Abstract
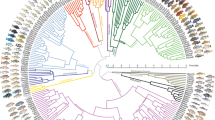
A fundamental challenge to our understanding of biodiversity is to explain why some groups of species undergo adaptive radiations, diversifying extensively into many and varied species, whereas others do not1,2. Both extrinsic environmental factors (for example, resource availability, climate) and intrinsic lineage-specific traits (for example, behavioural or morphological traits, genetic architecture) influence diversification, but few studies have addressed how such factors interact. Radiations of cichlid fishes in the African Great Lakes provide some of the most dramatic cases of species diversification. However, most cichlid lineages in African lakes have not undergone adaptive radiations. Here we compile data on cichlid colonization and diversification in 46 African lakes, along with lake environmental features and information about the traits of colonizing cichlid lineages, to investigate why adaptive radiation does and does not occur. We find that extrinsic environmental factors related to ecological opportunity and intrinsic lineage-specific traits related to sexual selection both strongly influence whether cichlids radiate. Cichlids are more likely to radiate in deep lakes, in regions with more incident solar radiation and in lakes where there has been more time for diversification. Weak or negative associations between diversification and lake surface area indicate that cichlid speciation is not constrained by area, in contrast to diversification in many terrestrial taxa3. Among the suite of intrinsic traits that we investigate, sexual dichromatism, a surrogate for the intensity of sexual selection, is consistently positively associated with diversification. Thus, for cichlids, it is the coincidence between ecological opportunity and sexual selection that best predicts whether adaptive radiation will occur. These findings suggest that adaptive radiation is predictable, but only when species traits and environmental factors are jointly considered.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Data deposits
Sequence data are deposited in the GenBank database under accession numbers listed in Supplementary Table 1.
References
Simpson, G. G. The Major Features of Evolution (Columbia Univ. Press, 1953)
Losos, J. B. Adaptive radiation, ecological opportunity, and evolutionary determinism. Am. Nat. 175, 623–639 (2010)
Kisel, Y. & Barraclough, T. G. Speciation has a spatial scale that depends on levels of gene flow. Am. Nat. 175, 316–334 (2010)
Schluter, D. The Ecology of Adaptive Radiation (Oxford Univ. Press, 2000)
Vamosi, S. M. The presence of other fish species affects speciation in threespine sticklebacks. Evol. Ecol. Res. 5, 717–730 (2003)
MacArthur, R. H. & Wilson, E. O. Equilibrium-theory of insular zoogeography. Evolution 17, 373–. (1963)
Price, S. A., Holzman, R., Near, T. J. & Wainwright, P. C. Coral reefs promote the evolution of morphological diversity and ecological novelty in labrid fishes. Ecol. Lett. 14, 462–469 (2011)
Mittelbach, G. G. et al. Evolution and the latitudinal diversity gradient: speciation, extinction and biogeography. Ecol. Lett. 10, 315–331 (2007)
Evans, K. L., Warren, P. H. & Gaston, K. J. Species-energy relationships at the macroecological scale: a review of the mechanisms. Biol. Rev. Camb. Philos. Soc. 80, 1–25 (2005)
Kraaijeveld, K., Kraaijeveld-Smit, F. J. L. & Maan, M. E. Sexual selection and speciation: the comparative evidence revisited. Biol. Rev. Camb. Philos. Soc. 86, 366–377 (2010)
Farrell, B. D. “Inordinate fondness” explained: why are there so many beetles? Science 281, 555–559 (1998)
Liem, K. F. Evolutionary strategies and morphological innovations: cichlid pharyngeal jaws. Syst. Zool. 22, 425–441 (1973)
Ricklefs, R. E. History and diversity: explorations at the intersection of ecology and evolution. Am. Nat. 170, S56–S70 (2007)
Fryer, G. Some aspects of evolution in Lake Nyasa. Evolution 13, 440–451 (1959)
Sturmbauer, C., Baric, S., Salzburger, W., Ruber, L. & Verheyen, E. Lake level fluctuations synchronize genetic divergences of cichlid fishes in African lakes. Mol. Biol. Evol. 18, 144–154 (2001)
Seehausen, O. & van Alphen, J. M. Can sympatric speciation by disruptive sexual selection explain rapid evolution of cichlid diversity in Lake Victoria? Ecol. Lett. 2, 262–271 (1999)
Seehausen, O. Evolution and ecological theory—chance, historical contingency and ecological determinism jointly determine the rate of adaptive radiation. Heredity 99, 361–363 (2007)
Greenwood, P. H. Towards a phyletic classification of the ‘genus’ Haplochromis (Pisces, Cichlidae) and related taxa. Part 1. Bull. Br. Mus. Nat. Hist. Zool. 35, 265–322 (1979)
Burnham, K. P. & Anderson, D. R. Model Selection and Multimodel Inference (Springer, 2002)
Ingram, T. Speciation along a depth gradient in a marine adaptive radiation. Proc. R. Soc. B 278, 613–618 (2011)
Seehausen, O. et al. Speciation through sensory drive in cichlid fish. Nature 455, 620–626 (2008)
Losos, J. B. & Schluter, D. Analysis of an evolutionary species–area relationship. Nature 408, 847–850 (2000)
Lande, R. Rapid origin of sexual isolation and character divergence in a cline. Evolution 36, 1–12 (1982)
Maan, M. E. & Seehausen, O. Ecology, sexual selection and speciation. Ecol. Lett. 14, 591–602 (2011)
Trivers, R. L. in Sexual Selection and the Descent of Man (ed. Campbell, B.) 136–179 (Aldine, 1972)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006)
Britton, T., Anderson, C. L., Jacquet, D., Lundqvist, S. & Bremer, K. Estimating divergence times in large phylogenetic trees. Syst. Biol. 56, 741–752 (2007)
Ives, A. R. & Garland, T. Phylogenetic logistic regression for binary dependent variables. Syst. Biol. 59, 9–26 (2010)
Kuwamura, T. in Fish communities in Lake Tanganyika (eds Kawanabe, H., Hori, M. & Nagoshi, M. ) 59–86 (Kyoto Univ. Press, 1997)
Worthington, E. B. & Ricardo, C. K. The fish of Lake Tanganyika (other than Cichlidae). Proc. Zool. Soc. Lond. 1936, 1061–1112 (1936)
Menard, S. Coefficients of determination for multiple logistic regression analysis. Am. Stat. 54, 17–24 (2000)
Quinn, G. P. & Keough, M. J. Experimental Design and Data Analysis for Biologists (Cambridge Univ. Press, 2002)
Hadfield, J. D. MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, 1–22 (2010)
Acknowledgements
We thank A. Ives, T. J. Davies and A. Mooers for analytical advice, P. McIntyre and C. Reidy Liermann for access to global energy data, S. Mwaiko, R. B. Stelkens, C. Katongo and U. Schliewen for unpublished DNA sequences, U. Schliewen, J. Jenson and O. Rittner for photographs, and A. McCune, D. Rabosky, R. Harrison, I. Lovette, E. Michel, G. Mittelbach, C. Melian, J. Brodersen, M. Maan, T. Ingram, B. Matthews, B. Dalziel, M. Pennell, J. Eastman, the Harmon laboratory group, the McCune laboratory group and the Seehausen laboratory group for discussions and comments on the manuscript. Bioinformatics facilities were supported by grants from the National Center for Research Resources (5P20RR016448-10) and the National Institute of General Medical Sciences (8 P20 GM103397-10) from the National Institutes of Health. This work was supported by the Swiss National Science Foundation project 31003A-118293 (to O.S.) and US National Science Foundation grant DEB 0919499 (to L.J.H.).
Author information
Authors and Affiliations
Contributions
C.E.W., L.J.H. and O.S. designed the study. O.S. and C.E.W. collected the data. C.E.W. conducted the analyses. C.E.W., L.J.H. and O.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary References, Supplementary Figures 1-6 and Supplementary Tables 1-6. (PDF 8016 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Wagner, C., Harmon, L. & Seehausen, O. Ecological opportunity and sexual selection together predict adaptive radiation. Nature 487, 366–369 (2012). https://doi.org/10.1038/nature11144
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11144
This article is cited by
-
East African cichlid fishes
EvoDevo (2023)
-
A continuous fish fossil record reveals key insights into adaptive radiation
Nature (2023)
-
The non-Haplochromis fish fauna in Uganda: an update on the distribution and a review of data gaps
Environmental Monitoring and Assessment (2023)
-
Testing Wickler’s hypothesis: cichlids are unable to distinguish eggs from egg spots in the wild
Hydrobiologia (2023)
-
Exceptional Genetic Differentiation at a Micro-geographic Scale in Apistogramma agassizii (Steindachner, 1875) from the Peruvian Amazon: Sympatric Speciation?
Evolutionary Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.