Abstract
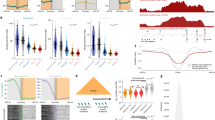
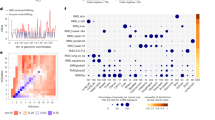
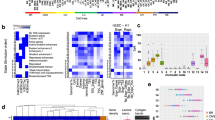
Cancer genome sequencing provides the first direct information on how mutation rates vary across the human genome in somatic cells1,2,3,4,5,6,7. Testing diverse genetic and epigenetic features, here we show that mutation rates in cancer genomes are strikingly related to chromatin organization. Indeed, at the megabase scale, a single feature—levels of the heterochromatin-associated histone modification H3K9me3—can account for more than 40% of mutation-rate variation, and a combination of features can account for more than 55%. The strong association between mutation rates and chromatin organization is upheld in samples from different tissues and for different mutation types. This suggests that the arrangement of the genome into heterochromatin- and euchromatin-like domains is a dominant influence on regional mutation-rate variation in human somatic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ley, T. J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008)
Pleasance, E. D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010)
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010)
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011)
Puente, X. S. et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature 475, 101–105 (2011)
Lee, W. et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 465, 473–477 (2010)
Hodgkinson, A., Chen, Y. & Eyre-Walker, A. The large-scale distribution of somatic mutations in cancer genomes. Hum. Mutat. 33, 136–143 (2012)
Wolfe, K. H., Sharp, P. M. & Li, W. H. Mutation rates differ among regions of the mammalian genome. Nature 337, 283–285 (1989)
Ellegren, H., Smith, N. G. C. & Webster, M. T. Mutation rate variation in the mammalian genome. Curr. Opin. Genet. Dev. 13, 562–568 (2003)
Cooper, D. N. & Krawczak, M. Cytosine methylation and the fate of CpG dinucleotides in vertebrate genomes. Hum. Genet. 83, 181–188 (1989)
Stamatoyannopoulos, J. et al. Human mutation rate associated with DNA replication timing. Nature Genet. 41, 393–395 (2009)
Prendergast, J. G. D. et al. Chromatin structure and evolution in the human genome. BMC Evol. Biol. 7, 72 (2007)
Lercher, M. J. & Hurst, L. D. Human SNP variability and mutation rate are higher in regions of high recombination. Trends Genet. 18, 337–340 (2002)
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010)
Schones, D. E. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Kong, A. et al. A high-resolution recombination map of the human genome. Nature Genet. 31, 241–247 (2002)
Dreszer, T. R. et al. The UCSC Genome Browser database: extensions and updates 2011. Nucleic Acids Res. 40, D918–D923 (2012)
Wang, Z. et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nature Genet. 40, 897–903 (2008)
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007)
Gossmann, T. I., Woolfit, M. & Eyre-Walker, A. Quantifying the variation in the effective population size within a genome. Genetics 189, 1389–1402 (2011)
Vavouri, T. & Lehner, B. Chromatin organization in sperm may be the major functional consequence of base composition variation in the human genome. PLoS Genet. 7, e1002036 (2011)
Peterson, C. L. & Côté, J. Cellular machineries for chromosomal DNA repair. Genes Dev. 18, 602–616 (2004)
Goodarzi, A. A. et al. ATM signaling facilitates repair of DNA double-strand breaks associated with heterochromatin. Mol. Cell 31, 167–177 (2008)
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007)
Wheeler, D. L. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 35, D5–D12 (2007)
A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Paten, B., Herrero, J., Beal, K., Fitzgerald, S. & Birney, E. Enredo and Pecan: genome-wide mammalian consistency-based multiple alignment with paralogs. Genome Res. 18, 1814–1828 (2008)
Zhang, Y., Shin, H., Song, J. S., Lei, Y. & Liu, X. S. Identifying positioned nucleosomes with epigenetic marks in human from ChIP-Seq. BMC Genomics 9, 537 (2008)
Broad Institute Sequencing Platform and Whole Genome Assembly Team et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–481 (2011)
Acknowledgements
This work was funded by an European Research Council (ERC) Starting Grant, European Union Framework 7 project 277899 4DCellFate, ERASysBioPLUS, Ministerio de Ciencia e Innovación (MICINN) grants BFU2008-00365 and BFU2011-26206, Agència de Gestió d’Ajuts Universitaris i de Recerca (AGAUR), the European Molecular Biology Organization (EMBO) Young Investigator Program, the EMBL-CRG Systems Biology Program and a Juan de la Cierva postdoctoral fellowship to B.S-B. We thank T. Vavouri and T. Warnecke for comments on the manuscript, and R.S. Hansen for assistance with analysing replication timing data.
Author information
Authors and Affiliations
Contributions
B.S.-B. performed all analyses. B.S.-B. and B.L. designed analyses and wrote the manuscript. B.L. conceived the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-11. (PDF 3636 kb)
Rights and permissions
About this article
Cite this article
Schuster-Böckler, B., Lehner, B. Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488, 504–507 (2012). https://doi.org/10.1038/nature11273
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11273
This article is cited by
-
Cell cycle gene alterations associate with a redistribution of mutation risk across chromosomal domains in human cancers
Nature Cancer (2024)
-
HJURP is recruited to double-strand break sites and facilitates DNA repair by promoting chromatin reorganization
Oncogene (2024)
-
Sequence dependencies and mutation rates of localized mutational processes in cancer
Genome Medicine (2023)
-
Comprehensive analyses of partially methylated domains and differentially methylated regions in esophageal cancer reveal both cell-type- and cancer-specific epigenetic regulation
Genome Biology (2023)
-
A spatial genome aligner for resolving chromatin architectures from multiplexed DNA FISH
Nature Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.