Abstract
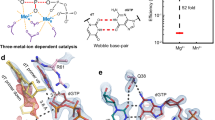
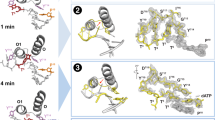
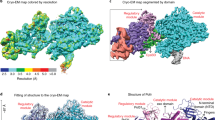
DNA synthesis has been extensively studied, but the chemical reaction itself has not been visualized. Here we follow the course of phosphodiester bond formation using time-resolved X-ray crystallography. Native human DNA polymerase η, DNA and dATP were co-crystallized at pH 6.0 without Mg2+. The polymerization reaction was initiated by exposing crystals to 1 mM Mg2+ at pH 7.0, and stopped by freezing at desired time points for structural analysis. The substrates and two Mg2+ ions are aligned within 40 s, but the bond formation is not evident until 80 s. From 80 to 300 s structures show a mixture of decreasing substrate and increasing product of the nucleotidyl-transfer reaction. Transient electron densities indicate that deprotonation and an accompanying C2′-endo to C3′-endo conversion of the nucleophile 3′-OH are rate limiting. A third Mg2+ ion, which arrives with the new bond and stabilizes the intermediate state, may be an unappreciated feature of the two-metal-ion mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Echols, H. & Goodman, M. F. Fidelity mechanisms in DNA replication. Annu. Rev. Biochem. 60, 477–511 (1991)
Rothwell, P. J. & Waksman, G. Structure and mechanism of DNA polymerases. Adv. Protein Chem. 71, 401–440 (2005)
Brody, R. S. & Frey, P. A. Unambiguous determination of the stereochemistry of nucleotidyl transfer catalyzed by DNA polymerase I from Escherichia coli . Biochemistry 20, 1245–1252 (1981)
Steitz, T. A. & Steitz, J. A. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl Acad. Sci. USA 90, 6498–6502 (1993)
Yang, W., Lee, J. Y. & Nowotny, M. Making and breaking nucleic acids: two-Mg2+-ion catalysis and substrate specificity. Mol. Cell 22, 5–13 (2006)
Johnson, K. A. Role of induced fit in enzyme specificity: a molecular forward/reverse switch. J. Biol. Chem. 283, 26297–26301 (2008)
Warshel, A. et al. Electrostatic basis for enzyme catalysis. Chem. Rev. 106, 3210–3235 (2006)
Pelletier, H., Sawaya, M. R., Kumar, A., Wilson, S. H. & Kraut, J. Structures of ternary complexes of rat DNA polymerase beta, a DNA template-primer, and ddCTP. Science 264, 1891–1903 (1994)
Doublie, S., Tabor, S., Long, A. M., Richardson, C. C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391, 251–258 (1998)
Huang, H., Chopra, R., Verdine, G. L. & Harrison, S. C. Structure of a covalently trapped catalytic complex of HIV-1 reverse transcriptase: implications for drug resistance. Science 282, 1669–1675 (1998)
Li, Y., Korolev, S. & Waksman, G. Crystal structures of open and closed forms of binary and ternary complexes of the large fragment of Thermus aquaticus DNA polymerase I: structural basis for nucleotide incorporation. EMBO J. 17, 7514–7525 (1998)
Franklin, M. C., Wang, J. & Steitz, T. A. Structure of the replicating complex of a pol α family DNA polymerase. Cell 105, 657–667 (2001)
Ling, H., Boudsocq, F., Woodgate, R. & Yang, W. Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication. Cell 107, 91–102 (2001)
Berman, A. J. et al. Structures of phi29 DNA polymerase complexed with substrate: the mechanism of translocation in B-family polymerases. EMBO J. 26, 3494–3505 (2007)
Johnson, S. J., Taylor, J. S. & Beese, L. S. Processive DNA synthesis observed in a polymerase crystal suggests a mechanism for the prevention of frameshift mutations. Proc. Natl Acad. Sci. USA 100, 3895–3900 (2003)
Evans, R. J. et al. Structure of PolC reveals unique DNA binding and fidelity determinants. Proc. Natl Acad. Sci. USA 105, 20695–20700 (2008)
Wang, F. & Yang, W. Structural insight into translesion synthesis by DNA Pol II. Cell 139, 1279–1289 (2009)
Batra, V. K. et al. Magnesium-induced assembly of a complete DNA polymerase catalytic complex. Structure 14, 757–766 (2006)
Patel, S. S., Wong, I. & Johnson, K. A. Pre-steady-state kinetic analysis of processive DNA replication including complete characterization of an exonuclease-deficient mutant. Biochemistry 30, 511–525 (1991)
Joyce, C. M. & Benkovic, S. J. DNA polymerase fidelity: kinetics, structure, and checkpoints. Biochemistry 43, 14317–14324 (2004)
Showalter, A. K. & Tsai, M. D. A reexamination of the nucleotide incorporation fidelity of DNA polymerases. Biochemistry 41, 10571–10576 (2002)
Shah, A. M., Li, S. X., Anderson, K. S. & Sweasy, J. B. Y265H mutator mutant of DNA polymerase β. Proper teometric alignment is critical for fidelity. J. Biol. Chem. 276, 10824–10831 (2001)
Fiala, K. A. & Suo, Z. Mechanism of DNA polymerization catalyzed by Sulfolobus solfataricus P2 DNA polymerase IV. Biochemistry 43, 2116–2125 (2004)
Cramer, J. & Restle, T. Pre-steady-state kinetic characterization of the DinB homologue DNA polymerase of Sulfolobus solfataricus . J. Biol. Chem. 280, 40552–40558 (2005)
Choi, J. Y. & Guengerich, F. P. Adduct size limits efficient and error-free bypass across bulky N2-guanine DNA lesions by human DNA polymerase η. J. Mol. Biol. 352, 72–90 (2005)
Rothwell, P. J., Mitaksov, V. & Waksman, G. Motions of the fingers subdomain of klentaq1 are fast and not rate limiting: implications for the molecular basis of fidelity in DNA polymerases. Mol. Cell 19, 345–355 (2005)
Zhang, H., Cao, W., Zakharova, E., Konigsberg, W. & De La Cruz, E. M. Fluorescence of 2-aminopurine reveals rapid conformational changes in the RB69 DNA polymerase-primer/template complexes upon binding and incorporation of matched deoxynucleoside triphosphates. Nucleic Acids Res. 35, 6052–6062 (2007)
Yang, W. & Woodgate, R. What a difference a decade makes: insights into translesion DNA synthesis. Proc. Natl Acad. Sci. USA 104, 15591–15598 (2007)
Biertümpfel, C. et al. Structure and mechanism of human DNA polymerase η. Nature 465, 1044–1048 (2010)
Zhao, Y., Biertümpfel, C., Gregory, M. T., Hua, Y. J., Hanaoka, F. & Wang, W. Structural basis of human DNA polymerase η-mediated chemoresistance to cisplatin. Proc. Natl Acad. Sci. USA 109, 7269–7274 (2012)
Castro, C. et al. Two proton transfers in the transition state for nucleotidyl transfer catalyzed by RNA- and DNA-dependent RNA and DNA polymerases. Proc. Natl Acad. Sci. USA 104, 4267–4272 (2007)
Garcia-Diaz, M., Bebenek, K., Krahn, J. M., Pedersen, L. C. & Kunkel, T. A. Role of the catalytic metal during polymerization by DNA polymerase lambda. DNA Repair 6, 1333–1340 (2007)
Le Du, M. H. et al. Artificial evolution of an enzyme active site: structural studies of three highly active mutants of Escherichia coli alkaline phosphatase. J. Mol. Biol. 316, 941–953 (2002)
Graham, D. L. et al. MgF3 − as a transition state analog of phosphoryl transfer. Chem. Biol. 9, 375–381 (2002)
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell 127, 1349–1360 (2006)
Cavanaugh, N. A., Beard, W. A. & Wilson, S. H. DNA polymerase β ribonucleotide discrimination: insertion, misinsertion, extension, and coding. J. Biol. Chem. 285, 24457–24465 (2010)
Wang, L., Broyde, S. & Zhang, Y. Polymerase-tailored variations in the water-mediated and substrate-assisted mechanism for nucleotidyl transfer: insights from a study of T7 DNA polymerase. J. Mol. Biol. 389, 787–796 (2009)
Wang, L., Yu, X., Hu, P., Broyde, S. & Zhang, Y. A water-mediated and substrate-assisted catalytic mechanism for Sulfolobus solfataricus DNA polymerase IV. J. Am. Chem. Soc. 129, 4731–4737 (2007)
Polesky, A. H., Dahlberg, M. E., Benkovic, S. J., Grindley, N. D. & Joyce, C. M. Side chains involved in catalysis of the polymerase reaction of DNA polymerase I from Escherichia coli . J. Biol. Chem. 267, 8417–8428 (1992)
Copeland, W. C. & Wang, T. S. Mutational analysis of the human DNA polymerase α. The most conserved region in α-like DNA polymerases is involved in metal-specific catalysis. J. Biol. Chem. 268, 11028–11040 (1993)
Zakharova, E., Wang, J. & Konigsberg, W. The activity of selected RB69 DNA polymerase mutants can be restored by manganese ions: the existence of alternative metal ion ligands used during the polymerization cycle. Biochemistry 43, 6587–6595 (2004)
Kim, E. E. & Wyckoff, H. W. Reaction mechanism of alkaline phosphatase based on crystal structures. Two-metal ion catalysis. J. Mol. Biol. 218, 449–464 (1991)
Romier, C., Dominguez, R., Lahm, A., Dahl, O. & Suck, D. Recognition of single-stranded DNA by nuclease P1: high resolution crystal structures of complexes with substrate analogs. Proteins 32, 414–424 (1998)
Garcin, E. D. et al. DNA apurinic–apyrimidinic site binding and excision by endonuclease IV. Nature Struct. Mol. Biol. 15, 515–522 (2008)
Shan, S., Yoshida, A., Sun, S., Piccirilli, J. A. & Herschlag, D. Three metal ions at the active site of the Tetrahymena group I ribozyme. Proc. Natl Acad. Sci. USA 96, 12299–12304 (1999)
Stahley, M. R. & Strobel, S. A. Structural evidence for a two-metal-ion mechanism of group I intron splicing. Science 309, 1587–1590 (2005)
Ding, X., Rasmussen, B. F., Petsko, G. A. & Ringe, D. Direct structural observation of an acyl-enzyme intermediate in the hydrolysis of an ester substrate by elastase. Biochemistry 33, 9285–9293 (1994)
Scott, W. G., Murray, J. B., Arnold, J. R., Stoddard, B. L. & Klug, A. Capturing the structure of a catalytic RNA intermediate: the hammerhead ribozyme. Science 274, 2065–2069 (1996)
Murray, J. B., Szoke, H., Szoke, A. & Scott, W. G. Capture and visualization of a catalytic RNA enzyme–product complex using crystal lattice trapping and X-ray holographic reconstruction. Mol. Cell 5, 279–287 (2000)
Yang, W. Nucleases: diversity of structure, function and mechanism. Q. Rev. Biophys. 44, 1–93 (2011)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot . Acta Crystallogr. D 66, 486–501 (2010)
Acknowledgements
We thank D. Leahy, M. Gellert and R. Craigie for editing the manuscript. The research was supported by the intramural research program of the National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health (W.Y., T.N. and Y.Z.); the Japan Society for the Promotion of Science Institutional Program for Young Researcher Overseas visits, Kumamoto University, and the Kumayaku Alumni Research Fund (T.N.); Chinese Ministry of Education scholarship (Y.Z.); National Natural Science Foundation of China (Y.-J.H.); and Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (Y.Y.).
Author information
Authors and Affiliations
Contributions
T.N. and Y.Z. carried out crystallography. Y.Z. did the mutagenesis and kinetic analyses. T.N. handled crystals, refined structures and analysed reaction time courses. W.Y. designed the project. T.N., Y.Z. and W.Y. prepared the manuscript. Y.Y. supervised T.N. and Y.-J.H. advised Y.Z. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3 and Supplementary Figures 1-5. (PDF 2963 kb)
Supplementary Movie 1
This file contains an animation of the chemical bond formation. (MOV 15373 kb)
Rights and permissions
About this article
Cite this article
Nakamura, T., Zhao, Y., Yamagata, Y. et al. Watching DNA polymerase η make a phosphodiester bond. Nature 487, 196–201 (2012). https://doi.org/10.1038/nature11181
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11181
This article is cited by
-
Monovalent metal ion binding promotes the first transesterification reaction in the spliceosome
Nature Communications (2023)
-
Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints
Nature Communications (2022)
-
In crystallo observation of three metal ion promoted DNA polymerase misincorporation
Nature Communications (2022)
-
Watching a double strand break repair polymerase insert a pro-mutagenic oxidized nucleotide
Nature Communications (2021)
-
Structural basis of DNA synthesis opposite 8-oxoguanine by human PrimPol primase-polymerase
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.