Abstract
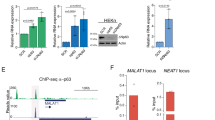
Several of the thousands of human long non-coding RNAs (lncRNAs) have been functionally characterized1,2,3,4; however, potential roles for lncRNAs in somatic tissue differentiation remain poorly understood. Here we show that a 3.7-kilobase lncRNA, terminal differentiation-induced ncRNA (TINCR), controls human epidermal differentiation by a post-transcriptional mechanism. TINCR is required for high messenger RNA abundance of key differentiation genes, many of which are mutated in human skin diseases, including FLG, LOR, ALOXE3, ALOX12B, ABCA12, CASP14 and ELOVL3. TINCR-deficient epidermis lacked terminal differentiation ultrastructure, including keratohyalin granules and intact lamellar bodies. Genome-scale RNA interactome analysis revealed that TINCR interacts with a range of differentiation mRNAs. TINCR–mRNA interaction occurs through a 25-nucleotide ‘TINCR box’ motif that is strongly enriched in interacting mRNAs and required for TINCR binding. A high-throughput screen to analyse TINCR binding capacity to approximately 9,400 human recombinant proteins revealed direct binding of TINCR RNA to the staufen1 (STAU1) protein. STAU1-deficient tissue recapitulated the impaired differentiation seen with TINCR depletion. Loss of UPF1 and UPF2, both of which are required for STAU1-mediated RNA decay, however, did not have differentiation effects. Instead, the TINCR–STAU1 complex seems to mediate stabilization of differentiation mRNAs, such as KRT80. These data identify TINCR as a key lncRNA required for somatic tissue differentiation, which occurs through lncRNA binding to differentiation mRNAs to ensure their expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009)
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009)
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007)
Martianov, I., Ramadass, A., Serra Barros, A., Chow, N. & Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 445, 666–670 (2007)
Pauli, A., Rinn, J. L. & Schier, A. F. Non-coding RNAs as regulators of embryogenesis. Nature Rev. Genet. 12, 136–149 (2011)
Wan, D. et al. Large-scale cDNA transfection screening for genes related to cancer development and progression. Proc. Natl Acad. Sci. USA 101, 15724–15729 (2004)
Sen, G. L., Reuter, J. A., Webster, D. E., Zhu, L. & Khavari, P. A. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463, 563–567 (2010)
Truong, A. B., Kretz, M., Ridky, T. W., Kimmel, R. & Khavari, P. A. p63 regulates proliferation and differentiation of developmentally mature keratinocytes. Genes Dev. 20, 3185–3197 (2006)
O’Driscoll, J. et al. A recurrent mutation in the loricrin gene underlies the ichthyotic variant of Vohwinkel syndrome. Clin. Exp. Dermatol. 27, 243–246 (2002)
Smith, F. J. et al. Loss-of-function mutations in the gene encoding filaggrin cause ichthyosis vulgaris. Nature Genet. 38, 337–342 (2006)
Virtanen, M., Smith, S. K., Gedde-Dahl, T., Jr, Vahlquist, A. & Bowden, P. E. Splice site and deletion mutations in keratin (KRT1 and KRT10) genes: unusual phenotypic alterations in Scandinavian patients with epidermolytic hyperkeratosis. J. Invest. Dermatol. 121, 1013–1020 (2003)
Elias, P. M. Stratum corneum defensive functions: an integrated view. J. Invest. Dermatol. 125, 183–200 (2005)
Eckl, K. M. et al. Molecular analysis of 250 patients with autosomal recessive congenital ichthyosis: evidence for mutation hotspots in ALOXE3 and allelic heterogeneity in ALOX12B. J. Invest. Dermatol. 129, 1421–1428 (2009)
Sakai, K. et al. ABCA12 is a major causative gene for non-bullous congenital ichthyosiform erythroderma. J. Invest. Dermatol. 129, 2306–2309 (2009)
Westerberg, R. et al. Role for ELOVL3 and fatty acid chain length in development of hair and skin function. J. Biol. Chem. 279, 5621–5629 (2004)
Denecker, G. et al. Caspase-14 protects against epidermal UVB photodamage and water loss. Nature Cell Biol. 9, 666–674 (2007)
Raj, A., van den Bogaard, P., Rifkin, S. A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nature Methods 5, 877–879 (2008)
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011)
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010)
Nagano, T. et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720 (2008)
Tsai, M. C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010)
Dugré-Brisson, S. et al. Interaction of Staufen1 with the 5′ end of mRNA facilitates translation of these RNAs. Nucleic Acids Res. 33, 4797–4812 (2005)
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011)
Kiebler, M. A. et al. The mammalian staufen protein localizes to the somatodendritic domain of cultured hippocampal neurons: implications for its involvement in mRNA transport. J. Neurosci. 19, 288–297 (1999)
St Johnston, D., Beuchle, D. & Nusslein-Volhard, C. Staufen, a gene required to localize maternal RNAs in the Drosophila egg. Cell 66, 51–63 (1991)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Furic, L., Maher-Laporte, M. & DesGroseillers, L. A genome-wide approach identifies distinct but overlapping subsets of cellular mRNAs associated with Staufen1- and Staufen2-containing ribonucleoprotein complexes. RNA 14, 324–335 (2008)
Merino, E. J., Wilkinson, K. A., Coughlan, J. L. & Weeks, K. M. RNA structure analysis at single nucleotide resolution by selective 2'-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005)
Bond, A. M. et al. Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nature Neurosci. 12, 1020–1027 (2009)
Young, T. L., Matsuda, T. & Cepko, C. L. The noncoding RNA Taurine Upregulated Gene 1 is required for differentiation of the murine retina. Curr. Biol. 15, 501–512 (2005)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Das, R., Laederach, A., Pearlman, S. M., Herschlag, D. & Altman, R. B. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005)
Deigan, K. E., Li, T. W., Mathews, D. H. & Weeks, K. M. Accurate SHAPE-directed RNA structure determination. Proc. Natl Acad. Sci. USA 106, 97–102 (2009)
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003)
Acknowledgements
This work was supported by US Veterans Affairs Office of Research and Development funding to P.A.K. and National Institutes of Health (National Institute of Arthritis and Musculoskeletal and Skin Diseases) grant AR49737 to P.A.K., and by NIH R01-HG004361 and California Institute for Regenerative Medicine to H.Y.C. H.Y.C. is an Early Career Scientist of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
M.K. designed and executed experiments, analysed data and wrote the manuscript. D.E.W., Z.S., C.C., A.Z., C.S.L., R.J.F., K.Q., J.C., D.J., G.X.Y.Z., G.E.K., A.F.G., R.C.S., R.A.F. and S.A. executed experiments, analysed data and contributed to design of experimentation. A.R. and J.L.R. helped design experiments and analysed data. P.A.K. and H.Y.C. designed experiments, analysed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 and Supplementary Tables 1-8. (PDF 966 kb)
Rights and permissions
About this article
Cite this article
Kretz, M., Siprashvili, Z., Chu, C. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013). https://doi.org/10.1038/nature11661
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11661
This article is cited by
-
Involvement of transcribed lncRNA uc.291 in hyperproliferative skin disorders
Biology Direct (2023)
-
Epigenetics, cryptorchidism, and infertility
Basic and Clinical Andrology (2023)
-
LncRNA INHEG promotes glioma stem cell maintenance and tumorigenicity through regulating rRNA 2’-O-methylation
Nature Communications (2023)
-
Epigenetic regulation in hematopoiesis and its implications in the targeted therapy of hematologic malignancies
Signal Transduction and Targeted Therapy (2023)
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.