Abstract
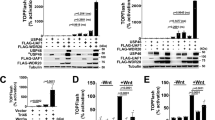
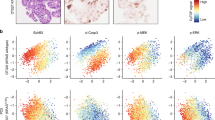
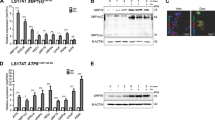
LGR5+ stem cells reside at crypt bottoms, intermingled with Paneth cells that provide Wnt, Notch and epidermal growth factor signals1. Here we find that the related RNF43 and ZNRF3 transmembrane E3 ubiquitin ligases are uniquely expressed in LGR5+ stem cells. Simultaneous deletion of the two genes encoding these proteins in the intestinal epithelium of mice induces rapidly growing adenomas containing high numbers of Paneth and LGR5+ stem cells. In vitro, growth of organoids derived from these adenomas is arrested when Wnt secretion is inhibited, indicating a dependence of the adenoma stem cells on Wnt produced by adenoma Paneth cells. In the HEK293T human cancer cell line, expression of RNF43 blocks Wnt responses and targets surface-expressed frizzled receptors to lysosomes. In the RNF43-mutant colorectal cancer cell line HCT116, reconstitution of RNF43 expression removes its response to exogenous Wnt. We conclude that RNF43 and ZNRF3 reduce Wnt signals by selectively ubiquitinating frizzled receptors, thereby targeting these Wnt receptors for degradation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
29 August 2012
An addition was made to the Acknowledgements.
References
Sato, T. et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011)
Cheng, H. & Leblond, C. P. Origin, differentiation and renewal of the four main epithelial cell types in the mouse small intestine. I. Columnar cell. Am. J. Anat. 141, 461–479 (1974)
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007)
Sato, T. et al. Single Lgr5 stem cells build crypt–villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009)
Yui, S. et al. Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nature Med. 18, 618–623 (2012)
van der Flier, L. G. et al. Transcription factor achaete scute-like 2 controls intestinal stem cell fate. Cell 136, 903–912 (2009)
El Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004)
Ireland, H. et al. Inducible cre-mediated control of gene expression in the murine gastrointestinal tract: effect of loss of β-catenin. Gastroenterology 126, 1236–1246 (2004)
Sansom, O. J. et al. Myc deletion rescues Apc deficiency in the small intestine. Nature 446, 676–679 (2007)
van Es, J. H. et al. Wnt signalling induces maturation of Paneth cells in intestinal crypts. Nature Cell Biol. 7, 381–386 (2005)
Carmon, K. S., Gong, X., Lin, Q., Thomas, A. & Liu, Q. R-spondins function as ligands of the orphan receptors LGR4 and LGR5 to regulate Wnt/β-catenin signaling. Proc. Natl Acad. Sci. USA 108, 11452–11457 (2011)
de Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011)
Kazanskaya, O. et al. The Wnt signaling regulator R-spondin 3 promotes angioblast and vascular development. Development 135, 3655–3664 (2008)
Chen, B. et al. Small molecule-mediated disruption of Wnt-dependent signaling in tissue regeneration and cancer. Nature Chem. Biol. 5, 100–107 (2009)
Koo, B. K. et al. Controlled gene expression in primary Lgr5 organoid cultures. Nature Methods 9, 81–83 (2012)
Liu, G., Bafico, A., Harris, V. K. & Aaronson, S. A. A novel mechanism for Wnt activation of canonical signaling through the LRP6 receptor. Mol. Cell. Biol. 23, 5825–5835 (2003)
Ivanov, I., Lo, K. C., Hawthorn, L., Cowell, J. K. & Ionov, Y. Identifying candidate colon cancer tumor suppressor genes using inhibition of nonsense-mediated mRNA decay in colon cancer cells. Oncogene 26, 2873–2884 (2007)
Li, V. S. et al. Wnt signaling through inhibition of β-catenin degradation in an intact Axin1 complex. Cell 149, 1245–1256 (2012)
Haglund, K. & Dikic, I. The role of ubiquitylation in receptor endocytosis and endosomal sorting. J. Cell Sci. 125, 265–275 (2012)
Mukai, A. et al. Balanced ubiquitylation and deubiquitylation of Frizzled regulate cellular responsiveness to Wg/Wnt. EMBO J. 29, 2114–2125 (2010)
Van der Flier, L. G. et al. The intestinal Wnt/TCF signature. Gastroenterology 132, 628–632 (2007)
Yagyu, R. et al. A novel oncoprotein RNF43 functions in an autocrine manner in colorectal cancer. Int. J. Oncol. 25, 1343–1348 (2004)
Hao, H. X. et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature 485, 195–200 (2012)
Niida, A. et al. DKK1, a negative regulator of Wnt signaling, is a target of the β-catenin/TCF pathway. Oncogene 23, 8520–8526 (2004)
Shimomura, Y. et al. APCDD1 is a novel Wnt inhibitor mutated in hereditary hypotrichosis simplex. Nature 464, 1043–1047 (2010)
Jho, E. H. et al. Wnt/β-catenin/Tcf signaling induces the transcription of Axin2, a negative regulator of the signaling pathway. Mol. Cell. Biol. 22, 1172–1183 (2002)
Berndt, J. D. et al. Mindbomb 1, an E3 ubiquitin ligase, forms a complex with RYK to activate Wnt/β-catenin signaling. J. Cell Biol. 194, 737–750 (2011)
Wu, J. et al. Whole-exome sequencing of neoplastic cysts of the pancreas reveals recurrent mutations in components of ubiquitin-dependent pathways. Proc. Natl Acad. Sci. USA 108, 21188–21193 (2011)
Ong, C. K. e. t. a. l. Exome sequencing of liver fluke-associated cholangiocarcinoma. Nature Genet. 44, 690–693 (2012)
March, H. N. et al. Insertional mutagenesis identifies multiple networks of cooperating genes driving intestinal tumorigenesis. Nature Genet. 43, 1202–1209 (2011)
Tauriello, D. V. et al. Loss of the tumor suppressor CYLD enhances Wnt/β-catenin signaling through K63-linked ubiquitination of Dvl. Mol. Cell 37, 607–619 (2010)
Acknowledgements
We thank A. A. Rolf, I. Kuper, D. V. F. Tauriello, M. van den Born, C. Kroon-Veenboer, H. Begthel, J. Korving and S. van den Brink for technical assistance, L. Lum for providing IWP1 and S. Bartfeld for the schematic drawing. This work was funded in part by grants from the European Research Council, EU/232814-StemCeLLMark and the National Research Foundation of Korea, NRF-2011-357-C00093 (B.-K.K.); EU/Health-F4-2007-200720 (M.v.d.W.); The Centre van Biomedical Genetics (D.E.S.); Ti Pharma/T3-106 (J.H.v.E.); the European Research Council, ERC-StG no.242958 (M.M.M.) and the KNAW/3V-fund.
Author information
Authors and Affiliations
Contributions
B.-K.K., M.M.M. and H.C. conceived and designed the experiments. B.-K.K., M.S., I.J., D.E.S., M.v.d.W. and J.H.v.E. performed the experiments. T.Y.L., S.M. and A.J.R.H. performed the mass spectrometry analysis. B.-K.K., M.S., I.J., M.M.M. and H.C. analysed the data. B.-K.K., M.M.M. and H.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.C. is an inventor on several patents involving the culture system in this paper. The other authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-12. (PDF 10932 kb)
Rights and permissions
About this article
Cite this article
Koo, BK., Spit, M., Jordens, I. et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature 488, 665–669 (2012). https://doi.org/10.1038/nature11308
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11308
This article is cited by
-
Role of the Ror family receptors in Wnt5a signaling
In Vitro Cellular & Developmental Biology - Animal (2024)
-
New insights in ubiquitin-dependent Wnt receptor regulation in tumorigenesis
In Vitro Cellular & Developmental Biology - Animal (2024)
-
Pan-cancer analysis identifies RNF43 as a prognostic, therapeutic and immunological biomarker
European Journal of Medical Research (2023)
-
LGR4 and LGR5 form distinct homodimers that only LGR4 complexes with RNF43/ZNRF3 to provide high affinity binding of R-spondin ligands
Scientific Reports (2023)
-
The USP46 deubiquitylase complex increases Wingless/Wnt signaling strength by stabilizing Arrow/LRP6
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.