Abstract
The evolutionary importance of hybridization and introgression has long been debated1. Hybrids are usually rare and unfit, but even infrequent hybridization can aid adaptation by transferring beneficial traits between species. Here we use genomic tools to investigate introgression in Heliconius, a rapidly radiating genus of neotropical butterflies widely used in studies of ecology, behaviour, mimicry and speciation2,3,4,5. We sequenced the genome of Heliconius melpomene and compared it with other taxa to investigate chromosomal evolution in Lepidoptera and gene flow among multiple Heliconius species and races. Among 12,669 predicted genes, biologically important expansions of families of chemosensory and Hox genes are particularly noteworthy. Chromosomal organization has remained broadly conserved since the Cretaceous period, when butterflies split from the Bombyx (silkmoth) lineage. Using genomic resequencing, we show hybrid exchange of genes between three co-mimics, Heliconius melpomene, Heliconius timareta and Heliconius elevatus, especially at two genomic regions that control mimicry pattern. We infer that closely related Heliconius species exchange protective colour-pattern genes promiscuously, implying that hybridization has an important role in adaptive radiation.
Similar content being viewed by others
Main
The butterfly genus Heliconius (Nymphalidae: Heliconiinae) is associated with a suite of derived life-history and ecological traits, including pollen feeding, extended lifespan, augmented ultraviolet colour vision, ‘trap-lining’ foraging behaviour, gregarious roosting and complex mating behaviours, and provides outstanding opportunities for genomic studies of adaptive radiation and speciation4,6. The genus is best known for the hundreds of races with different colour patterns seen among its 43 species, with repeated examples of both convergent evolution among distantly related species and divergent evolution between closely related taxa3. Geographic mosaics of multiple colour-pattern races, such as in Heliconius melpomene (Fig. 1), converge to similar mosaics in other species, and this led to the hypothesis of mimicry2. Heliconius are unpalatable to vertebrate predators and Müllerian mimicry of warning colour patterns enables species to share the cost of educating predators3. As a result of its dual role in mimicry and mate selection, divergence in wing pattern is also associated with speciation and adaptive radiation3,5. A particularly recent radiation is the melpomene–silvaniform clade, in which mimetic patterns often seem to be polyphyletic (Fig. 1a). Most species in this clade occasionally hybridize in the wild with other clade members7. Gene genealogies at a small number of loci indicate introgression between species8, and one non-mimetic species, Heliconius heurippa, has a hybrid origin9. Adaptive introgression of mimicry loci is therefore a plausible explanation for parallel evolution of multiple mimetic patterns in the melpomene–silvaniform clade.
a, Phylogenetic relationship of sequenced species and subspecies in the melpomene–silvaniform clade of Heliconius. Heliconius elevatus falls in the silvaniform clade, but it mimics colour patterns of melpomene–timareta clade taxa. Most other silvaniforms mimic unrelated ithomiine butterflies24. b, Geographic distribution of postman and rayed H. melpomene races studied here (blue, yellow and purple), and the entire distribution of H. melpomene (grey). The H. timareta races investigated have limited distributions (red) indicated by arrows and mimic sympatric races of H. melpomene. Heliconius elevatus and the other silvaniform species are distributed widely across the Amazon basin (Supplementary Information, section 22).
A Heliconius melpomene melpomene stock from Darién, Panama (Fig. 1), was inbred through five generations of sib mating. We sequenced a single male to ×38 coverage (after quality filtering) using combined 454 and Illumina technologies (Supplementary Information, sections 1–8). The complete draft genome assembly, which is 269 megabases (Mb) in size, consists of 3,807 scaffolds with an N50 of 277 kb and contains 12,669 predicted protein-coding genes. Restriction-site-associated DNA (RAD) linkage mapping was used to assign and order 83% of the sequenced genome onto the 21 chromosomes (Supplementary Information, section 4). These data permit a considerably improved genome-wide chromosomal synteny comparison with the silkmoth Bombyx mori10,11.
Using 6,010 orthologues identified between H. melpomene and B. mori, we found that 11 of 21 H. melpomene linkage groups show homology to single B. mori chromosomes and that ten linkage groups have major contributions from two B. mori chromosomes (Fig. 2a and Supplementary Information, section 8), revealing several previously unidentified chromosomal fusions. These fusions on the Heliconius lineage most probably occurred after divergence from the sister genus Eueides4, which has the lepidopteran modal karyotype of n = 31 (ref. 12). Three chromosomal fusions are evident in Bombyx (B. mori chromosomes 11, 23 and 24; Fig. 2a), as required for evolution of the Bombyx n = 28 karyotype from the ancestral n = 31 karyotype. Heliconius and Bombyx lineages diverged in the Cretaceous, more than 100 million years ago11, so the gross chromosomal structures of Lepidoptera genomes have remained highly conserved compared with those of flies or vertebrates13,14. By contrast, small-scale rearrangements were frequent. In the comparison with Bombyx, we estimate there to be 0.05–0.13 breaks per megabase per million years, and in that with Danaus plexippus (Monarch butterfly), we estimate there to be 0.04–0.29 breaks per megabase per million years. Although lower than previously suggested for Lepidoptera15, these rates are comparable to those in Drosophila (Supplementary Information, section 8).
a, Maps of the 21 Heliconius chromosomes (colour) and of the 28 Bombyx chromosomes (grey) based on positions of 6,010 orthologue pairs demonstrate highly conserved synteny and a shared n = 31 ancestor (Supplementary Information, section 8). Dotted lines within chromosomes indicate major chromosomal fusions. b, Maximum-likelihood tree showing expansions of chemosensory protein (CSP) genes in the two butterfly genomes.
The origin of butterflies was associated with a switch from nocturnal to diurnal behaviour, and a corresponding increase in visual communication16. Heliconius have increased visual complexity through expression of a duplicate ultraviolet opsin6, in addition to the long-wavelength-, blue- and ultraviolet-sensitive opsins in Bombyx. We might therefore predict reduced complexity of olfactory genes, but in fact Heliconius and Danaus17 genomes have more chemosensory genes than any other insect genome: 33 and 34, respectively (Supplementary Information, section 9). For comparison, there are 24 in Bombyx and 3–4 in Drosophila18. Lineage-specific expansions of chemosensory genes were evident in both Danaus and Heliconius (Fig. 2b). By contrast, all three lepidopteran genomes have similar numbers of odorant binding proteins and olfactory receptors (Supplementary Information, section 9). Hox genes are involved in body plan development and show strong conservation across animals. We identified four additional Hox genes located between the canonical Hox genes pb and zen, orthologous to shx genes in B. mori19 (Supplementary Information, section 10). These Hox gene duplications in the butterflies and Bombyx have a common origin and are independent of the two tandem duplications known in dipterans (zen2 and bcd). Immunity-related gene families are similar across all three lepidopterans (Supplementary Information, section 11), whereas there are extensive duplications and losses within dipterans20.
The Heliconius reference genome allowed us to perform rigorous tests for introgression among melpomene–silvaniform clade species. We used RAD resequencing to reconstruct a robust phylogenetic tree based on 84 individuals of H. melpomene and its relatives, sampling on average 12 Mb, or 4%, of the genome (Fig. 1a and Supplementary Information, sections 12–18). We then tested for introgression between the sympatric co-mimetic postman butterfly races of Heliconius melpomene amaryllis and H. timareta ssp. nov. (Fig. 1) in Peru, using ‘ABBA/BABA’ single nucleotide sites and Patterson’s D-statistics (Fig. 3a), originally developed to test for admixture between Neanderthals and modern humans21,22 (Supplementary Information, section 12). Genome-wide, we found an excess of ABBA sites, giving a significantly positive Patterson’s D of 0.037 ± 0.003 (two-tailed Z-test for D = 0, P = 1 × 10−40), indicating greater genome-wide introgression between the sympatric mimetic taxa H. melpomene amaryllis and H. timareta ssp. nov. than between H. melpomene aglaope and H. timareta ssp. nov., which do not overlap spatially (Fig. 1b). On the basis of these D-statistics, we estimate that 2–5% of the genome was exchanged21 between H. timareta and H. melpomene amaryllis, to the exclusion of H. melpomene aglaope. (Supplementary Information, section 12). Exchange was not random. Of the 21 chromosomes, 11 have significantly positive D-statistics, and the strongest signals of introgression were found on the two chromosomes containing known mimicry loci B/D and N/Yb (Fig. 3b and Supplementary Information, section 15).
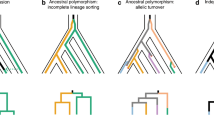
a, ABBA and BABA nucleotide sites employed in the test are derived (– – B –) in H. timareta compared with the silvaniform outgroup (– – – A), but differ among H. melpomene amaryllis and H. melpomene aglaope (either ABBA or BABA). As this almost exclusively restricts attention to sites polymorphic in the ancestor of H. timareta and H. melpomene, equal numbers of ABBA and BABA sites are expected under a null hypothesis of no introgression22, as depicted in the two gene genealogies. b, Distribution among chromosomes of Patterson’s D-statistic (±s.e.), which measures excess of ABBA sites over BABA sites22, here for the comparison: H. m. aglaope, H. m. amaryllis, H. timareta ssp. nov., silvaniform. Chromosomes containing the two colour-pattern regions (B/D, red; N/Yb, yellow) have the two highest D-statistics; the combinatorial probability of this occurring by chance is 0.005. The excess of ABBA sites (0 < D < 1) indicates introgression between sympatric H. timareta and H. m. amaryllis.
Perhaps the best-known case of Müllerian mimicry is the geographic mosaic of ∼30 bold postman and rayed colour-pattern races of H. melpomene (Fig. 1b and Supplementary Information, section 22), which mimic a near-identical colour-pattern mosaic in Heliconius erato (Fig. 1a), among other Heliconius species. Mimicry variation is mostly controlled by a few loci with strong effects. Mimetic pattern differences between the postman H. m. amaryllis and the rayed H. m. aglaope races studied here (Fig. 1a) are controlled by the B/D (red pattern) and N/Yb (yellow pattern) loci23,24. These loci are located on the two chromosomes that show the highest D-statistics in our RAD analysis (Fig. 3b). To test whether mimicry loci might be introgressed between co-mimetic H. timareta and H. melpomene7 (Fig. 1a), we resequenced the colour-pattern regions B/D (0.7 Mb) and N/Yb (1.2 Mb), and 1.8 Mb of unlinked regions across the genome, from both postman and ray-patterned H. melpomene and H. timareta from Peru and Colombia, and six silvaniform outgroup taxa (Fig. 1a and Supplementary Information, section 12). To test for introgression at the B/D mimicry locus, we compared rayed H. m. aglaope and postman H. m. amaryllis as the ingroup with postman H. timareta ssp. nov. (Fig. 3a) and found large, significant peaks of shared, fixed ABBA nucleotide sites combined with an almost complete lack of BABA sites (Fig. 4b). This provides evidence that blocks of shared sequence variation in the B/D region were exchanged between postman H. timareta and postman H. melpomene in the genomic region known to determine red mimicry patterns between races of H. melpomene23,24 (Fig. 4a).
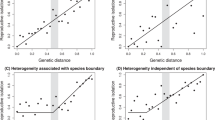
a, Genetic divergence between H. melpomene races aglaope (rayed) and amaryllis (postman) across a hybrid zone in northeast Peru. Divergence, FST, is measured along the B/D region (Supplementary Information 14) and peaks in the region known to control red wing pattern elements between the genes kinesin and optix23. b, c, Distribution of fixed ABBA and BABA sites (see Fig. 3a) along B/D for two comparisons. Excesses of ABBA in b and BABA in c are highly significant (two-tailed Z-tests for D = 0; D = 0.90 ± 0.13, P = 5 × 10−14 and D = −0.91 ± 0.10, P = 9 × 10−24, respectively), indicating introgression. d, e, f, Genealogical change along B/D investigated with maximum likelihood based on 50-kb windows. Three representative tree topologies are shown. Topology A, the species tree, is found within the white windows. In topologies B (dark green window) and C (light green windows) taxa group by colour pattern rather than by species. Within striped windows, H. melpomene and/or H. timareta are paraphyletic but the taxa do not group by colour pattern. Support is shown for nodes with >50% bootstrap support (Supplementary Information, section 19). bp, base pair.
For a reciprocal test, we used the same H. melpomene races as the ingroup to compare with rayed Heliconius timareta florencia at the B/D region. In this case, correspondingly large and significant peaks of BABA nucleotide sites are accompanied by an almost complete absence of ABBA sites (Fig. 4c), indicating that variation at the same mimicry locus was also shared between rayed H. timareta and rayed H. melpomene. Equivalent results in the N/Yb colour-pattern region, controlling yellow colour-pattern differences, are in the expected directions for introgression and are highly significant for the test using postman H. timareta ssp. nov. (P = 6 × 10−34), but are not significant in rayed H. t. florencia (P = 0.13; Supplementary Information, section 17). By contrast, hardly any ABBA or BABA sites are present in either comparison across 1.8 Mb in 55 genomic scaffolds that are unlinked to the colour-pattern regions (Supplementary Information, section 21). These concordant but reciprocal patterns of fixed ABBA and BABA substitutions occur almost exclusively within large genomic blocks at two different colour-pattern loci (449 and 99 sites for B/D and N/Yb, respectively; Fig. 4b, c and Supplementary Information, section 17). These patterns would be very hard to explain in terms of convergent functional-site evolution or random coalescent fluctuations. Instead, our results imply that derived colour-pattern elements have introgressed recently between both rayed and postman forms of H. timareta and H. melpomene.
To test whether colour-pattern loci might be shared more broadly across the clade, we used sliding-window phylogenetic analyses along the colour-pattern regions. For regions flanking and unlinked to colour-pattern loci, tree topologies are similar to the predominant signal recovered from the genome as a whole (Supplementary Information, section 18). Races of H. melpomene and H. timareta each form separate monophyletic sister groups and both are separated from the more distantly related silvaniform species (Fig. 4d). By contrast, topologies within the region of peak ABBA/BABA differences group individuals by colour pattern, and the species themselves become polyphyletic (Fig. 4e, f and Supplementary Information, sections 19 and 20). Remarkably, the rayed H. elevatus, a member of the silvaniform clade according to genome average relationships (Fig. 1a and Supplementary Information, section 18), groups with rayed races of unrelated H. melpomene and H. timareta in small sections within both B/D and N/Yb colour-pattern loci (Fig. 4e and Supplementary Information, sections 19 and 20). These results are again most readily explained by introgression and fixation of mimicry genes.
We have developed a de novo reference genome sequence that will facilitate evolutionary and ecological studies in this key group of butterflies. We have demonstrated repeated exchange of large (∼100-kb) adaptive regions among multiple species in a recent radiation. Our genome-scale analysis provides considerably greater power than previous tests of introgression8,25,26,27. Our evidence suggests that H. elevatus, like H. heurippa9, was formed during a hybrid speciation event. The main genomic signal from this rayed species places it closest to Heliconius pardalinus butleri (Fig. 1a), but colour-pattern genomic regions resemble those of rayed races of H. melpomene (Fig. 4e and Supplementary Information, sections 18–21). Colour pattern is important in mating behaviour in Heliconius5, and the transfer of mimetic pattern may have enabled the divergent sibling species H. elevatus to coexist with H. pardalinus across the Amazon basin. Although it was long suspected that introgression might be important in evolutionary radiations1, our results from the most diverse terrestrial biome on the planet suggest that adaptive introgression is more pervasive than previously realized.
The annotated genome version 1.1 is available on the Heliconius Genome Consortium’s genome browser at http://butterflygenome.org/ and this version will also be included in the next release of ENSEMBL Genomes. A full description of methods can be found in Supplementary Information.
Accession codes
Data deposits
The genome sequence has been submitted to the European Nucleotide Archive under accession numbers HE667773–HE672081. Additional short read sequences have been submitted to the European Nucleotide Archive under accession numbers ERP000993 and ERP000991.
References
Seehausen, O. Hybridization and adaptive radiation. Trends Ecol. Evol. 19, 198–207 (2004)
Bates, H. W. Contributions to an insect fauna of the Amazon valley. Lepidoptera: Heliconidae. Trans. Linn. Soc. Lond. 23, 495–566 (1862)
Turner, J. R. G. Adaptation and evolution in Heliconius: a defense of neo-Darwinism. Annu. Rev. Ecol. Syst. 12, 99–121 (1981)
Brown, K. S. The biology of Heliconius and related genera. Annu. Rev. Entomol. 26, 427–457 (1981)
Jiggins, C. D., Naisbit, R. E., Coe, R. L. & Mallet, J. Reproductive isolation caused by colour pattern mimicry. Nature 411, 302–305 (2001)
Briscoe, A. D. et al. Positive selection of a duplicated UV-sensitive visual pigment coincides with wing pigment evolution in Heliconius butterflies. Proc. Natl Acad. Sci. USA 107, 3628–3633 (2010)
Mallet, J. in Speciation and Patterns of Diversity (eds Butlin, R. K., Schluter, D. & Bridle, J. R. ) 177–194 (Cambridge Univ. Press, 2009)
Kronforst, M. R. Gene flow persists millions of years after speciation in Heliconius butterflies. BMC Evol. Biol. 8, 98 (2008)
Salazar, C. et al. Genetic evidence for hybrid trait speciation in Heliconius butterflies. PLoS Genet. 6, e1000930 (2010)
International. Silkworm Genome Consortium. The genome of a lepidopteran model insect, the silkworm Bombyx mori . Insect Biochem. Mol. Biol. 38, 1036–1045 (2008)
Pringle, E. G. et al. Synteny and chromosome evolution in the Lepidoptera: evidence from mapping in Heliconius melpomene . Genetics 177, 417–426 (2007)
Robinson, R. Lepidoptera Genetics 557–598 (Pergamon, 1971)
Deng, Q., Zeng, Q., Qian, Y., Li, C. & Yang, Y. Research on the karyotype and evolution of the Drosophila melanogaster species group. J. Genet. Genomics 34, 196–213 (2007)
Kemkemer, C. et al. Gene synteny comparisons between different vertebrates provide new insights into breakage and fusion events during mammalian karyotype evolution. BMC Evol. Biol. 9, 84 (2009)
d’Alençon, E. et al. Extensive synteny conservation of holocentric chromosomes in Lepidoptera despite high rates of local genome rearrangements. Proc. Natl Acad. Sci. USA 107, 7680–7685 (2010)
Vane-Wright, R. I. & Boppré, M. Visual and chemical signalling in butterflies: functional and phylogenetic perspectives. Philos. Trans. R. Soc. Lond., B 340, 197–205 (1993)
Zhan, S., Merlin, C., Boore, J. L. & Reppert, S. M. The monarch butterfly genome yields insights into long-distance migration. Cell 147, 1171–1185 (2011)
Vieira, F. G. & Rozas, J. Comparative genomics of the odorant-binding and chemosensory protein gene families across the Arthropoda: origin and evolutionary history of the chemosensory system. Genome Biol. Evol. 3, 476–490 (2011)
Chai, C. L. et al. A genomewide survey of homeobox genes and identification of novel structure of the Hox cluster in the silkworm, Bombyx mori . Insect Biochem. Mol. Biol. 38, 1111–1120 (2008)
Sackton, T. B. et al. Dynamic evolution of the innate immune system in Drosophila . Nature Genet. 39, 1461–1468 (2007)
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Durand, E. Y., Patterson, N., Reich, D. & Slatkin, M. Testing for ancient admixture between closely related populations. Mol. Biol. Evol. 28, 2239–2252 (2011)
Reed, R. D. et al. optix drives the repeated convergent evolution of butterfly wing pattern mimicry. Science 333, 1137–1141 (2011)
Nadeau, N. J. et al. Evidence for genomic islands of divergence among hybridizing species and subspecies of Heliconius butterflies obtained by large-scale targeted sequencing. Phil. Trans. R. Soc. B 367, 343–353 (2012)
Bull, V. et al. Polyphyly and gene flow between non-sibling Heliconius species. BMC Biol. 4, 11 (2006)
Kim, M. et al. Regulatory genes control a key morphological and ecological trait transferred between species. Science 322, 1116–1119 (2008)
Song, Y. et al. Adaptive introgression of anticoagulant rodent poison resistance by hybridization between Old World mice. Curr. Biol. 21, 1296–1301 (2011)
Acknowledgements
We thank the governments of Colombia, Peru and Panama for permission to collect the butterflies. Sequencing was funded by contributions from consortium members. We thank M. Abanto for assistance in raising the inbred line. Individual laboratories were funded by the Leverhulme Trust (C.D.J.), the John Fell Fund and Christ Church College, Oxford (L.C.F.), The Royal Society (M.J., C.D.J.), the NSF (W.O.M., M.R.K., R.D.R., S.M., A.D.B.), the NIH (M.R.K., S.L.S., J.A.Y.), the CNRS (M.J.), the ERC (M.J., P.W.H.H.), the Banco de la República and COLCIENCAS (M.L.) and the BBSRC (J.M., C.D.J., M.L.B. and R.H.f.-C.).
Author information
Authors and Affiliations
Consortia
Contributions
Consortium leaders: C.D.J., W.O.M. Heliconius Genome Consortium Principal Investigators: R.H.f.-C., M.R.K., M.J., J.M., S.M., R.D.R, M.L.B., L.E.G., M.L., G.L. Introgression study leader: J.M. Lead investigators: K.K.D., J.R.W., N.J.N., A.W., J.W.D., A.D.B., L.C.F., D.S.T.H., S.M., C.S., J.J.L., A.V.Z. Sequencing: S.R., S.E.S., A.L.B., M.T., K. Gharbi, C.E., M.L.B., R.A.G., Y.H., J.C.J., C.K., T.M., D.M.M., F.O., L.-L.P., J.Q., R.L.T., K.C.W., Y.-Q.W. Assembly: A.V.Z., J.A.Y., S.L.S., A.P., K. Gordon. RAD map and assembly verification: J.W.D., S.W.B., M.L.B., L.S.M., D.D.K., J.R.W., P.A.W. Geographic distribution map: N.R. Annotation: J.R.W., D.S.T.H., D.W., D.L., K.J.H., S.A., P.A.W., P.K. Genome browser and databases: D.S.T.H., J.J.L. Manual annotation and evolutionary analyses: A.D.B., E.J.-J., F.Y. (olfactory proteins); L.C.F., P.W.H.H., J.R.W. (Hox genes); A.S., T.D., D.M., S.M. (microRNAs); W.J.P., F.M.J. (immune genes); R.T.J., R.C. (P450 genes); H.V., S.-J.A., D.G.H. (uridine diphosphate glucuronosyltransferase genes); Y.P. (ribosomal proteins); S.W.B., M.L.B., A.D.B., N.L.C., B.A.C., L.C.F., H.M.H., C.D.J., F.M.J., M.J., D.D.K., M.R.K., J.M., A.M., S.P.M., N.J.N., W.J.P, R.P., M.A.S., A.T.-T., A.W., F.Y. (manual annotation group); B.A.C., D.A.R. (transposable elements); D.A.B. (orthologue predictions); A.W., J.W.D., D.G.H., K. Gordon (synteny); K.K.D., N.J.N., J.W.D., S.H.M., C.S., C.D.J., M.J., J.M. (introgression analysis). K.K.D. and J.R.W. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Contents, Supplementary Text, Supplementary Figures as listed in the Supplementary Contents, Supplementary Tables as listed in the Supplementary Contents and Supplementary References. (PDF 13571 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
The Heliconius Genome Consortium. Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487, 94–98 (2012). https://doi.org/10.1038/nature11041
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11041
This article is cited by
-
Surprise hybrid origins of a butterfly species
Nature (2024)
-
Unraveling the complex evolutionary history of lepidopteran chromosomes through ancestral chromosome reconstruction and novel chromosome nomenclature
BMC Biology (2023)
-
Hybridisation between host races broadens the host range of offspring in Eotetranychus asiaticus (Acari: Tetranychidae)
Experimental and Applied Acarology (2023)
-
Geographic Variation in Genomic Signals of Admixture Between Two Closely Related European Sepsid Fly Species
Evolutionary Biology (2023)
-
One’s trash is someone else’s treasure: sequence read archives from Lepidoptera genomes provide material for genome reconstruction of their endosymbionts
BMC Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.