytids and microsporidians
BMC Biology (2023)
-
Estimating transcriptome complexities across eukaryotes
BMC Genomics (2023)
-
Molecular dissection of the soluble photosynthetic antenna from the cryptophyte alga Hemiselmis andersenii
Communications Biology (2023)
-
Spatial fragmentation in the distribution of diatom endosymbionts from the taxonomically clarified dinophyte Kryptoperidinium triquetrum (= Kryptoperidinium foliaceum, Peridiniales)
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.