Abstract
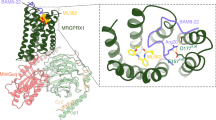
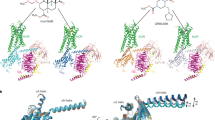
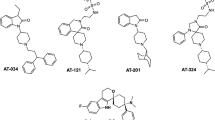
Members of the opioid receptor family of G-protein-coupled receptors (GPCRs) are found throughout the peripheral and central nervous system, where they have key roles in nociception and analgesia. Unlike the ‘classical’ opioid receptors, δ, κ and μ (δ-OR, κ-OR and μ-OR), which were delineated by pharmacological criteria in the 1970s and 1980s, the nociceptin/orphanin FQ (N/OFQ) peptide receptor (NOP, also known as ORL-1) was discovered relatively recently by molecular cloning and characterization of an orphan GPCR1. Although it shares high sequence similarity with classical opioid GPCR subtypes (∼60%), NOP has a markedly distinct pharmacology, featuring activation by the endogenous peptide N/OFQ, and unique selectivity for exogenous ligands2,3. Here we report the crystal structure of human NOP, solved in complex with the peptide mimetic antagonist compound-24 (C-24) (ref. 4), revealing atomic details of ligand–receptor recognition and selectivity. Compound-24 mimics the first four amino-terminal residues of the NOP-selective peptide antagonist UFP-101, a close derivative of N/OFQ, and provides important clues to the binding of these peptides. The X-ray structure also shows substantial conformational differences in the pocket regions between NOP and the classical opioid receptors κ (ref. 5) and μ (ref. 6), and these are probably due to a small number of residues that vary between these receptors. The NOP–compound-24 structure explains the divergent selectivity profile of NOP and provides a new structural template for the design of NOP ligands.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mollereau, C. et al. ORL1, a novel member of the opioid receptor family. Cloning, functional expression and localization. FEBS Lett. 341, 33–38 (1994)
Meunier, J. C. et al. Isolation and structure of the endogenous agonist of opioid receptor-like ORL1 receptor. Nature 377, 532–535 (1995)
Reinscheid, R. K. et al. Orphanin FQ: a neuropeptide that activates an opioidlike G protein-coupled receptor. Science 270, 792–794 (1995)
Goto, Y. et al. Identification of a novel spiropiperidine opioid receptor-like 1 antagonist class by a focused library approach featuring 3D-pharmacophore similarity. J. Med. Chem. 49, 847–849 (2006)
Wu, H. et al. Structure of the human kappa opioid receptor in complex with JDTic. Nature advance online publication 10.1038/nature10939 (21 March 2012)
Manglik, A. et al. Crystal structure of the μ-opioid receptor bound to a morphinan antagonist. Nature advance online publication 10.1038/nature10954 (21 March 2012)
Lambert, D. G. The nociceptin/orphanin FQ receptor: a target with broad therapeutic potential. Nature Rev. Drug Discov. 7, 694–710 (2008)
Chu, R. et al. Redesign of a four-helix bundle protein by phage display coupled with proteolysis and structural characterization by NMR and X-ray crystallography. J. Mol. Biol. 323, 253–262 (2002)
Varty, G. B. et al. The anxiolytic-like effects of the novel, orally active nociceptin opioid receptor agonist 8-[bis(2-methylphenyl)methyl]-3-phenyl-8-azabicyclo[3.2.1]octan-3-ol (SCH 221510). J. Pharmacol. Exp. Ther. 326, 672–682 (2008)
Fischetti, C. et al. Pharmacological characterization of the nociceptin/orphanin FQ receptor non peptide antagonist Compound 24. Eur. J. Pharmacol. 614, 50–57 (2009)
Volta, M., Viaro, R., Trapella, C., Marti, M. & Morari, M. Dopamine-nociceptin/orphanin FQ interactions in the substantia nigra reticulata of hemiparkinsonian rats: involvement of D2/D3 receptors and impact on nigro-thalamic neurons and motor activity. Exp. Neurol. 228, 126–137 (2011)
Chavkin, C. & Goldstein, A. Specific receptor for the opioid peptide dynorphin: structure–activity relationships. Proc. Natl Acad. Sci. USA 78, 6543–6547 (1981)
Katritch, V., Cherezov, V. & Stevens, R. C. Diversity and modularity of G protein-coupled receptor structures. Trends Pharmacol. Sci. 33, 17–27 (2012)
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010)
Fredriksson, R., Lagerstrom, M. C., Lundin, L. G. & Schioth, H. B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003)
Lapalu, S. et al. Comparison of the structure-activity relationships of nociceptin and dynorphin A using chimeric peptides. FEBS Lett. 417, 333–336 (1997)
Doré, A. S. et al. Structure of the adenosine A2A receptor in complex with ZM241385 and the xanthines XAC and caffeine. Structure 19, 1283–1293 (2011)
Mouledous, L., Topham, C. M., Moisand, C., Mollereau, C. & Meunier, J. C. Functional inactivation of the nociceptin receptor by alanine substitution of glutamine 286 at the C terminus of transmembrane segment VI: evidence from a site-directed mutagenesis study of the ORL1 receptor transmembrane-binding domain. Mol. Pharmacol. 57, 495–502 (2000)
Akuzawa, N., Takeda, S. & Ishiguro, M. Structural modelling and mutation analysis of a nociceptin receptor and its ligand complexes. J. Biochem. 141, 907–916 (2007)
Topham, C. M., Mouledous, L., Poda, G., Maigret, B. & Meunier, J. C. Molecular modelling of the ORL1 receptor and its complex with nociceptin. Protein Eng. 11, 1163–1179 (1998)
Trapella, C. et al. Structure-activity studies on the nociceptin/orphanin FQ receptor antagonist 1-benzyl-N-{3-[spiroisobenzofuran-1(3H),4'-piperidin-1-yl]propyl} pyrrolidine-2-carboxamide. Bioorg. Med. Chem. 17, 5080–5095 (2009)
Guerrini, R. et al. Further studies on nociceptin-related peptides: discovery of a new chemical template with antagonist activity on the nociceptin receptor. J. Med. Chem. 43, 2805–2813 (2000)
Calo, G. et al. [Nphe1,Arg14,Lys15]nociceptin-NH2, a novel potent and selective antagonist of the nociceptin/orphanin FQ receptor. Br. J. Pharmacol. 136, 303–311 (2002)
Tancredi, T. et al. The interaction of highly helical structural mutants with the NOP receptor discloses the role of the address domain of nociceptin/orphanin FQ. Chemistry 11, 2061–2070 (2005)
Orsini, M. J. et al. The nociceptin pharmacophore site for opioid receptor binding derived from the NMR structure and bioactivity relationships. J. Biol. Chem. 280, 8134–8142 (2005)
Bes, B. & Meunier, J. C. Identification of a hexapeptide binding region in the nociceptin (ORL1) receptor by photo-affinity labelling with Ac-Arg-Bpa-Tyr-Arg-Trp-Arg-NH2. Biochem. Biophys. Res. Commun. 310, 992–1001 (2003)
Meng, F. et al. Moving from the orphanin FQ receptor to an opioid receptor using four point mutations. J. Biol. Chem. 271, 32016–32020 (1996)
Meng, F. et al. Creating a functional opioid alkaloid binding site in the orphanin FQ receptor through site-directed mutagenesis. Mol. Pharmacol. 53, 772–777 (1998)
Heckman, K. L. & Pease, L. R. Gene splicing and mutagenesis by PCR-driven overlap extension. Nature Protocols 2, 924–932 (2007)
Kimple, A. J. et al. Structural determinants of G-protein alpha subunit selectivity by regulator of G-protein signaling 2 (RGS2). J. Biol. Chem. 284, 19402–19411 (2009)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nature Protocols 4, 706–731 (2009)
Cherezov, V., Peddi, A., Muthusubramaniam, L., Zheng, Y. F. & Caffrey, M. A robotic system for crystallizing membrane and soluble proteins in lipidic mesophases. Acta Crystallogr. D 60, 1795–1807 (2004)
Otwinowski Z & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
The PyMOL Molecular Graphics System. v.1.4.1. (2011)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Totrov, M. & Abagyan, R. Flexible protein-ligand docking by global energy optimization in internal coordinates. Proteins 29 (suppl.). 215–220 (1997)
Katritch, V. et al. Analysis of full and partial agonists binding to β 2-adrenergic receptor suggests a role of transmembrane helix V in agonist-specific conformational changes. J. Mol. Recognit. 22, 307–318 (2009)
Acknowledgements
This work was supported by PSI:Biology grant U54 GM094618 for biological studies and structure production, NIH Roadmap grant P50 GM073197 for technology development and R01 DA017204, R01 DA27170, and the NIMH Psychoactive Drug Screening Program (X.-P.H., E.V. and B.L.R.) and the Michael Hooker Chair of Pharmacology (B.L.R.), University of Ferrara (FAR grant to G.C.), Italian Ministry of University (FIRB Futuro in Ricerca 2010 grant to C.T.). We thank J. Francis for suggesting the idea to pursue the NOP receptor; J. Velasquez for help on molecular biology; T. Trinh, K. Allin and M. Chu for help on baculovirus expression; A. Walker and E. Abola for assistance with manuscript preparation; J. Smith, R. Fischetti and N. Sanishvili for assistance in development and use of the minibeam and beamtime at GM/CA-CAT beamline 23-ID at the Advanced Photon Source, which is supported by National Cancer Institute grant Y1-CO-1020 and National Institute of General Medical Sciences grant Y1-GM-1104.
Author information
Authors and Affiliations
Contributions
A.A.T. optimized the constructs, purified and crystallized the receptor in LCP, optimized crystallization conditions, grew crystals for data collection, collected the data and refined the structure, and prepared the manuscript. W.L. assisted with LCP experiments, performed fluorescence recovery after photobleaching assays, collected diffraction data, and assisted with preparing the manuscript. E.C. assisted in construct optimization, assisted with LCP experiments, collected diffraction data, and assisted with preparing the manuscript. V.K. performed the receptor docking and prepared the manuscript. H.W. assisted with membrane preparations, provided advice on crystallization strategies, and assisted with preparing the manuscript. E.V. and X.-P.H. performed ligand-binding and site-directed mutagenesis studies. C.T., R.G. and G.C. suggested the use of and synthesized numerous ligands for crystallization and pharmacological studies, and assisted with preparing the manuscript. B.L.R. supervised the pharmacology and mutagenesis studies and prepared the manuscript. V.C. assisted with the crystallization in LCP, collected diffraction data, processed diffraction data, and prepared the manuscript. R.C.S. was responsible for the overall project strategy and management and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.C.S. is a founder and Board of Directors member of Receptos, a GPCR drug discovery company.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5, Supplementary Figures 1-8 and additional references.The correct Supplementary Information was added on 29th May 2012. (PDF 1713 kb)
Rights and permissions
About this article
Cite this article
Thompson, A., Liu, W., Chun, E. et al. Structure of the nociceptin/orphanin FQ receptor in complex with a peptide mimetic. Nature 485, 395–399 (2012). https://doi.org/10.1038/nature11085
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11085
This article is cited by
-
Exploring the binding interactions of NOP receptor with designed natural phytochemical-neuropeptide conjugates: an in silico and SPR study
Applied Biological Chemistry (2024)
-
Nociceptin Receptor-Related Agonists as Safe and Non-addictive Analgesics
Drugs (2023)
-
Regulation of N-type calcium channels by nociceptin receptors and its possible role in neurological disorders
Molecular Brain (2022)
-
Universal platform for the generation of thermostabilized GPCRs that crystallize in LCP
Nature Protocols (2022)
-
N/OFQ modulates orofacial pain induced by tooth movement through CGRP-dependent pathways
BMC Neuroscience (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.