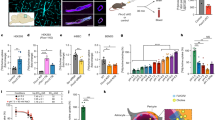
Abstract
Defects in the availability of haem substrates or the catalytic activity of the terminal enzyme in haem biosynthesis, ferrochelatase (Fech), impair haem synthesis and thus cause human congenital anaemias1,2. The interdependent functions of regulators of mitochondrial homeostasis and enzymes responsible for haem synthesis are largely unknown. To investigate this we used zebrafish genetic screens and cloned mitochondrial ATPase inhibitory factor 1 (atpif1) from a zebrafish mutant with profound anaemia, pinotage (pnt tq209 ). Here we describe a direct mechanism establishing that Atpif1 regulates the catalytic efficiency of vertebrate Fech to synthesize haem. The loss of Atpif1 impairs haemoglobin synthesis in zebrafish, mouse and human haematopoietic models as a consequence of diminished Fech activity and elevated mitochondrial pH. To understand the relationship between mitochondrial pH, redox potential, [2Fe–2S] clusters and Fech activity, we used genetic complementation studies of Fech constructs with or without [2Fe–2S] clusters in pnt, as well as pharmacological agents modulating mitochondrial pH and redox potential. The presence of [2Fe–2S] cluster renders vertebrate Fech vulnerable to perturbations in Atpif1-regulated mitochondrial pH and redox potential. Therefore, Atpif1 deficiency reduces the efficiency of vertebrate Fech to synthesize haem, resulting in anaemia. The identification of mitochondrial Atpif1 as a regulator of haem synthesis advances our understanding of the mechanisms regulating mitochondrial haem homeostasis and red blood cell development. An ATPIF1 deficiency may contribute to important human diseases, such as congenital sideroblastic anaemias and mitochondriopathies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schultz, I. J., Chen, C., Paw, B. H. & Hamza, I. Iron and porphyrin trafficking in heme biogenesis. J. Biol. Chem. 285, 26753–26759 (2010)
Iolascon, A., De Falco, L. & Beaumont, C. Molecular basis of inherited microcytic anemia due to defects in iron acquisition or heme synthesis. Haematologica 94, 395–408 (2009)
Severance, S. & Hamza, I. Trafficking of heme and porphyrins in metazoa. Chem. Rev. 109, 4596–4616 (2009)
Anderson, K. E., Sassa, S., Bishop, D. F. & Desnick, R. J. in The Online Metabolic & Molecular Basis of Inherited Disease (ed. Bishop, D. F. ) 1–153 (The McGraw-Hill Press, 2011)
Shin, J. T. & Fishman, M. C. From zebrafish to human: modular medical models. Annu. Rev. Genomics Hum. Genet. 3, 311–340 (2002)
Ransom, D. G. et al. Characterization of zebrafish mutants with defects in embryonic hematopoiesis. Development 123, 311–319 (1996)
Postlethwait, J. H. et al. Vertebrate genome evolution and the zebrafish gene map. Nature Genet. 18, 345–349 (1998)
Campanella, M., Parker, N., Tan, C. H., Hall, A. M. & Duchen, M. R. IF1: setting the pace of the F1F0-ATP synthase. Trends Biochem. Sci. 34, 343–350 (2009)
Ando, C. & Ichikawa, N. Glutamic acid in the inhibitory site of mitochondrial ATPase inhibitor, IF1, participates in pH sensing in both mammals and yeast. J. Biochem. 144, 547–553 (2008)
Shaw, G. C. et al. Mitoferrin is essential for erythroid iron assimilation. Nature 440, 96–100 (2006)
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009)
Campanella, M. et al. Regulation of mitochondrial structure and function by the F1F0-ATPase inhibitor protein, IF1 . Cell Metab. 8, 13–25 (2008)
Richardson, D. R. et al. Mitochondrial iron trafficking and the integration of iron metabolism between the mitochondrion and cytosol. Proc. Natl Acad. Sci. USA 107, 10775–10782 (2010)
Nilsson, R. et al. Discovery of genes essential for heme biosynthesis through large-scale gene expression analysis. Cell Metab. 10, 119–130 (2009)
Walker, J. E. The regulation of catalysis in ATP synthase. Curr. Opin. Struct. Biol. 4, 912–918 (1994)
Lanzilotta, W. N. & Dailey, H. A. Human ferrochelatase. In Handbook of Metalloproteins (ed. Messerschmidt, A. ) Vol. 4/5: 138–146 (John Wiley & Sons, 2011)
Lange, H., Kispal, G. & Lill, R. Mechanism of iron transport to the site of heme synthesis inside yeast mitochondria. J. Biol. Chem. 274, 18989–18996 (1999)
Froschauer, E. M., Schweyen, R. J. & Wiesenberger, G. The yeast mitochondrial carrier proteins Mrs3p/Mrs4p mediate iron transport across the inner mitochondrial membrane. Biochim. Biophys. Acta 1788, 1044–1050 (2009)
Park, D. et al. Continued clearance of apoptotic cells critically depends on the phagocyte Ucp2 protein. Nature 477, 220–224 (2011)
Crooks, D. R., Ghosh, M. C., Haller, R. G., Tong, W. H. & Rouault, T. A. Posttranslational stability of the heme biosynthetic enzyme ferrochelatase is dependent on iron availability and intact iron-sulfur cluster assembly machinery. Blood 115, 860–869 (2010)
Medlock, A. E. & Dailey, H. A. Examination of the activity of carboxyl-terminal chimeric constructs of human and yeast ferrochelatases. Biochemistry 39, 7461–7467 (2000)
Childs, S. et al. Zebrafish dracula encodes ferrochelatase and its mutation provides a model for erythropoietic protoporphyria. Curr. Biol. 10, 1001–1004 (2000)
Hu, J., Dong, L. & Outten, C. E. The redox environment in the mitochondrial intermembrane space is maintained separately from the cytosol and matrix. J. Biol. Chem. 283, 29126–29134 (2008)
Yu, D. et al. miR-451 protects against erythroid oxidant stress by repressing 14–3-3 ζ. Genes Dev. 24, 1620–1633 (2010)
North, T. E. et al. PGE2-regulated wnt signaling and N-acetylcysteine are synergistically hepatoprotective in zebrafish acetaminophen injury. Proc. Natl Acad. Sci. USA 107, 17315–17320 (2010)
Cao, Y. A. et al. Heme oxygenase-1 deletion affects stress erythropoiesis. PLOS One 6, e20634 (2011)
Formentini, L., Sanchez-Arago, M., Sanchez-Cenizo, L. & Cuezva, J. M. The mitochondrial ATPase inhibitory factor 1 triggers a ROS-mediated retrograde prosurvival and proliferative response. Mol. Cell 45, 731–742 (2012)
Sheftel, A. D., Richardson, D. R., Prchal, J. & Ponka, P. Mitochondrial iron metabolism and sideroblastic anemia. Acta Haematol. 122, 120–133 (2009)
Camaschella, C. Hereditary sideroblastic anemias: pathophysiology, diagnosis, and treatment. Semin. Hematol. 46, 371–377 (2009)
Li, L. & Kaplan, J. A mitochondrial-vacuolar signaling pathway in yeast that affects iron and copper metabolism. J. Biol. Chem. 279, 33653–33661 (2004)
Acknowledgements
We thank members of our laboratory (M. Cassim, A. Kaplan, G. Hildick-Smith and H. Anderson) and colleagues (S. L. Alper, K. Pepper and N. S. Trede) for critical review of the manuscript, T. C. Law for pnt adult blood characterization, H. Mulhern for help with the electron microscopy, and Christopher Lawrence and his team for the zebrafish husbandry. This research was supported in part by the Cooley’s Anemia Foundation (D.I.S. and C.C.), the March of Dimes Foundation (B.H.P.), the American Heart Association (J.D.C. and A.E.M.), the Dutch National Science Fund (I.J.S.), the Fondation Soldati pour la Recherche en Cancerologie (G.V.), the Burroughs Welcome Fund (N.N.D.), the NIDDK (D.I.S., B.H.P., A.N., J.K., H.A.D. and S.M.H.) and the NHLBI (D.I.S. and B.H.P).
Author information
Authors and Affiliations
Contributions
D.I.S. and B.H.P. originally conceived the project, designed and performed the experiments, analysed data and wrote the manuscript. N.T.-M., A.S., L.L., D.M.W. and J.K. measured 59Fe uptake in mitochondria and complexed in haem, PPIX levels, Fech activity, xanthine oxidase and aconitase activities and the haem levels in a yeast knockout for Inh1 and participated in scientific discussions. J.D.C., I.J.S., E.L.P. and N.B.L. did zebrafish embryo microinjections. S.K.H., G.V., C.C. and W.C. helped with zebrafish colony maintenance and protein experiments. Y.Z. helped with high resolution meiotic mapping. A.N., S.N.H. and B.L.E. helped with silencing ATPIF1 in human primary CD34+ cells. S.M.H. helped with silencing Atpif1 in MFPL cells. A.E.M., T.A.D. and H.A.D. created Fech constructs for injection, measured Fech activity as a function of pH, analysed Fech structure for [2Fe–2S] clusters and participated in scientific discussions. D.F., J.M.W., M.C. and N.N.D. helped to design and measure mitochondrial physiological parameters. C.B. analysed pnt adult blood parameters. D.R.C. and M.D.F. helped with electron microscopic analysis of mitochondrial structure and disease.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Sequences are available in GenBank/EMBL/DDBJ as follows: zebrafish atpif1a (NM_001089521.1), zebrafish atpif1b (NM_001044859), mouse Atpif1 (NC_000070.5) and human ATPIF1 (NC_005104.2).
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7, Supplementary Methods, a Supplementary Discussion and Supplementary References. (PDF 1249 kb)
Rights and permissions
About this article
Cite this article
Shah, D., Takahashi-Makise, N., Cooney, J. et al. Mitochondrial Atpif1 regulates haem synthesis in developing erythroblasts. Nature 491, 608–612 (2012). https://doi.org/10.1038/nature11536
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11536
This article is cited by
-
ATP synthase inhibitory factor subunit 1 regulates islet β-cell function via repression of mitochondrial homeostasis
Laboratory Investigation (2022)
-
A pathway coordinated by DELE1 relays mitochondrial stress to the cytosol
Nature (2020)
-
Reduction of the ATPase inhibitory factor 1 (IF1) leads to visual impairment in vertebrates
Cell Death & Disease (2018)
-
Knockout of the ATPase inhibitory factor 1 protects the heart from pressure overload-induced cardiac hypertrophy
Scientific Reports (2017)
-
Integrated analysis identified an intestinal-like and a diffuse-like gene sets that predict gastric cancer outcome
Tumor Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.