Abstract
Unravelling the genomic landscape of divergence between lineages is key to understanding speciation1. The naturally hybridizing collared flycatcher and pied flycatcher are important avian speciation models2,3,4,5,6,7 that show pre- as well as postzygotic isolation8,9. We sequenced and assembled the 1.1-Gb flycatcher genome, physically mapped the assembly to chromosomes using a low-density linkage map10 and re-sequenced population samples of each species. Here we show that the genomic landscape of species differentiation is highly heterogeneous with approximately 50 ‘divergence islands’ showing up to 50-fold higher sequence divergence than the genomic background. These non-randomly distributed islands, with between one and three regions of elevated divergence per chromosome irrespective of chromosome size, are characterized by reduced levels of nucleotide diversity, skewed allele-frequency spectra, elevated levels of linkage disequilibrium and reduced proportions of shared polymorphisms in both species, indicative of parallel episodes of selection. Proximity of divergence peaks to genomic regions resistant to sequence assembly, potentially including centromeres and telomeres, indicate that complex repeat structures may drive species divergence. A much higher background level of species divergence of the Z chromosome, and a lower proportion of shared polymorphisms, indicate that sex chromosomes and autosomes are at different stages of speciation. This study provides a roadmap to the emerging field of speciation genomics.
Similar content being viewed by others
Main
As lineages diverge, a combination of pre- as well as postzygotic reproductive isolation barriers will eventually arise1. Divergence is likely to start from specific loci that may precede and cause the evolution of reproductive incompatibility. Hybridization between diverging lineages may therefore create a genomic mosaic of regions where interspecific gene flow occurs at different rates (the genic view of speciation11), with introgression expected to be weak in genomic regions involved in speciation. Revealing the genomic regions with elevated levels of divergence will eventually deepen our knowledge of the speciation process. However, more than 150 years after the publication of On the Origin of Species12, the genetic basis of speciation is still largely unresolved13,14. We know little about the identity, number and effect size of loci involved in population divergence, their genomic distribution and the type of mutations involved. Advances in sequencing technology now open a promising avenue for the study of genomic divergence, even for non-model vertebrate species with gigabase (Gb)-sized genomes.
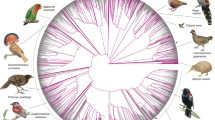
The collared flycatcher Ficedula albicollis and the pied flycatcher Ficedula hypoleuca (Fig. 1) are important study organisms for key aspects of evolutionary ecology and biology2,3,4,5,6,7. Diverged less than 2 million years ago, their history has been shaped by repeated cycles of glaciation in Eurasia where periods of allopatric divergence in refugia probably alternated with periods of secondary contact during which gene flow and selection were vital components; they still hybridize in areas of sympatry (Supplementary Figure 2). To study the genetic basis of species divergence in this system, we sequenced and assembled the flycatcher genome, and physically placed, ordered and oriented sequence scaffolds along chromosomes through linkage-map data. This was followed by re-sequencing of genomes and transcriptomes of population samples of both species (Supplementary Methods, Supplementary Fig. 1), allowing base-pair (bp)-resolution of the pattern of differentiation on a genomic level and providing a roadmap for studies in the emerging field of speciation genomics.
a, Male collared flycatcher. b, Male pied flycatcher. Note that the male collared flycatcher has a white neck collar and a more pronounced white forehead and wing patches. Photographs courtesy of Johan Träff.
The final assembly encompassed 1.13 Gb with an N50 scaffold size of 7.3 Mb and with 89% of the assembly contained within no more than 200 scaffolds larger than 1 Mb (Supplementary Tables 1–7). The sequenced bird was heterozygous at 3.66 million positions, corresponding to an average of one segregating site every 330 bp. A low-density linkage map of collared flycatcher10 anchors 73% of the assembly to chromosomes and orients scaffolds along them (Supplementary Fig. 3). Based on conserved chromosomal organization between flycatcher and zebra finch (Supplementary Fig. 4) we were able to anchor and orient additional scaffolds, thereby physically positioning 1.00 Gb of the assembly (89%) in the genome (Supplementary Fig. 5). This illustrates that physical assembly of Gb-sized genomes sequenced with short reads is possible with modest linkage information and when assisted with genome information from a related species. The flycatcher genome contained 18,735 predicted protein-coding genes, of which 18,649 (>99.5%) were identified as expressed based on RNA–seq data from a variety of tissues.
We then sequenced the genomes of 10 unrelated males of each species (mean coverage 5.69x ± 2.01 s.d.; Supplementary Table 8) and found 9.86- and 7.13-million segregating sites in collared flycatchers and pied flycatchers, respectively. The fact that 3.81 million of these single nucleotide polymorphisms (SNPs) (53.4% and 38.6% of the total in each species, respectively) were shared between species confirms their close genetic relationship and provides an unusual access to genomic data of two species before complete lineage sorting. The mean pairwise nucleotide difference in interspecific comparisons (dxy) for 50-kb windows was 0.0046 ± 0.0011, which was only marginally higher than the mean pairwise nucleotide diversity (π) in intraspecific comparisons (πcoll: 0.0036 ± 0.0010; πpied: 0.0021 ± 0.00076); individuals of the two species are thus genetically not much more different from each other than are individuals within species. Acknowledging that population samples of 10 individuals from each species provide low power for detecting rare alleles, an indication of species divergence can be obtained by noting at how many sites all collared flycatchers were homozygous for one allele and all pied flycatchers homozygous for another (which we refer to as sites of fixed differences, df). We found 239,745 such sites, which corresponds to 1 fixed difference every 4.7 kb. Of these, 1,513 sites were located within protein-coding regions and it is realistic that a proportion of these positions represent the genetic basis for key phenotypic differences between the two species.
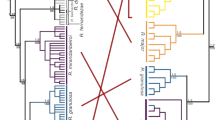
The genomic landscape of species divergence was highly heterogeneous, with a fraction of windows showing highly elevated divergence up to 50 times higher than the genomic median (0.00013) and mean (0.00022) (Supplementary Fig. 6). The distribution of autosomal windows with elevated divergence showed a non-random pattern with approximately 50 well-defined clusters of high FST (the fixation index, a measure of population differentiation) and df (‘divergence peaks’ or ‘genomic islands of divergence’) (Fig. 2a and Table 1). Average peak size was in the range of several hundred kb (median, 400 kb; mean, 625 kb; range, ∼100 kb to 3 Mb; Supplementary Fig. 7) and, in total, divergence peaks covered 2.7% of the genome yet containing 25% of all fixed differences. An immediate feature of the distribution of these 50 peaks was that they were non-randomly distributed across the genome (Kolmogorov–Smirnov test, D = 0.4516, P = 0.0002), irrespective of chromosome size and despite substantial heterogeneity in chromosome size, there were in most cases one to three peaks per chromosome (Fig. 2a). Moreover, peaks were highly overrepresented in the very end of chromosomes and six microchromosomes had peaks in both ends. Another feature of divergence islands was that they lie at the end of scaffolds, thereby not forming a continuous and symmetric signal in the assembly. As a consequence, peaks in the end of chromosomes were generally ‘one-tailed’, whereas peaks within the interior of chromosomes were usually formed by adjacent scaffolds with maximum divergence juxtaposed to the assembly gap between scaffolds (Fig. 2b and Supplementary Fig. 8). This peculiar pattern raises the issue of whether peaks are artefacts associated with the scaffolding process or read mapping. However, several observations convincingly argue against this (Supplementary Notes).
a, Distribution of divergence measured as the density of fixed differences per bp for 200-kb windows across the genome. Chromosomes are listed in numerical order and are separated by gaps. Red horizontal bars show the approximate location of centromeres in homologous chromosomes of zebra finch. Open read symbols are used to indicate that avian microchromosomes are generally acro- or telocentric; both ends of these chromosomes are labelled as the orientation is not known. For chromosomes 4, 6 and 8, there is a lack of an in situ mapped marker 5′ of the centromere in zebra finch. b, Distribution of population genomic parameters along an example chromosome (chromosome 4A). The plots show the density of fixed differences per bp (df) (yellow), FST (red), the total between-species sequence divergence (dxy), nucleotide diversity (π) for each species, the proportion of shared polymorphisms among sites polymorphic in at least one species (purple), the proportion of private polymorphisms among sites polymorphic within species (private and shared polymorphisms shown in the same panel), Tajima’s D, and linkage disequilibrium (r2). For π, private polymorphisms, D and r2, species-specific estimates are given for collared flycatcher in blue and for pied flycatcher in green. Assigned scaffolds are shown under the plot: black, denoting scaffolds ordered and oriented by the collared flycatcher linkage map; grey, scaffolds ordered with the collared flycatcher linkage map and oriented through comparative mapping with zebra finch; white, scaffolds ordered and oriented through comparative mapping with zebra finch.
If selection has driven population divergence in regions of high differentiation, we might expect to see reduced levels of within-species diversity in these regions in one of the species. Species-specific estimates of π showed that this was essentially always the case (Fig. 2b, Supplementary Fig. 8), with mean π in divergence islands less than one-third of the genomic background level (Table 1). As FST by its nature is a function of within-species diversity15, this association might be trivial. However, because the vast majority of divergence islands were seen with FST as well as df, regions of high divergence were characterized both by a high frequency of sequence differences between the two species and a low frequency of sequence differences within species. A noteworthy consequence of these coinciding features was that dxy did not exceed background levels in divergence islands (Fig. 2b and Supplementary Fig. 8). Further indication of selection in regions of high divergence was given by the observations of allele-frequency spectra being skewed towards rare alleles and strong signals of linkage disequilibrium (Table 1, Fig. 2b and Supplementary Notes).
How the abovementioned signs of selection are distributed between the two species is relevant for interpretation of the heterogeneous genomic landscape of species divergence. For almost all regions of elevated divergence, both species showed reduced nucleotide diversity (Fig. 2b and Supplementary Fig. 8). This cannot be explained by a loss of diversity in the ancestral population because it would not lead to the observed high incidence of fixed differences. Moreover, taking dS (the synonymous substitution rate) as a proxy for mutation rate, we found that regions of high divergence were not low in variability because of low mutation rate (generalized linear model, including chromosome length, peak versus non-peak; z = 0.598, P = 0.550). The association of high divergence with low diversity speaks further against mapping artefacts, which can increase divergence between species but should also lead to elevated diversity within species. Taken together, these results suggest that selection has acted to reduce genetic variability in the very same regions in the two lineages independently.
Genomic regions with reduced levels of interspecific recombination are hindered from gene flow, facilitating the build-up of reproductive incompatibilities. Moreover, divergent selection may enhance differentiation over larger genomic regions when the intraspecific recombination rate is low (divergence hitchhiking16,17). To assess the relationship between recombination and divergence, we estimated the population recombination rate (ρ, which equals Ner, where Ne is the effective population size and r is the per-generation recombination rate), ρ/π (as reduced diversity, Ne, within divergence islands will lower the estimates of ρ and contribute to differences in ρ between divergence peaks and the genomic background even if r would be similar) and used genetic distances from the collared flycatcher linkage map related to the physical distance between markers according to the genome assembly to assess the relationship between recombination and divergence (Supplementary Methods and Supplementary Table 9). These tests provided no strong evidence of reduced recombination rate in the proximity of divergence islands (Supplementary Notes).
The flycatcher karyotype has not been established, thus the location of centromeres is not known. However, avian microchromosomes are generally acro- or telocentric18. An attempt to approximate the location of centromeres on flycatcher macrochromosomes was made using information on the location of centromeres in the karyotype of zebra finch, coupled with the high degree of flycatcher–zebra-finch synteny conservation10 (Supplementary Fig. 4). This reveals considerable overlap between the presumed location of flycatcher centromeres and divergence islands (Fig. 2a). This is particularly apparent when considering the enrichment of divergence islands at the ends of microchromosomes. Moreover, the fact that several microchromosomes showed divergence peaks in both ends further suggests that at least some telomeric regions are highly differentiated between species.
Limited pedigree data from multiple generations of flycatcher hybrid descendants demonstrates fitness reduction and suggests that the current rate of introgression is low9. Nevertheless, genetic data from a few loci have previously indicated detectable levels of ongoing gene flow8. To further address this issue we carried out deep sequencing of 24 intronic regions spread across the genome in sympatric population samples of the 2 species (Supplementary Methods and Supplementary Table 10). Assuming an isolation–migration model, the maximum likelihood estimate of the rate of gene flow from pied flycatcher to collared flycatcher was 1.7 × 10−6 per gene and generation (90% posterior density distribution 0.1–2.8 × 10−6), while the rate for the opposite direction was much lower (4.5 × 10−9). From the analysis of nested models we can reject a model without gene flow from pied flycatcher to collared flycatcher (likelihood ratio test, P < 0.01) and estimate the rate at Nem = 0.38 (where m is the migration rate), or roughly one migrant every three generations.
If divergence islands are involved in reproductive isolation, they might be expected to be shielded from gene flow in areas of sympatry, while other genome regions may get introgressed. One way of addressing this possibility is to study the distribution of private and shared polymorphisms to infer differential rates of lineage sorting, such as those caused by variation in gene flow across the genome. We found a very clear pattern at the point at which the proportion of shared polymorphisms drops significantly in all divergence islands, from a mean background level of 32.8% to 18.3% in islands (Wilcoxon test, W = 632,244, P ≪ 0.001; Fig. 2b and Supplementary Fig. 8). This observation indicates more advanced lineage sorting within than outside regions of elevated divergence, and is consistent with a role for gene flow in homogenizing background levels of divergence. Moreover, we found that the proportion of private polymorphisms was significantly higher in divergence islands than elsewhere in the genome (pied flycatcher: 35.3% in non-islands versus 56.1% in islands, W = 319,712, P ≪ 0.001; collared flycatcher: 60.2% in non- islands vs. 75.4% in islands, W = 608,994, P ≪ 0.001; Fig. 2b and Supplementary Fig. 8), consistent with restricted gene flow in islands. Furthermore, the genomic background level of the proportion of private polymorphisms was considerably higher in collared flycatcher than in pied flycatcher (W = 19,001,201, P ≪ 0.001, Fig. 2b and Supplementary Fig. 8), in agreement with the direction of gene flow recorded.
Birds have female heterogamety (males, ZZ; females, ZW) and as all sequencing was carried out using male birds, read coverage is expected to be similar for autosomes and the Z chromosome. Estimates of diversity and divergence should therefore be directly comparable between chromosome categories. The Z chromosome showed greater than sevenfold higher mean divergence (df , 0.0016 ± 0.00060) than autosomes (0.00022 ± 0.00036, W = 23,706,977, P < 2 × 10−16) and significantly higher FST (0.623 ± 0.076 versus 0.350 ± 0.110, W = 23,274,298, P < 2 × 10−16) (Fig. 3). Divergence was more uniformly distributed along the Z chromosome and did not show the distinct islands of divergence characterizing most autosomes; the Z chromosome contained approximately 35% of all fixed sites in the genome. Moreover, estimates of π, Tajima’s D and r2 (an estimate of linkage disequilibrium) were also more uniform along the Z chromosome (Fig. 3 and Supplementary Fig. 8). Despite the higher mean divergence, we note that divergence at individual windows on the Z chromosome did not exceed that within autosomal divergence islands. High sex-linked divergence is thus a consequence of increased background level rather than more extreme divergence in individual regions. Reports of a disproportionately large effect of the X chromosome on hybrid sterility and of reduced introgression of sex-linked genes19 have fed the idea that sex chromosomes are particularly important for the build-up of reproductive isolation. This is supported by data from female heterogametic organisms20, including higher FST seen for a handful of Z-linked markers than for autosomal markers in flycatchers21. In flycatchers, mating patterns also suggest sex-linkage of male plumage traits and species recognition5,21, traits that may evolve under the influence of divergent selection. Our observations could be taken to suggest that natural or sexual selection at multiple loci associated with reproductive isolation on the Z chromosome has erased the signal from individual divergence islands by broadly increasing divergence to a higher background level. From this perspective, the Z chromosome could be seen to represent a more advanced stage of species differentiation, with islands turning into plateaus or divergence hitchhiking turning into genome hitchhiking22. We note in this context that total sequence differentiation in interspecific comparisons was higher for the Z chromosome (mean dxy = 0.0057 ± 0.0011) than for autosomes (0.0045 ± 0.0010; W = 19,256,489, P < 2 × 10−16). Moreover, more advanced divergence of the Z chromosome compared to autosomes was also supported by a significantly lower proportion of shared polymorphisms in the former (15.2%) than among the latter (32.3%; W = 1,408,692, P < < 0.001) (Fig. 3).
The plots show the density of fixed differences per kb (df), FST, nucleotide diversity (π; collared flycatcher in blue and pied flycatcher in green) and the proportion of shared polymorphisms.
The 50 regions defined as divergence peaks contained a total of 530 protein-coding genes. An assessment of gene ontology among these genes did not reveal any functional category to be significantly over-represented (Supplementary Table 11). Moreover, we found no indication that proteins encoded by genes within divergence peaks would be faster evolving than other proteins in the genome (general linearized model of dN/dS, where dN is the rate of non-synonymous substitution, including chromosome length; peak versus non-peak; z = 0.837, P = 0.403). As many peak regions contained more than one gene it is possible that unrelated features of linked genes blur characteristics common to genes under selection. However, we found that genes differentially expressed between species were significantly more common in peak regions (246 out of 346 genes; 71.1%) than in the rest of the genome (4,180 out of 7,134 genes, 58.6%; χ2 = 11.2, P < 10−3; Supplementary Table 12). One possible explanation to this observation is that standing variation at cis-acting regulatory elements in an ancestral population has segregated via linkage to loci under divergent selection in peak regions.
The collared flycatcher and the pied flycatcher probably started to diverge in allopatry in glacial refugia of the Mediterranean area during the Pleistocene epoch, candidate regions being the Iberian and Apennine peninsulas, respectively. Subsequent secondary contact during repeated cycles of interglacial periods allowed gene flow, just as hybridization and gene flow occurs in contemporary areas of sympatry. According to this scenario, allopatric divergence may have been followed repeatedly by genomic homogenization in sympatry. The highly heterogeneous nature of genomic divergence between the two species is compatible with such a model, with some genomic regions refractory to gene flow. Our data show that these regions are localized, numerous, diverged far beyond the background level and present on almost all chromosomes, essentially shedding light on several of the central questions on the genomic landscape of species divergence. The consistent observation in both flycatcher species of reduced diversity in divergence islands would suggest that the same loci, or closely linked loci, have been subject to directional selection in both lineages independently. This, together with the juxtaposition of the most extreme divergence and gaps in the genome assembly, raise the possibility that centromeres or other heterochromatic repeats themselves actually are drivers of species divergence. The meiotic drive model of speciation invokes an arms race between centromeric alleles for deposition into the single resultant oocyte of female meiosis23, in which selection acts on allelic variation in the ability to attract microtubuli of an asymmetric spindle pole. This could lead to rapid evolution of repeat sequences as well as of proteins involved with spindle-fibre attachment to centromeres, possibilities that are supported by empirical data24, and may hinder proper chromosome segregation or pairing during hybrid meiosis25. Interestingly, the Drosophila Zhr locus causing female lethality is itself a heterochromatic satellite-DNA block26, and the Drosophila ods-site homeobox (OdsH) speciation gene has been linked to hybrid male sterility through its binding to evolutionary labile heterochromatic repeats27. Similar actions are suggested for other speciation genes13. There is evidence for segregation distortion in chicken chromosomes involving loci with centromeric or telomeric locations28. The genomic distribution of divergence peaks in flycatchers is compatible with an involvement of telomeres as well, as some chromosomes showed divergence signals in both ends. Telomeres have been shown to have an evolutionary conserved role during meiosis in which they cluster on the nuclear envelope, forming a ‘telomere bouquet’, and enable chromosome movements to promote homologous synapsis29. It is noteworthy in this respect that birds have more extensive arrays of telomeric repeats than other vertebrates, and show structural polymorphism of telomeres within species30, setting the stage for a meiotic drive also in these types of repeats. As meiotic drives are characterized by repeated episodes of selection, this would be compatible with the relative large size of divergence islands observed.
To conclude, this study presents the genome sequence of an avian speciation model and unravels the genomic landscape of species divergence in unprecedented detail. The results show strong heterogeneity in sequence differentiation in a species pair in which lineage sorting is incomplete. The potential connection of species divergence to key repetitive elements of chromosomes calls for a shift in focus, with the quest for genetic basis of reproductive isolation extended to include sequences other than protein-coding genes. For further dissection of the mechanism driving species divergence in this and other systems it will be important to obtain detailed maps of how the rate of recombination varies along chromosomes, based on large-scale genotyping in pedigrees. Together with modelling (under varying intensity and character of selection), this can address to what extent sweeps or background selection, possibly aided by low recombination, are expected to increase divergence and over what distances. As the size of genomic islands of divergence will also be affected by variation in Ne and the rate of migration, these are also factors that need to be integrated in models. Moreover, extensive genotyping in pedigrees would be a means to test for segregation distortion introduced by meiotic drives.
Methods Summary
Genome sequencing was carried out with Illumina technology using DNA from a single wild-caught male collared flycatcher. Sequences from paired-end and mate-pair reads of multiple libraries (200–21,000 bp) were assembled using SOAPDENOVO in subsequent steps with increasing insert size of libraries. Scaffolds were physically anchored to chromosomes by the aid of a collared-flycatcher linkage map and with comparative map information from the zebra-finch genome. Protein-coding genes of the flycatcher genome were retrieved through a combination of mapping reads to zebra-finch gene templates, using flycatcher expressed sequence tag (EST) evidence and ab initio prediction. Levels of gene expression were measured across a suite of tissues (embryonic, adult somatic and gonadal tissues) using RNA–seq with Illumina technology, and differentially expressed genes were identified with BAYSEQ. Population genomic analyses were based on data from re-sequencing of 10 individuals each of collared flycatcher and pied flycatcher, in which reads were mapped to the assembly using BWA (Burrows-Wheeler Aligner) software. After analysis and data processing using a combination of software tools, sequence variants were identified with the Genome Analysis Toolkit (GATK; Broad Institute). Divergence and diversity parameters were estimated using ‘haploidized’ data by randomly choosing one allele from heterozygous genotypes.
Accession codes
Data deposits
The flycatcher genome sequence and associated data is available under the accession numbers AGTO00100000 (GenBank) and ERP001377 (Short Read Archive).
References
Coyne, J. A. & Orr, H. A. Speciation. (Sinauer Associates, 2004)
Ellegren, H., Gustafsson, L. & Sheldon, B. C. Sex ratio adjustment in relation to paternal attractiveness in a wild bird population. Proc. Natl Acad. Sci. USA 93, 11723–11728 (1996)
Saetre, G.-P. et al. A sexually selected character displacement in flycatchers reinforces premating isolation. Nature 387, 589–592 (1997)
Qvarnström, A., Part, T. & Sheldon, B. C. Adaptive plasticity in mate preference linked to differences in reproductive effort. Nature 405, 344–347 (2000)
Veen, T. et al. Hybridization and adaptive mate choice in flycatchers. Nature 411, 45–50 (2001)
Merilä, J., Kruuk, L. E. B. & Sheldon, B. C. Cryptic evolution in a wild bird population. Nature 412, 76–79 (2001)
Saether, S. A. et al. Sex chromosome-linked species recognition and evolution of reproductive isolation in flycatchers. Science 318, 95–97 (2007)
Borge, T., Lindroos, K., Nadvornik, P., Syvanen, A. C. & Saetre, G. P. Amount of introgression in flycatcher hybrid zones reflects regional differences in pre and post-zygotic barriers to gene exchange. J. Evol. Biol. 18, 1416–1424 (2005)
Wiley, C., Qvarnstrom, A., Andersson, G., Borge, T. & Saetre, G. P. Postzygotic isolation over multiple generations of hybrid descendens in a natural hybrid zone: how well do single-generation estimates reflect reproductive isolation? Evolution 63, 1731–1739 (2009)
Backström, N. et al. A gene-based genetic linkage map of the collared flycatcher (Ficedula albicollis) reveals extensive synteny and gene-order conservation during 100 million years of avian evolution. Genetics 179, 1479–1495 (2008)
Wu, C.-I. The genic view of the process of speciation. J. Evol. Biol. 14, 851–865 (2001)
Darwin, C. On the Origin of Species by Means of Natural Selection, or the Preservation of Favoured Races in the Struggle for Life. (John Murray, 1859)
Presgraves, D. C. The molecular evolutionary basis of species formation. Nature Rev. Genet. 11, 175–180 (2010)
Nosil, P. & Schluter, D. The genes underlying the process of speciation. Trends Ecol. Evol. 26, 160–167 (2011)
Charlesworth, B. Measures of divergence between populations and the effect of forces that reduce variability. Mol. Biol. Evol. 15, 538–543 (1998)
Feder, J. L. & Nosil, P. The efficacy of divergence hitchhiking in generating genomic islands during ecological speciation. Evolution 64, 1729–1747 (2010)
Via, S. & West, J. The genetic mosaic suggests a new role for hitchhiking in ecological speciation. Mol. Ecol. 17, 4334–4345 (2008)
Shields, G. F. Comparative avian cytogenetics: a review. Condor 84, 45–58 (1982)
Coyne, J. A. Genetics and speciation. Nature 355, 511–515 (1992)
Jiggins, C. D. et al. Sex-linked hybrid sterility in a butterfly. Evolution 55, 1631–1638 (2001)
Saetre, G.-P. et al. Sex chromosome evolution and speciation in Ficedula flycatchers. Proc. R. Soc. Lond. B 270, 53–59 (2003)
Nosil, P. & Feder, J. L. Genomic divergence during speciation: causes and consequences. Phil. Trans. R. Soc. B 367, 332–342 (2012)
Henikoff, S. & Malik, H. S. Centromeres: selfish drivers. Nature 417, 227 (2002)
Malik, H. S. & Henikoff, S. Major evolutionary transitions in centromere complexity. Cell 138, 1067–1082 (2009)
Fishman, L. & Saunders, A. Centromere-associated female meiotic drive entails male fitness costs in monkeyflowers. Science 322, 1559–1562 (2008)
Ferree, P. M. & Barbash, D. A. Species-specific heterochromatin prevents mitotic chromosome segregation to cause hybrid lethality in Drosophila. PLoS Biol. 7, e1000234 (2009)
Bayes, J. J. & Malik, H. S. Altered heterochromatin binding by a hybrid sterility protein in Drosophila sibling species. Science 326, 1538–1541 (2009)
Axelsson, E. et al. Segregation distortion in chicken and the evolutionary consequences of female meiotic drive in birds. Heredity 105, 290–298 (2010)
Tsai, J.-H. & McKee, B. D. Homologous pairing and the role of pairing centers in meiosis. J. Cell Sci. 124, 1955–1963 (2011)
Delany, M. E., Gessaro, T. M., Rodrigue, K. L. & Daniels, L. M. Chromosomal mapping of chicken mega-telomere arrays to GGA9, 16, 28 and W using a cytogenomic approach. Cytogenet. Genome Res. 117, 54–63 (2007)
Acknowledgements
Financial support was obtained from a European Research Council Advanced Investigator Grant (NEXTGENMOLECOL), a Knut and Alice Wallenberg Scholar Grant, and from the Swedish Research Council to H.E. R.B. was funded by the Swiss National Science Foundation (grants PBLAB3-134299 and PBLAB1-140171). We are grateful to M. Lascoux, M. Noor and T. Slotte for helpful discussion and comments. We thank the Uppsala University SNP and SEQ Technology Platform for help with DNA sequencing, and the Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX), and the associated Next generation sequencing Cluster and Storage (UPPNEX) project, funded by the Knut and Alice Wallenberg Foundation and the Swedish National Infrastructure for Computing (SNIC), for computer resources.
Author information
Authors and Affiliations
Contributions
L.S., P.I.O. and A.K. carried out the bioinformatic analyses; R.B. and T.K. performed population genomic analyses and interpreted the data; H.E., N.B., K.N.-B. and J.B.W.W. interpreted the data; S.U. performed analyses of differential gene expression; H.M. generated the RNA–seq data; N.B. collected and processed the samples; A.Q. facilitated sampling of collared flycatchers; H.E., L.S., R.B., P.I.O., T.K., A.K. and S.U. wrote the Supplementary Information, with input from the other authors; H.E. conceived and designed the study, supervised the project, and wrote the main paper with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11, Supplementary Methods, Supplementary Notes, Supplementary Tables 1-12 and Supplementary References. (PDF 12916 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Ellegren, H., Smeds, L., Burri, R. et al. The genomic landscape of species divergence in Ficedula flycatchers. Nature 491, 756–760 (2012). https://doi.org/10.1038/nature11584
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11584
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.