Abstract
For 10,000 years pigs and humans have shared a close and complex relationship. From domestication to modern breeding practices, humans have shaped the genomes of domestic pigs. Here we present the assembly and analysis of the genome sequence of a female domestic Duroc pig (Sus scrofa) and a comparison with the genomes of wild and domestic pigs from Europe and Asia. Wild pigs emerged in South East Asia and subsequently spread across Eurasia. Our results reveal a deep phylogenetic split between European and Asian wild boars ∼1 million years ago, and a selective sweep analysis indicates selection on genes involved in RNA processing and regulation. Genes associated with immune response and olfaction exhibit fast evolution. Pigs have the largest repertoire of functional olfactory receptor genes, reflecting the importance of smell in this scavenging animal. The pig genome sequence provides an important resource for further improvements of this important livestock species, and our identification of many putative disease-causing variants extends the potential of the pig as a biomedical model.
Similar content being viewed by others
Main
The domestic pig (Sus scrofa) is a eutherian mammal and a member of the Cetartiodactyla order, a clade distinct from rodent and primates, that last shared a common ancestor with humans between 79 and 97 million years (Myr) ago1,2 (http://www.timetree.net). Molecular genetic evidence indicates that Sus scrofa emerged in South East Asia during the climatic fluctuations of the early Pliocene 5.3–3.5 Myr ago. Then, beginning ∼10,000 years ago, pigs were domesticated in multiple locations across Eurasia3 (Frantz, L. A. F. et al., manuscript submitted).
Here we provide a high-quality draft pig genome sequence developed under the auspices of the Swine Genome Sequencing Consortium4,5, established using bacterial artificial chromosome (BAC)6 and whole-genome shotgun (WGS) sequences (see Methods and Supplementary Information). The assembly (Sscrofa10.2) comprises 2.60 gigabases (Gb) assigned to chromosomes with a further 212 megabases (Mb) in unplaced scaffolds (Table 1 and Supplementary Tables 1–3).
Genome annotation
A de novo repeat discovery and annotation strategy (Supplementary Fig. 8) revealed a total of 95 novel repeat families, including: 5 long interspersed elements (LINEs), 6 short interspersed elements (SINEs), 8 satellites and 76 long terminal repeats (LTRs). The relative content of repetitive elements (∼40%, Supplementary Figs 9 and 10) is lower than reported for other mammalian genomes. The main repetitive element groups are the LINE1 and glutamic acid transfer RNA (tRNAGlu)-derived SINEs or PRE (porcine repetitive element). The expansion of PRE is specific to the porcine lineage. Phylogenetic analysis of LINE1 and PRE (Supplementary Figs 13 and 14) indicates that only a single lineage of each is currently active and that the main expansion of both LINE1 and PRE occurred in the first half of the Tertiary period. Smaller expansions, particularly in LINE1, have occurred since, but recent activity is very low (Supplementary Information).
Annotation of genes, transcripts and predictions of orthologues and paralogues was performed using the Ensembl analysis pipeline7 (Table 1 and Supplementary Figs 3–7). Further annotation for non-protein-coding RNAs (ncRNAs) was undertaken with another analysis pipeline (Supplementary Information and Supplementary Table 4).
Evolution of the porcine genome
Evolution of genes and gene families
To examine the mutation rate and type of protein-coding genes that show accelerated evolution in pigs, we identified ∼9,000 as 1:1 orthologues within a group of six mammals (human, mouse, dog, horse, cow and pig). This orthologous gene set was used to identify proteins that show accelerated evolution in each of these six mammalian lineages (Supplementary Information). The observed number of synonymous substitutions per synonymous site (dS) for the pig lineage (0.160) is similar to that of the other mammals (0.138–0.201) except for the mouse (0.458), indicating similar evolutionary rates in pigs and other mammals. The observed dN/dS ratio (ratio of the rate of non-synonymous substitutions to the rate of synonymous substitutions) of 0.144 is between those of humans (0.163) and mice (0.116), indicating an intermediate level of purifying selection pressure in the pig. Genes showing increased dN/dS ratios in each lineage were analysed using DAVID8 to examine whether these rapidly evolving genes were enriched for specific biological processes. Most lineages show different fast-evolving pathways, but some pathways are shared (Fig. 1).
KEGG pathways with genes that show accelerated evolution for each of the six mammals used in the dN/dS analysis. The bar charts show the individual dN/dS and dS values for each of the six mammals. The dN/dS and dS values refer to the time period of each of the six individual lineages. The number of proteins that show significantly accelerated dN/dS ratios in each lineage varies from 84 in the mouse to 311 in the pig lineage. Pathways significantly (P < 0.05) enriched within this group of genes are also shown with the number of genes shown in brackets. HPI, Helicobacter pylori infection.
Immune genes are known to be actively evolving in mammals9,10. Because many immune genes were not included in the analysis of 1:1 orthologues, we examined a randomly selected subset of 158 immunity-related pig proteins for evidence of accelerated evolution (Supplementary information). Twenty-seven of these genes (17%) demonstrated accelerated evolution (Supplementary Table 8). A parallel analysis of 143 human and 145 bovine orthologues revealed very similar rates of evolution (18% in human and 12% in cattle, respectively). Using a branch-site analysis, we detected accelerated evolution of amino acids in PRSS12, CD1D and TRAF3 specific to pig (positive selection on pig branch), as well as amino acids in TREM1, IL1B and SCARA5 specific to pig and cow (positive selection on the cetartiodactyl branch).
Further analysis of porcine immune genes (Supplementary Table 5) revealed evidence for specific gene duplications and gene-family expansions (Supplementary Tables 6 and 7). The analysis of this second cetartiodactyl genome indicates that some expansions are cetartiodactyl-specific (cathelicidin) whereas others are ruminant/bovine-specific (β-defensins, C-type lyzozymes) or potentially porcine-specific (type I interferon, δ subfamily).
Pigs have at least 39 type I interferon (IFN) genes, which is twice the number identified in humans and significantly more than in mice. We also detected 16 pseudogenes in this family. Cattle have 51 type I IFNs (13 pseudogenes), indicating that both bovine and porcine type I IFN families have undergone expansion. This is particularly important for interferon subtypes δ (IFND), ω (IFNW) and τ (IFNT); pigs and cattle are evolving species-specific subtypes of IFND and IFNT, respectively. Both species are expanding the IFNW family and share many more IFNW isoforms than other species. Thus, expansion of interferon genes is not ruminant-specific as proposed earlier10, although duplication within some specific sub-families seems to be either bovine- or porcine-specific.
Within the immunity-related genes annotated, we found evidence for duplication of six immune-related genes: IL1B, CD36, CD68, CD163, CRP and IFIT1, and one non-immune gene, RDH16. The CD36 gene is also duplicated in the bovine genome, whereas the IL1B gene duplication, where evidence for a partial duplication was reported previously11, is unique in mammals. Other key immune genes in the major histocompatibility complex, immunoglobulin, T-cell-receptor and natural killer cell receptor loci have been characterized in detail12,13,14,15,16,17,18,19 (Supplementary Information).
Another significant porcine genome expansion is the olfactory receptor gene family. We identified 1,301 porcine olfactory receptor genes and 343 partial olfactory receptor genes20. The fraction of pseudogenes within these olfactory receptor sequences (14%) is the lowest observed in any species so far. This large number of functional olfactory receptor genes most probably reflects the strong reliance of pigs on their sense of smell while scavenging for food.
Conservation of synteny and evolutionary breakpoints
Alignment of the porcine genome against seven other mammalian genomes (Supplementary Information) identified homologous synteny blocks (HSBs). Using porcine HSBs and stringent filtering criteria, 192 pig-specific evolutionary breakpoint regions (EBRs) were located. The number of porcine EBRs (146, Supplementary Table 11 and Supplementary Fig. 16) is comparable to the number of bovine-lineage-specific EBRs (100) reported earlier using a slightly lower resolution (500 kilobases (kb)), indicating that both lineages evolved with an average rate of ∼2.1 large-scale rearrangements per million years after the divergence from a common cetartiodactyl ancestor ∼60 Myr ago2. This rate compares to ∼1.9 rearrangements per million years within the primate lineage (Supplementary Table 11). A total of 20 and 18 cetartiodactyl EBRs (shared by pigs and cattle) were detected using the pig and human genomes as a reference, respectively.
Pig-specific EBRs were enriched for LTR endogenous retrovirus 1 (LTR-ERV1) transposons and satellite repeats (Supplementary Table 12), indicating that these two families of repetitive sequences have contributed to chromosomal evolution in the pig lineage. Different families of transposable elements seem to have been active in the cetartiodactyl ancestor. The cetartiodactyl EBRs are enriched for LINE1 elements and tRNAGlu-derived SINEs. tRNAGlu-derived SINEs, previously found over-represented in cetartiodactyl EBRs defined in the bovine genome10, originated in the common ancestor of cetartiodactyls21. Our observation that these elements are also enriched in porcine EBRs strongly supports the hypothesis that active transposable elements promote lineage-specific genomic rearrangements.
A stringent set of porcine to human one-to-one orthologues using the MetaCore database revealed that porcine EBRs and adjacent intervals are enriched for genes involved in sensory perception of taste (P < 8.9 × 10−6; FDR <0.05), indicating that taste phenotypes may have been affected by events associated with genomic rearrangements. Pigs have a limited ability to taste NaCl22. SCNN1B, a gene encoding a sodium channel involved in the perception of salty tastes, is located in a porcine-specific EBR. Another gene, ITPR3, encoding a receptor for inositol triphosphate and a calcium channel involved in the perception of umami and sweet tastes, has been affected by the insertion of several porcine-specific SINE mobile elements into its 3′ untranslated region (3′ UTR), consistent with our observation of a higher density of transposable elements in EBRs. In addition to 8 bitter taste receptor genes annotated by Ensembl and which were used in the gene enrichment analysis, we identified 9 intact genes, to give a total number of 17 TAS2R receptors in the pig (Supplementary Table 13). This compares to 18 intact bitter taste receptors in cattle, 19 in horse, 15 in dog and 25 in humans23,24. Of the 14 bitter taste receptor genes that were mapped to a specific pig chromosome (SSC), 10 were found near 2 EBRs on SSC5 and SSC18 (Supplementary Tables 13 and 15). We also found that at least four taste receptors (TAS1R2, TAS2R1, TAS2R40 and TAS2R39) have been under relaxed selection (Supplementary Information). Pigs are not sensitive to bitter tastes and tolerate higher concentrations of bitter compounds than humans22,25. Thus, pigs can eat food that is unpalatable to humans. A review of the porcine taste transduction network (Supplementary Fig. 17) revealed additional genes affected by rearrangements that affect ‘apical and taste receptor cell’ processes. Together with the observed over-representation of genes related to ‘adrenergic receptor activity’ and ‘angiotensin and other binding’ categories in the pig EBRs (Supplementary Fig. 18), our data indicate that chromosomal rearrangements significantly contributed to adaptation in the suid lineage.
Population divergence and domestication
Divergence between Asian and European wild boar
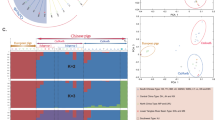
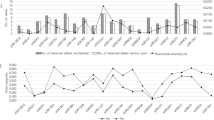
We investigated the evolution within Sus scrofa in Eurasia by sequencing ten individual unrelated wild boars from different geographical areas. In total, 17,210,760 single nucleotide polymorphisms (SNPs) were identified among these ten wild boars. The number of SNPs segregating in the four Asian wild boars (11,472,192) was much higher than that observed in the six European wild boars (6,407,224) with only 2,212,288 shared SNPs. This higher nucleotide diversity was visible in the distribution of heterozygous sites of the Asian compared to the European wild boar genomes (Fig. 2). Phylogenomic analyses of complete genome sequences from these wild boars and six domestic pigs revealed distinct Asian and European lineages (Supplementary Fig. 23) that split during the mid-Pleistocene 1.6–0.8 Myr ago (Calabrian stage, Frantz, L. A. F. et al., manuscript submitted). Colder climates during the Calabrian glacial intervals probably triggered isolation of populations across Eurasia. Admixture analyses (Supplementary Information) within Eurasian Sus scrofa disclosed gene flow between the northern Chinese and European populations consistent with pig migration across Eurasia, between Europe and northern China, throughout the Pleistocene. Our demographic analysis on the whole-genome sequences of European and Asian wild boars (Fig. 3) revealed an increase in the European population after pigs arrived from China. During the Last Glacial Maximum (LGM; ∼20,000 years ago)26, however, Asian and European populations both suffered population bottlenecks. The drop in population size was more pronounced in Europe than Asia (Fig. 3), suggesting a greater impact of the LGM in northern European regions and probably resulting in the observed lower genetic diversity in modern European wild boar.
Shown is the distribution of the heterozygosity as the log2(SNPs) per 10k bin. a, Wild Sus scrofa: blue, south China; green, north China; orange, Italian; red, Dutch. b, Breeds: blue, Chinese breeds (Jiangquhai, Meishan, Xiang); red–yellow, European breeds (Hampshire, large white, landrace). Note that the Hampshire breed is a North American breed of European origin.
Demographic history was inferred using a hidden Markov model (HMM) approach as implemented in pairwise sequentially Markovian coalescence (PSMC)45. In the absence of known mutation rates for pig, we used the default mutation rate for human (μ) of 2.5 × 10−8. For the generation time (g) we used an estimate of 5 years. The Last Glacial Maximum (LGM) is highlighted in grey. WBnl, wild boar Netherlands; WBit, wild boar Italy; WBNch, wild boar north China; WBSch, wild boar south China.
The deep phylogenetic split between European and Asian wild boars is further supported by our observation of 1,272,737 fixed differences between the six European and four Asian wild boars, 1,706 of which are non-synonymous mutations in 1,191 different genes. Genes involved in sensory perception, immunity and host defence were among the most rapidly evolving genes (Supplementary Table 28), further strengthening the conclusions from our analysis of immunity-related pig proteins. This conclusion is further supported by our observation that these genes are also enriched in porcine segmental duplications (Supplementary Information).
To investigate further whether specific regions in the genome of European and Asian wild boar have been under positive selection, a selective sweep analysis was performed on the ten wild boar genome sequences using an approach similar to that recently described in the comparison of Neanderthal and Homo sapiens genomes27. Regions in the genome under strong positive selection after the divergence of these two populations are expected to share fewer derived alleles. Using stringent criteria (Supplementary Information), we identified a total of 251 putative selective sweep regions, with an average size of 111,269 base pairs (bp), together comprising around 1% of the genome and harbouring 365 annotated protein-coding genes (Supplementary Table 26). Many of these regions (112) do not contain any currently annotated protein-coding exons. In contrast, the 10 putative selective sweep regions located between positions 39–43 Mb on SSC3 together harbour 93 genes. This SSC3 region (Supplementary Fig. 25) is directly adjacent to the centromere and exhibits low recombination rates28. Low recombining regions have been shown to be more prone to selective sweeps and meiotic drive29,30. Although similar large putative selective sweep regions close to the centromere were only observed on SSC6 between positions 56.2–57.5 Mb, on most chromosomes selective sweep regions tended to cluster in the central part of chromosomes, thus exhibiting a clear correlation with regions of low recombination (Supplementary Fig. 27). As expected, regions with the highest nucleotide differentiation between European and Asian wild boars were observed in high recombination regions towards the end of the chromosomes on both metacentric and acrocentric chromosomes28.
The putative selective sweep regions displayed significant over-representation of genes involved in RNA splicing and RNA processing, indicating possible changes in the regulation of genes at the level of RNA processing (Supplementary Table 27). Several of these genes (CELF1, CELF6, WDR83, RBM39, RBM6, HNRNPA1, HNRNPM) are involved in alternative splicing, and, small differences in expression might affect alternatively spliced transcripts of specific genes. Evolution of regulatory splicing factors such as heterogeneous ribonucleoprotein particle (hnRNP) proteins has been proposed as an evolutionary model for alternative splicing31, and genetic variation in such factors can affect alternative splicing and result in different phenotypes or disease32. Our observation that specific genes involved in splicing show accelerated evolution in the pig lineage (Fig. 1) supports this hypothesis. Of particular interest is the selective sweep region observed at position 26 Mb on SSC3 around the ERI2 gene (Fig. 4), which encodes ERI1 exoribonuclease family member 2. Different gene variants have been fixed in European and Asian wild boar coding for proteins that differ at two amino acid positions: Cys52Arg and His358Leu encoded by exons 3 and 9 of the ERI2 gene, respectively. The precise function of ERI2 is unknown but the ERI1 exoribonuclease family members have been shown to be involved in mRNA decay33 and in Caenorhabditis elegans ERI-1 has been shown to be involved in the degradation of microRNAs (miRNAs)34.
The y axis shows the log-transformed value of the ratio for the observed/expected derived allele frequency using a sliding window at a bin size of 50,000 bp. The x axis shows the position on SSC3 in base pairs.
Independent domestication and admixture events in domestic breeds
A phylogenetic tree constructed using four European wild boar and domestic pigs and six East Asian wild boar and domestic pigs revealed a clear distinction between European and Asian breeds, thus substantiating the hypothesis that pigs were independently domesticated in western Eurasia and East Asia3. An admixture analysis revealed European influence in Asian breeds, and a ∼35% Asian fraction in European breeds (Supplementary Table 24). These results are consistent with the known exchange of genetic material between European and Asian pig breeds35. We also observed that European breeds form a paraphyletic clade, which cannot be solely explained by varying degrees of Asian admixture (Supplementary Information). Within each continent, our analysis revealed different degrees of relatedness between breeds and their respective wild relatives (Supplementary Table 20).
During domestication, pigs were often allowed to roam in a semi-managed state and recurrent admixture between wild and domesticated individuals was not uncommon, especially in Europe35. Thus, the most likely explanation for the paraphyletic pattern seen in domestic individuals is a long history of genetic exchange between wild and domestic pigs.
The pig as a biomedical model
The pig is an important biomedical model and the ability to generate transgenics and knockouts in combination with somatic nuclear cloning procedures has resulted in a number of models for specific human diseases36. Naturally occurring mutations also offer opportunities to use pigs as biomedical models37,38. To explore the potential for natural models further, predicted porcine protein sequences were compared with their human orthologues. We observed 112 positions where the porcine protein has the same amino acid that is implicated in a human disease (Supplementary Table 29). Most of these changes in humans have been shown to increase risk in multifactorial traits such as obesity (ADRB3, SDC3) and diabetes (PPP1RA, SLC30A8, ZNF615) or shown to result in relatively mild phenotypes (for example, dyslexia: KIAA0319) or late-onset diseases such as Parkinson’s disease (LRRK2, SNCA) and Alzheimer’s disease (TUBD1, BLMH, CEP192, PLAU). These porcine variants are of interest, as they will allow detailed characterization in an experimental model organism whose physiology is very similar to that of human.
Among 32,548 non-synonymous mutations identified by sequencing 48 individual pigs, representing 8 different European and Asian breeds and wild boars39, 6 protein variants implicated in human disease were identified (Supplementary Table 30). In addition, another 157 nonsense mutations in 142 genes were identified, 11 of which have also been implicated in human disease (Supplementary Table 31). Most of these 11 variants were only observed in a heterozygous state and those for which homozygous individuals were observed probably result in either a mild phenotype (ASS1, mild form of citrullinaemia in humans) or in phenotypes unlikely to affect the fitness of wild boars (RBBP8, pancreatic carcinomas). Our estimate for the average number of nonsense mutations per individual (∼30) is smaller than that observed in humans40 despite the observed threefold higher nucleotide diversity in pigs39. This is in agreement with the higher effective population size in the pig compared to that for the human population, which exhibited a strong bottleneck followed by an exponential increase in size during recent history41.
When considering pig-to-human xenotransplantation, porcine endogenous retroviruses (PERVs) pose a risk of zoonotic infection. The pig genome contains fewer endogenous retroviruses than many vertebrates, including humans and mice, and most PERVs were characterized as defective. However, the potential risk posed by re-activation of rare replication-competent PERVs and defective PERVs by recombination remains, as shown for murine ERVs (XMRV)42. Most PERVs consist of γ and γ-like groups (68%), with β-retroviral ERVs comprising the second most abundant group (Supplementary Fig. 15). Our phylogenetic study shows a particularly close relationship between the most intact γ1 group of PERVs (γ1) and murine γ-ERVs, suggesting a potential recent instance of murine-to-porcine transmission of γ1 ERVs (Supplementary Fig. 15). We identified 20 almost intact PERV γ1 loci (Supplementary Table 10), none of which contained a complete set of gag, pol or env open reading frames, indicating that these proviruses are not replicable. We also identified four β-retroviral PERVs, each containing defects, primarily in env. These were distantly related to intracisternal type A particle (IAP) proviruses of mice and the mouse mammary tumour virus (MMTV)-like (HML) proviruses of humans. None of the above loci was shared in more than 120 pigs tested, indicating considerable PERV polymorphisms.
Conclusion
The draft pig genome sequence reported here has illuminated the evolution of Sus scrofa and confirmed its speciation in South East Asia and subsequent domestication at multiple regions across Eurasia. The high-quality annotated reference genome sequence has already proven to be a critical framework for comparing individual genomes39,43,44 and its value is further illustrated in associated papers published elsewhere (http://www.biomedcentral.com/series/swine). The genome sequence also provides a valuable resource enabling effective uses of pigs both in agricultural production and in biomedical research.
Methods Summary
Assembly
We constructed a hybrid de novo assembly based primarily on sequences from BAC clones sequenced clone-by-clone and supplemented with Illumina whole-genome shotgun (WGS) reads. BAC clones were selected from the high-resolution physical (BAC contig) map6 with CHORI-242 library clones prepared from DNA from a single Duroc sow (Duroc 2-14) chosen preferentially. The WGS sequence data were generated using DNA isolated from the same animal. The BAC-derived sequence data were assembled into sequence contigs using Phrap on a clone-by-clone basis and subsequently independently assembled WGS contigs (Supplementary Information) were used to extend BAC clone-derived sequence contigs and to close gaps between clone-derived contigs. Further details and other methods are described in Supplementary Information.
Accession codes
Primary accessions
European Nucleotide Archive
GenBank/EMBL/DDBJ
Data deposits
The final assembly (Sscrofa10.2) has been deposited in the public sequence databases (GenBank/EMBL/DDBJ) under accession number AEMK01000000. The primary source of the Sscrofa10.2 assembly is the NCBI ftp site (ftp://ftp.ncbi.nih.gov/genbank/genomes/Eukaryotes/vertebrates_mammals/Sus_scrofa/Sscrofa10.2/). The chromosomes are CM000812–CM00830 and CM001155. They are built from 5,343 placed scaffolds, with GenBank accession numbers GL878569-GL882503 and JH114391-JH118402. The 4,562 unplaced scaffolds of Sscrofa10.2 have accessions in the ranges GL892100–GL896682 and JH118403–JH118999. Illumina sequences for the sequenced wild boars and individuals of the other breeds, aligned against build10.2, have been deposited in the European Nucleotide Archive (ENA) under project number ERP001813.
References
Kumar, S. & Hedges, S. B. A molecular timescale for vertebrate evolution. Nature 392, 917–920 (1998)
Meredith, R. W. et al. Impacts of the Cretaceous Terrestrial Revolution and KPg extinction on mammal diversification. Science 334, 521–524 (2011)
Larson, G. et al. Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307, 1618–1621 (2005)
Schook, L. B. et al. Swine Genome Sequencing Consortium (SGSC): a strategic roadmap for sequencing the pig genome. Comp. Funct. Genomics 6, 251–255 (2005)
Archibald, A. L. et al. Pig genome sequence – analysis and publication strategy. BMC Genomics 11, 438 (2010)
Humphray, S. J. et al. A high utility integrated map of the pig genome. Genome Biol. 8, R139 (2007)
Flicek, P. et al. Ensembl 2012. Nucleic Acids Res. 40, D84–D90 (2012)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Barreiro, L. B. & Quintana-Murci, L. From evolutionary genetics to human immunology: how selection shapes host defense genes. Nature Rev. Genet. 11, 17–30 (2010)
Bovine Genome Sequencing and Analysis Consortium. The genome sequence of taurine cattle: a window to ruminant biology and evolution. Science 324, 522–528 (2009)
Vandenbroeck, K. et al. Gene sequence, cDNA construction, expression in Escherichia coli and genetically approached purification of porcine interleukin-1 beta. Eur. J. Biochem. 217, 45–52 (1993)
Renard, C. et al. The genomic sequence and analysis of the swine major histocompatibility complex. Genomics 88, 96–110 (2006)
Tanaka-Matsuda, M., Ando, A., Rogel-Gaillard, C., Chardon, P. & Uenishi, H. Difference in number of loci of swine leukocyte antigen classical class I genes among haplotypes. Genomics 93, 261–273 (2009)
Schwartz, J. C., Lefranc, M. P. & Murtaugh, M. P. Evolution of the porcine (Sus scrofa domestica) immunoglobulin κ locus through germline gene conversion. Immunogenetics 64, 303–311 (2012)
Schwartz, J. C., Lefranc, M. P. & Murtaugh, M. P. Organization, complexity and allelic diversity of the porcine (Sus scrofa domestica) immunoglobulin lambda locus. Immunogenetics 64, 399–407 (2012)
Uenishi, H. et al. Genomic structure around joining segments and constant regions of swine T-cell receptor α/δ (TRA/TRD) locus. Immunology 109, 515–526 (2003)
Uenishi, H. et al. Genomic sequence encoding diversity segments of the pig TCR δ chain gene demonstrates productivity of highly diversified repertoire. Mol. Immunol. 46, 1212–1221 (2009)
Eguchi-Ogawa, T., Toki, D. & Uenishi, H. Genomic structure of the whole D-J-C clusters and the upstream region coding V segments of the TRB locus in pig. Dev. Comp. Immunol. 33, 1111–1119 (2009)
Sambrook, J. G. et al. Identification of a single killer immunoglobulin-like receptor (KIR) gene in the porcine leukocyte receptor complex on chromosome 6q. Immunogenetics 58, 481–486 (2006)
Nguyen, D. T. et al. The complete swine olfactory subgenome: expansion of olfactory receptor gene repertoire in the pig genome. BMC Genomics (in the press)
Shimamura, M., Abe, H., Nikaido, M., Ohshima, K. & Okada, N. Genealogy of families of SINEs in cetaceans and artiodactyls: the presence of a huge superfamily of tRNA(Glu)-derived families of SINEs. Mol. Biol. Evol. 16, 1046–1060 (1999)
Hellekant, G. & Danilova, V. Taste in domestic pig, Sus scrofa. J. Anim. Physiol. Anim. Nutr. (Berl.) 82, 8–24 (1999)
Fischer, A., Gilad, Y., Man, O. & Pääbo, S. Evolution of bitter taste receptors in humans and apes. Mol. Biol. Evol. 22, 432–436 (2005)
Dong, D., Jones, G. & Zhang, S. Dynamic evolution of bitter taste receptor genes in vertebrates. BMC Evol. Biol. 9, 12 (2009)
Nelson, S. L. & Sanregret, J. D. Response of pigs to bitter-tasting compounds. Chem. Senses 22, 129–132 (1997)
Yokoyama, Y., Lambeck, K., De Deckker, P., Johnston, P. & Fifield, L. K. Timing of the Last Glacial Maximum. Nature 406, 713–716 (2000)
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Tortereau, F. et al. Sex specific differences in recombination rate in the pig are correlated with GC content. BMC Genomics (in the press)
Barton, N. H. Genetic hitchhiking. Phil. Trans. R. Soc. Lond. B 355, 1553–1562 (2000)
Lyttle, T. W. Cheaters sometimes prosper: distortion of mendelian segregation by meiotic drive. Trends Genet. 9, 205–210 (1993)
Ast, G. How did alternative splicing evolve? Nature Rev. Genet. 5, 773–782 (2004)
Garcia-Blanco, M. A., Baraniak, A. P. & Lasda, E. L. Alternative splicing in disease and therapy. Nature Biotechnol. 22, 535–546 (2004)
Kupsco, J. M., Wu, M.-J., Marzluff, W. F., Thapar, R. & Duronio, R. J. Genetic and biochemical characterization of Drosophila Snipper: A promiscuous member of the metazoan 3′hExo/ERI-1 family of 3′ to 5′ exonucleases. RNA 12, 2103–2117 (2006)
Kennedy, S., Wang, D. & Ruvkun, G. A conserved siRNA-degrading RNase negatively regulates RNA interference in C. elegans. Nature 427, 645–649 (2004)
White, S. From globalized pig breeds to capitalist pigs: a study in animals cultures and evolutionary history. Environ. Hist. 16, 94–120 (2011)
Walters, E. M. et al. Completion of the swine genome will simplify the production of swine as a large animal biomedical model. BMC Med. Genomics (in the press)
Gillard, E. F. et al. A substitution of cysteine for arginine 614 in the ryanodine receptor is potentially causative of human malignant hyperthermia. Genomics 11, 751–755 (1991)
Murgiano, L., Tammen, I., Harlizius, B. & Drögemüller, C. A de novo germline mutation in MYH7 causes a progressive dominant myopathy in pigs. BMC Genomics (in the press)
Bosse, M. et al. Regions of homozygosity in the porcine genome: Consequence of demography and the recombination landscape. PLoS Genet. 8, e1003100 (2012)
MacArthur, D. G. et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 335, 823–828 (2012)
Keinan, A. & Clark, A. G. Recent explosive human population growth has resulted in an excess of rare variants. Science 336, 740–743 (2012)
Paprotka, T. et al. Recombinant origin of the retrovirus XMRV. Science 333, 97–101 (2011)
Fang, X. et al. The sequence and analysis of an inbred pig genome. GigaScience (in the press)
Rubin, C. J. et al. Strong signatures of selection in the domestic pig genome. Proc. Natl. Acad. Sci. USA (in the press)
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011)
Acknowledgements
The authors recognize the contributions of the following individuals towards the establishment of the Swine Genome Sequencing Consortium and their leadership in realizing this effort: J. Jen, P. J. Burfening, D. Hamernik, R. A. Easter, N. Merchen, R. D. Green, J. Cassady, B. Harlizius, M. Boggess and M. Stratton. Also the authors acknowledge A. Hernandez, C. Wright at the University of Illinois Keck Center for Comparative and Functional Genomics; N. Bruneau and Prof. Ning Li for their contribution to PERV studies; D. Goodband and D. Berman for their efforts in genome annotation; D. Grafham of the Welcome Trust Sanger Institute for his efforts in the genome assembly and J. Hedegaard, M. Nielsen and R. O. Nielsen for their contribution on the miRNA analysis. We also recognize contributions from the National Institute of Agrobiological Sciences and the Institute of Japan Association for Techno-innovation in Agriculture, Forestry and Fisheries, Tsukuba, Japan, H. Shinkai, T. Eguchi-Ogawa, K. Suzuki, D. Toki, T. Matsumoto, N. Fujishima-Kanaya, A. Mikawa, N. Okumura, M. Tanaka-Matsuda, K. Kurita, H. Sasaki, K. Kamiya, A. Kikuta, T. Bito and N. Fujitsuka. We acknowledge support from the USDA CSREES/NIFA Swine Genome Coordination Program, College of Agricultural, Consumer and Environmental Sciences, University of Illinois; College of Agriculture and Life Sciences, Iowa State University; North Carolina Agricultural Research Service; USA National Pork Board; Iowa Pork Producers Association; North Carolina Pork Council; Danish government; TOPIGS Research Center IPG The Netherlands; INRA Genescope, France; Wellcome Trust Sanger Institute and BGI. We are grateful to the genome team at NCBI for their assistance in checking the Sscrofa10.2 assembly and for their independent annotation of the sequence. This project was also partially supported by grants: BBSRC grants (Ensembl): BB/E010520/1, BB/E010520/2, BB/I025328/1; EC FP6 ‘Cutting edge genomics for sustainable animal breeding (SABRE)’; EC FP7 ‘Quantomics’; C. J. Martin Overseas Based Biomedical Fellowship from the Australian NHMRC (575585); BBSRC (BB/H005935/1); Next-Generation BioGreen 21 Program (PJ009019, PJ0081162012), RDA, Republic of Korea; Consolider programme from Ministry of Research (Spain); NIH R13 RR020283A; NIH R13 RR032267A; ILLU 535-314; ILLU 538-379; ILLU-538-312; ILLU-538-34; CSREES, NIFA for funding genome coordination activities; NIH grant 5 P41LM006252; MAFF grants (IRPPIAUGT-AG 1101/1201); USDA-NRI-2009-35205-05192; USDA-NRSP8 Bioinformatics Coordination and Pig Genome Coordination funds; US-UK Fulbright Commission; Next-Generation BioGreen 21 (no. PJ0080892011) Program, RDA, Republic of Korea; USDA-ARS Project Plan 1235-51000-055-00D; USDA-ARS Project Plan 1265-32000-098-00D; USDA-NRI-2006-35204-17337 USDA AFRI NIFA/DHS 2010-39559-21860; NIH P20-RR017686; USDA ARS; USDA-NRSP8 Bioinformatics; USDA ARS Beltsville Area Summer Undergraduate Fellowships; BBSRC grant BB/G004013/1; NSFC Outstanding Youth grant (31025026); The Swedish Research Council FORMAS; The Swedish Wenner-Gren Foundations; European Commission FP6 funded project LSHB-CT-2006-037377; BioGreen21, RDA grant PJ00622901; BioGreen21, RDA grant PJ00622902; BioGreen21, RDA grant PJ00622903; BioGreen21, RDA grant PJ00622903. The research leading to these results has received funding from the European Research Council under the European Community’s Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement no. 249894 (SelSweep); NIH P20-RR017686; NIH NIDA P30 DA018310; NIH NIDA R21 DA027548; NIAS, RDA grant PJ001758; BioGreen21, RDA grant PJ006229; NIAS, RDA grant 20040301034467; ANR grant ANR07-GANI-001 DeliSus; Danish funding agencies: FTP/DFF (09-066598); DSF/Strategic Growth Technologies (09-067036); the Lundbeck foundation (374/06); DCSC (Scientific Computing); The Funds for International Cooperation from the Ministry of Science and Technology of China 2002AA229061; PL-Grid project: POIG.02.03.00-00-007/08-00 ‘Genome Assembly’; USDA-NIFA-CREES AG 2006-35216-16668; AG 2002-34480-11828; AG 2003-34480-13172; AG 2004-34480-14417; AG 2005-34480-15939; AG 2006-34480-17150; AG 2008-34480-19328; AG 2009-34480-19875; AG 2002-35205-12712; somatic cell genomics: Integrating QTL Discovery and Validation; AG 2008-35205-18769; AG 2009-65205-05642; AG 2004-3881-02193; AG 2011-67015-30229; AG 58-5438-2-313; AG 58-5438-7-317; and AG 58-0208-7-149; NIH grant 5 P41 LM006252.
Author information
Authors and Affiliations
Contributions
Manuscript main text: A.L.A., M.A.M.G., L.B.S., H.U., C.K.T., Y.T., M.F.R., C.P., S.L., D.M., H.-J.M., D.M.L., H.Ki., L.A.F.F., G.L.M.C.; project coordination: A.L.A., M.A.M.G., L.B.S., M.F.R., D.M., J.R., C.Chu., H.U., M.C., K.E.; project initiation: A.L.A., M.A.M.G., L.B.S., M.F.R., D.M., M.F., C.W.B., P.C., G.A.R., M.Y., J.R., L.B.; library preparation and sequencing: S.J.H., C.S., C.Cl., S.M., L.M., M.J., Y.Lu, X.X., P.N., Jia.Z., G.Z., A.L.A., R.C.C., T.M., H.Ka., K.-T.L., T.-H.K., H.-S.P., E.-W.P., J.-H.K., S.-H.C., S.-J.O., Ji.W., Ju.W., J.-T.J.; genome assembly: A.L.A., M.C., S.L., C.S., P.D., H.-J.M., H.U., D.M., B.S., T.F., Y.Li, N.D., R.R.-G., R.L., K.H., W.C.; repetitive DNA analysis: G.J.F. (leader), J.J., F.DeS., H.-J.M.; gene content and genome evolution: S.F., B.L.A., S.W., S.S.; conservation of synteny and evolutionary breakpoints: D.M.L. (leader), J.N., L.A., B.C., H.A.L., J.M., J.K., D.K.G., K.E.F.; speciation: L.A.F.F., M.A.M.G., O.M., H.-J.M., J.G.S.; divergence of Asian and European wild boar: H.-J.M., M.Bo., M.A.M.G., L.A.F.F.; annotation: S.S., B.L.A., T.M., C.K.T., Y.S., M.By., R.C., J.R., E.F., Z.-L.H., W.L., M.P.-E.; RNA analysis: O.M., R.P.M.A.C., H.U., C.A., H.T., B.T., P.S., M.F., J.G., C.B., F.P., H.H., Z.B., J.F.; neuropeptides: J.V.S., B.R.S., S.R.-Z.; pig domestication: L.A.F.F., R.P.M.A.C., H.-J.M., M.Bo., S.O., G.L., L.R., J.G.S.; population admixture: L.A.F.F., J.G.S.; biomedical models: B.D., L.R., K.S., M.A.M.G.; immune response: C.K.T., (co-leader) C.R.-G. (co-leader), H.D.D., J.E.L., A.A., B.B., J.S., D.B., F.B., M.By., S.B., C.Che., D.C.-S., R.C., E.F., E.G., J.G.R.G., J.L.H., T.H., Z.-L.H., R.K., J.K.L., K.M., M.P.M., T.M., G.P., J.M.R., J.S., H.U., Jie Z., S.Z.; olfactory and taste receptor analysis: C.P. (leader), D.T.N., K.L.; dN/dS analysis: H.Ki. (leader), H.A., K.-W.K.; PERV and retroviral insertions: C.R.-G., A.H., P.J., J.B., G.S., L.S., R.W., Y.T. (leader); segmental duplications: O.M., Y.P., Z.-Q.D., M.F.R.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text 1-11, Supplementary Figures 1-28, Supplementary Tables 1-7, 9-15, 18-25, 27-28, 30 (see separate files for Supplementary Tables 8, 16, 17, 26, 29 and 31) and additional references.This file was replaced on 23 January 2013 as figure 2 had corrupted in the original version posted online. (PDF 5571 kb)
Supplementary Table 8
This file contains Supplementary Table 8. (XLS 463 kb)
Supplementary Table 16
This file contains Supplementary Table 16. (XLS 154 kb)
Supplementary Table 17
This file contains Supplementary Table 17. (XLS 70 kb)
Supplementary Table 26
This file contains Supplementary Table 26. (XLS 58 kb)
Supplementary Table 29
This file contains Supplementary Table 29. (XLS 55 kb)
Supplementary Table 31
This file contains Supplementary Table 31. (XLS 116 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Groenen, M., Archibald, A., Uenishi, H. et al. Analyses of pig genomes provide insight into porcine demography and evolution. Nature 491, 393–398 (2012). https://doi.org/10.1038/nature11622
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11622
This article is cited by
-
Mapping and functional characterization of structural variation in 1060 pig genomes
Genome Biology (2024)
-
Genetic introgression from commercial European pigs to the indigenous Chinese Lijiang breed and associated changes in phenotypes
Genetics Selection Evolution (2024)
-
The pig as an optimal animal model for cardiovascular research
Lab Animal (2024)
-
Population genetics reveals new introgression in the nucleus herd of min pigs
Genes & Genomics (2024)
-
Genome-wide scan for runs of homozygosity in South American Camelids
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.