Abstract
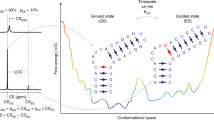
The visualization of RNA conformational changes has provided fundamental insights into how regulatory RNAs carry out their biological functions. The RNA structural transitions that have been characterized so far involve long-lived species that can be captured by structure characterization techniques. Here we report the nuclear magnetic resonance visualization of RNA transitions towards ‘invisible’ excited states (ESs), which exist in too little abundance (2–13%) and for too short a duration (45–250 μs) to allow structural characterization by conventional techniques. Transitions towards ESs result in localized rearrangements in base-pairing that alter building block elements of RNA architecture, including helix–junction–helix motifs and apical loops. The ES can inhibit function by sequestering residues involved in recognition and signalling or promote ATP-independent strand exchange. Thus, RNAs do not adopt a single conformation, but rather exist in rapid equilibrium with alternative ESs, which can be stabilized by cellular cues to affect functional outcomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Palmer, A. G. & Massi, F. Characterization of the dynamics of biomacromolecules using rotating-frame spin relaxation NMR spectroscopy. Chem. Rev. 106, 1700–1719 (2006)
Baldwin, A. J., Hansen, D. F., Vallurupalli, P. & Kay, L. E. Measurement of methyl axis orientations in invisible, excited states of proteins by relaxation dispersion NMR spectroscopy. J. Am. Chem. Soc. 131, 11939–11948 (2009)
Neudecker, P. et al. Structure of an intermediate state in protein folding and aggregation. Science 336, 362–366 (2012)
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007)
Sugase, K., Dyson, H. J. & Wright, P. E. Mechanism of coupled folding and binding of an intrinsically disordered protein. Nature 447, 1021–1025 (2007)
Korzhnev, D. M., Religa, T. L., Banachewicz, W., Fersht, A. R. & Kay, L. E. A transient and low-populated protein-folding intermediate at atomic resolution. Science 329, 1312–1316 (2010)
Li, P., Martins, I. R. S., Amarasinghe, G. K. & Rosen, M. K. Internal dynamics control activation and activity of the autoinhibited Vav DH domain. Nature Struct. Biol. 15, 613–618 (2008)
Boehr, D. D., McElheny, D., Dyson, H. J. & Wright, P. E. The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313, 1638–1642 (2006)
Hansen, A. L., Nikolova, E. N., Casiano-Negroni, A. & Al-Hashimi, H. M. Extending the range of microsecond-to-millisecond chemical exchange detected in labeled and unlabeled nucleic acids by selective carbon R(1rho) NMR spectroscopy. J. Am. Chem. Soc. 131, 3818–3819 (2009)
Massi, F., Johnson, E., Wang, C., Rance, M. & Palmer, A. G. NMR R1ρ rotating-frame relaxation with weak radio frequency fields. J. Am. Chem. Soc. 126, 2247–2256 (2004)
Korzhnev, D. M., Orekhov, V. Y. & Kay, L. E. Off-resonance R1ρ NMR studies of exchange dynamics in proteins with low spin-lock fields: An application to a Fyn SH3 domain. J. Am. Chem. Soc. 127, 713–721 (2005)
Nikolova, E. N. et al. Transient Hoogsteen base pairs in canonical duplex DNA. Nature 470, 498–502 (2011)
Hoogstraten, C. G., Wank, J. R. & Pardi, A. Active site dynamics in the lead-dependent ribozyme. Biochemistry 39, 9951–9958 (2000)
Johnson, J. E. & Hoogstraten, C. G. Extensive backbone dynamics in the GCAA RNA tetraloop analyzed using C-13 NMR spin relaxation and specific isotope labeling. J. Am. Chem. Soc. 130, 16757–16769 (2008)
Blad, H., Reiter, N. J., Abildgaard, F., Markley, J. L. & Butcher, S. E. Dynamics and metal ion binding in the U6 RNA intramolecular stem-loop as analyzed by NMR. J. Mol. Biol. 353, 540–555 (2005)
Dethoff, E. A. et al. Characterizing complex dynamics in the transactivation response element apical loop and motional correlations with the bulge by NMR, molecular dynamics, and mutagenesis. Biophys. J. 95, 3906–3915 (2008)
Bannwarth, S. & Gatignol, A. HIV-1 TAR RNA: the target of molecular interactions between the virus and its host. Curr. HIV Res. 3, 61–71 (2005)
Jaeger, J. A. & Tinoco, I., Jr An NMR study of the HIV-1 TAR element hairpin. Biochemistry 32, 12522–12530 (1993)
Kulinski, T. et al. The apical loop of the HIV-1 TAR RNA hairpin is stabilized by a cross-loop base pair. J. Biol. Chem. 278, 38892–38901 (2003)
Farès, C., Amata, I. & Carlomagno, T. 13C-detection in RNA bases: revealing structure-chemical shift relationships. J. Am. Chem. Soc. 129, 15814–15823 (2007)
Ghose, R., Marino, J. P., Wiberg, K. B. & Prestegard, J. H. Dependence of 13C chemical shifts on glycosidic torsional angles in ribonucleic acids. J. Am. Chem. Soc. 116, 8827–8828 (1994)
Nozinovic, S., Furtig, B., Jonker, H. R., Richter, C. & Schwalbe, H. High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA. Nucleic Acids Res. 38, 683–694 (2010)
Snoussi, K. & Leroy, J.-L. Imino proton exchange and base-pair kinetics in RNA duplexes. Biochemistry 40, 8898–8904 (2001)
Parisien, M. & Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008)
Legault, P. & Pardi, A. Unusual dynamics and pKa shift at the active site of a lead-dependent ribozyme. J. Am. Chem. Soc. 119, 6621–6628 (1997)
Feng, S. & Holland, E. C. HIV-1 tat trans-activation requires the loop sequence within tar. Nature 334, 165–167 (1988)
Berkhout, B. & Jeang, K. T. trans activation of human immunodeficiency virus type 1 is sequence specific for both the single-stranded bulge and loop of the trans-acting-responsive hairpin: a quantitative analysis. J. Virol. 63, 5501–5504 (1989)
Richter, S., Cao, H. & Rana, T. M. Specific HIV-1 TAR RNA loop sequence and functional groups are required for human cyclin T1-Tat-TAR ternary complex formation. Biochemistry 41, 6391–6397 (2002)
Yoshizawa, S., Fourmy, D. & Puglisi, J. Recognition of the codon-anticodon helix by ribosomal RNA. Science 285, 1722–1725 (1999)
Schmeing, T. M. & Ramakrishnan, V. What recent ribosome structures have revealed about the mechanism of translation. Nature 461, 1234–1242 (2009)
Shandrick, S. et al. Monitoring molecular recognition of the ribosomal decoding site. Angew. Chem. Int. Ed. 43, 3177–3182 (2004)
Fourmy, D., Recht, M., Blanchard, S. & Puglisi, J. Structure of the A site of Escherichia coli 16S ribosomal RNA complexed with an aminoglycoside antibiotic. Science 274, 1367–1371 (1996)
Romanowska, J., Setny, P. & Trylska, J. Molecular dynamics study of the ribosomal A-site. J. Phys. Chem. B 112, 15227–15243 (2008)
O’Connor, M., Thomas, C. L., Zimmermann, R. A. & Dahlberg, A. E. Decoding fidelity at the ribosomal A and P sites: influence of mutations in three different regions of the decoding domain in 16S rRNA. Nucleic Acids Res. 25, 1185–1193 (1997)
Dahlquist, K. D. & Puglisi, J. D. Interaction of translation initiation factor IF1 with the E. coli ribosomal A site. J. Mol. Biol. 299, 1–15 (2000)
Kipper, K., Hetényi, C., Sild, S., Remme, J. & Liiv, A. Ribosomal intersubunit bridge B2a is involved in factor-dependent translation initiation and translational processivity. J. Mol. Biol. 385, 405–422 (2009)
Moore, M. D. & Hu, W.-S. HIV-1 RNA dimerization: It takes two to tango. AIDS Rev. 11, 91–102 (2009)
Clever, J. L. & Parslow, T. G. Mutant human immunodeficiency virus type 1 genomes with defects in RNA dimerization or encapsidation. J. Virol. 71, 3407–3414 (1997)
Rist, M. J. & Marino, J. P. Mechanism of nucleocapsid protein catalyzed structural isomerization of the dimerization initiation site of HIV-1. Biochemistry 41, 14762–14770 (2002)
Mujeeb, A. et al. Nucleocapsid protein-mediated maturation of dimer initiation complex of full-length SL1 stemloop of HIV-1: sequence effects and mechanism of RNA refolding. Nucleic Acids Res. 35, 2026–2034 (2007)
Turner, K. B., Hagan, N. A. & Fabris, D. Understanding the isomerization of the HIV-1 dimerization initiation domain by the nucleocapsid protein. J. Mol. Biol. 369, 812–828 (2007)
Takahashi, K. et al. Structural requirement for the two-step dimerization of human immunodeficiency virus type 1 genome. RNA 6, 96–102 (2000)
Sun, X., Zhang, Q. & Al-Hashimi, H. M. Resolving fast and slow motions in the internal loop containing stem-loop 1 of HIV-1 that are modulated by Mg2+ binding: role in the kissing-duplex structural transition. Nucleic Acids Res. 35, 1698–1713 (2007)
Yuan, Y., Kerwood, D. J., Paoletti, A. C., Shubsda, M. F. & Borer, P. N. Stem of SL1 RNA in HIV-1: structure and nucleocapsid protein binding for a 1 x 3 internal loop. Biochemistry 42, 5259–5269 (2003)
Lawrence, D. C., Stover, C. C., Noznitsky, J., Wu, Z. & Summers, M. F. Structure of the intact stem and bulge of HIV-1 Ψ-RNA stem-loop SL1. J. Mol. Biol. 326, 529–542 (2003)
Ulyanov, N. B. NMR structure of the full-length linear dimer of stem-loop-1 RNA in the HIV-1 dimer initiation site. J. Biol. Chem. 281, 16168–16177 (2006)
Breaker, R. R. Prospects for riboswitch discovery and analysis. Mol. Cell 43, 867–879 (2011)
Dethoff, E. A., Chugh, J., Mustoe, A. M. & Al-Hashimi, H. M. Functional complexity and regulation through RNA dynamics. Nature 482, 322–330 (2012)
Fourmy, D., Yoshizawa, S. & Puglisi, J. D. Paromomycin binding induces a local conformational change in the A-site of 16 S rRNA. J. Mol. Biol. 277, 333–345 (1998)
Delaglio, F. et al. Nmrpipe—a multidimensional spectral processing system based on Unix Pipes. J. Biomol. NMR 6, 277–293 (1995)
Spyracopoulos, L. A suite of Mathematica notebooks for the analysis of protein main chain 15N NMR relaxation data. J. Biomol. NMR 36, 215–224 (2006)
Meinhold, D. W. & Wright, P. E. Measurement of protein unfolding/refolding kinetics and structural characterization of hidden intermediates by NMR relaxation dispersion. Proc. Natl Acad. Sci. USA 108, 9078–9083 (2011)
Vallurupalli, P., Bouvignies, G. & Kay, L. E. Increasing the exchange time-scale that can be probed by CPMG relaxation dispersion NMR. J. Phys. Chem. B 115, 14891–14900 (2011)
Acknowledgements
E.A.D., K.P. and J.C contributed equally to this work. We thank members of the Al-Hashimi laboratory for input. We acknowledge the Michigan Economic Development Cooperation and the Michigan Technology Tri-Corridor for the support of the purchase of a 600 MHz spectrometer. K.P. is supported by a postdoctoral Fellowship from the Swedish Research Council (VR-K2011-78PK-21662-0-12). This work was supported by the US National Institutes of Health (R01 AI066975) and by a Rackham Graduate Student Research Grant awarded by the University of Michigan.
Author information
Authors and Affiliations
Contributions
H.M.A., E.A.D., K.P. and J.C. conceived the approaches to structurally characterize RNA ES and wrote the paper. E.A.D. and K.P. performed all experiments and data analyses for HIV TAR and SL1m, respectively. J.C. with assistance from A.C.-N. performed all experiments and data analyses for the A-site.
Corresponding author
Ethics declarations
Competing interests
H.M.A. is an advisor to and holds an ownership interest in Nymirum Inc., which is an RNA-based drug discovery company. The research reported in this article was performed by the University of Michigan faculty and students and was funded by an NIH contract to H.M.A.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-8, Supplementary Tables 1-2, a Supplementary Discussion and additional references. (PDF 7758 kb)
Rights and permissions
About this article
Cite this article
Dethoff, E., Petzold, K., Chugh, J. et al. Visualizing transient low-populated structures of RNA. Nature 491, 724–728 (2012). https://doi.org/10.1038/nature11498
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11498
This article is cited by
-
Dynamic basis for dA•dGTP and dA•d8OGTP misincorporation via Hoogsteen base pairs
Nature Chemical Biology (2023)
-
Rational design of hairpin RNA excited states reveals multi-step transitions
Nature Communications (2022)
-
Chemical shift prediction of RNA imino groups: application toward characterizing RNA excited states
Nature Communications (2021)
-
A quantitative model predicts how m6A reshapes the kinetic landscape of nucleic acid hybridization and conformational transitions
Nature Communications (2021)
-
Mutate-and-chemical-shift-fingerprint (MCSF) to characterize excited states in RNA using NMR spectroscopy
Nature Protocols (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.