Abstract
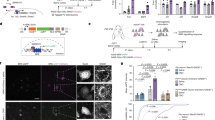
Autism spectrum disorders comprise a range of neurodevelopmental disorders characterized by deficits in social interaction and communication, and by repetitive behaviour1. Mutations in synaptic proteins such as neuroligins2,3, neurexins4, GKAPs/SAPAPs5 and ProSAPs/Shanks6,7,8,9,10 were identified in patients with autism spectrum disorder, but the causative mechanisms remain largely unknown. ProSAPs/Shanks build large homo- and heteromeric protein complexes at excitatory synapses and organize the complex protein machinery of the postsynaptic density in a laminar fashion11,12. Here we demonstrate that genetic deletion of ProSAP1/Shank2 results in an early, brain-region-specific upregulation of ionotropic glutamate receptors at the synapse and increased levels of ProSAP2/Shank3. Moreover, ProSAP1/Shank2−/− mutants exhibit fewer dendritic spines and show reduced basal synaptic transmission, a reduced frequency of miniature excitatory postsynaptic currents and enhanced N-methyl-d-aspartate receptor-mediated excitatory currents at the physiological level. Mutants are extremely hyperactive and display profound autistic-like behavioural alterations including repetitive grooming as well as abnormalities in vocal and social behaviours. By comparing the data on ProSAP1/Shank2−/− mutants with ProSAP2/Shank3αβ−/− mice, we show that different abnormalities in synaptic glutamate receptor expression can cause alterations in social interactions and communication. Accordingly, we propose that appropriate therapies for autism spectrum disorders are to be carefully matched to the underlying synaptopathic phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
13 June 2011
A present address was added to the affiliations.
References
Abrahams, B. S. & Geschwind, D. H. Advances in autism genetics: on the threshold of a new neurobiology. Nature Rev. Genet. 9, 341–355 (2008)
Jamain, S. et al. Mutations of the X-linked genes encoding neuroligins NLGN3 and NLGN4 are associated with autism. Nature Genet. 34, 27–29 (2003)
Etherton, M. R., Tabuchi, K., Sharma, M., Ko, J. & Südhof, T. C. An autism-associated point mutation in the neuroligin cytoplasmic tail selectively impairs AMPA receptor-mediated synaptic transmission in hippocampus. EMBO J. 30, 2908–2919 (2011)
Kim, H. G. et al. Disruption of neurexin 1 associated with autism spectrum disorder. Am. J. Hum. Genet. 82, 199–207 (2008)
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010)
Moessner, R. et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am. J. Hum. Genet. 81, 1289–1297 (2007)
Gauthier, J. et al. Novel de novo SHANK3 mutation in autistic patients. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 150B, 421–424 (2009)
Durand, C. M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nature Genet. 39, 25–27 (2007)
Berkel, S. et al. Mutations in the SHANK2 synaptic scaffolding gene in autism spectrum disorder and mental retardation. Nature Genet. 42, 489–491 (2010)
Leblond, C. S. et al. Genetic and functional analyses of SHANK2 mutations provide evidence for a multiple hit model of autism spectrum disorders. PLoS Genet. 8, e1002521 (2012)
Baron, M. K. et al. An architectural framework that may lie at the core of the postsynaptic density. Science 311, 531–535 (2006)
Grabrucker, A. M. et al. Concerted action of zinc and ProSAP/Shank in synaptogenesis and synapse maturation. EMBO J. 30, 569–581 (2011)
Toro, R. et al. Key role for gene dosage and synaptic homeostasis in autism spectrum disorders. Trends Genet. 26, 363–372 (2010)
Grabrucker, A. M., Schmeisser, M. J., Schoen, M. & Boeckers, T. M. Postsynaptic ProSAP/Shank scaffolds in the cross-hair of synaptopathies. Trends Cell Biol. 21, 594–603 (2011)
Hamdan, F. F. et al. Excess of de novo deleterious mutations in genes associated with glutamatergic systems in nonsyndromic intellectual disability. Am. J. Hum. Genet. 88, 306–316 (2011)
Bozdagi, O. et al. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol. Autism 1, 15 (2010)
Peca, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011)
Wang, X. et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum. Mol. Genet. 20, 3093–3108 (2011)
Bangash, M. A. et al. Enhanced polyubiquitination of Shank3 and NMDA receptor in a mouse model of autism. Cell 145, 758–772 (2011)
Chao, H. T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010)
Gemelli, T. et al. Postnatal loss of methyl-CpG binding protein 2 in the forebrain is sufficient to mediate behavioral aspects of Rett syndrome in mice. Biol. Psychiatry 59, 468–476 (2006)
Boeckers, T. M. et al. Proline-rich synapse-associated protein-1/cortactin binding protein 1 (ProSAP1/CortBP1) is a PDZ-domain protein highly enriched in the postsynaptic density. J. Neurosci. 19, 6506–6518 (1999)
Berkel, S. et al. Inherited and de novo SHANK2 variants associated with autism spectrum disorder impair neuronal morphogenesis and physiology. Hum. Mol. Genet. 21, 344–357 (2012)
Tabuchi, K. et al. A neuroligin-3 mutation implicated in autism increases inhibitory synaptic transmission in mice. Science 318, 71–76 (2007)
Pearson, B. L. et al. Motor and cognitive stereotypies in the BTBR T+tf/J mouse model of autism. Genes Brain Behav. 10, 228–235 (2011)
Hung, A. Y. et al. Smaller dendritic spines, weaker synaptic transmission, but enhanced spatial learning in mice lacking Shank1. J. Neurosci. 28, 1697–1708 (2008)
Wohr, M., Roullet, F. I., Hung, A. Y., Sheng, M. & Crawley, J. N. Communication impairments in mice lacking Shank1: reduced levels of ultrasonic vocalizations and scent marking behavior. PLoS ONE 6, e20631 (2011)
Silverman, J. L. et al. Sociability and motor functions in Shank1 mutant mice. Brain Res. 1380, 120–137 (2011)
Ey, E., Leblond, C. S. & Bourgeron, T. Behavioral profiles of mouse models for autism spectrum disorders. Autism Res. 4, 5–16 (2011)
Schmeisser, M. J., Grabrucker, A. M., Bockmann, J. & Boeckers, T. M. Synaptic cross-talk between N-methyl-D-aspartate receptors and LAPSER1-beta-catenin at excitatory synapses. J. Biol. Chem. 284, 29146–29157 (2009)
Boeckers, T. M. et al. Proline-rich synapse-associated protein-1/cortactin binding protein 1 (ProSAP1/CortBP1) is a PDZ-domain protein highly enriched in the postsynaptic density. J. Neurosci. 19, 6506–6518 (1999)
Grabrucker, A. M. et al. Concerted action of zinc and ProSAP/Shank in synaptogenesis and synapse maturation. EMBO J. 30, 569–581 (2011)
Zitzer, H., Hoenck, H. H., Baechner, D., Richter, D. & Kreienkamp, H. J. Somatostatin receptor interacting protein defines a novel family of multidomain proteins present in human and rodent brain. J. Biol. Chem. 274, 32997–33001 (1999)
Schmitz, D., Mellor, J., Breustedt, J. & Nicoll, R. A. Presynaptic kainate receptors impart an associative property to hippocampal mossy fiber long-term potentiation. Nature Neurosci. 6, 1058–1063 (2003)
Wishaw, I. Q., Haun, F. & Kolb, B. in Modern Techniques in Neuroscience (eds Windhorst, U. & Johansson, H. ) 1243–1275 (Springer, 1999)
Rogers, D. C. et al. Behavioral and functional analysis of mouse phenotype: SHIRPA, a proposed protocol for comprehensive phenotype assessment. Mamm. Genome 8, 711–713 (1997)
Nadler, J. J. et al. Automated apparatus for quantitation of social approach behaviors in mice. Genes Brain Behav. 3, 303–314 (2004)
Bourgeron, T., Jamain, S. & Granon, S. in Contemporary Clinical Neuroscience: Transgenic and Knockout Models of Neuropsychiatric Disorders (eds Fisch, G.S. & Flint, J. ) 151–174 (Humana Press, 2006)
Montag-Sallaz, M., Schachner, M. & Montag, D. Misguided axonal projections, NCAM180 mRNA upregulation, and altered behavior in mice deficient for the close homolog of L1 (CHL1). Mol. Cell. Biol. 22, 7967–7981 (2002)
Scattoni, M. L. Unusual repertoire of vocalizations in adult BTBR T+tf/J mice during three types of social encounters. Genes Brain Behav. 10, 44–56 (2011)
Acknowledgements
We thank M. Manz, R. Zienecker, S. Gerlach-Arbeiter, N. Damm, H. Riederer, C. Jean, S. Rieckmann, S. Hochmuth and K. Sowa for technical assistance. M.J.S., A.-L.J. and P.T.U. are members of the International Graduate School in Molecular Medicine at Ulm University. M.J.S. is further supported by Baustein 3.2 (L.SBN.0081), E.E. by the Fondation de France and the Agence Nationale de la Recherche (ANR) FLEXNEURIM (ANR09BLAN034003), S.W. and A.V.S. by the Deutsche Forschungsgemeinschaft (DFG) (GRK 1123), A.M.G. by Baustein 3.2 (L.SBN.0083), S.A.S by the DFG (EXC 257), D.S. by the DFG (SFB 618, SFB 665, EXC 257), the Bundesministerium für Bildung und Forschung (BMBF) (BCCN, BFNL) and the Einstein Foundation, M.R.K. by the DFG (SFB 779), C.S.L., R.T., N.T., A.LS. and T.B. by the ANR (ANR-08-MNPS-037-01 - SynGen), Neuron-ERANET (EUHF-AUTISM), Fondation Orange and the Fondation FondaMentale, P.F. by the Bettencourt-Schueller Fondation, R.T., T.B., P.F. by the CNRS Neuroinformatic, E.D.G. by the DFG (SFB 779) and the BMBF (EraNET Neuron), and T.M.B. by the DFG (Bo 1718/3-1 and 1718/4-1; SFB 497/B8).
Author information
Authors and Affiliations
Contributions
M.J.S., E.E., J.B., C.S., D.B., S.t.D., K.H.S., D.M., D.S., M.R.K., T.B., E.D.G. and T.M.B. designed the outline of this study. J.B. and B.V.S. generated, and J.B., C.S. and S.A.S. supervised breeding of, the ProSAP1/Shank2-mutant mice. J.B. supervised breeding of the ProSAP2/Shank3-mutant mice. M.J.S., A.K., A-L.J., P.T.U. and A.M.G. performed all the biochemistry, real-time PCR, Golgi stainings, electron microscopy, transfection of primary neurons and immunohistochemistry, E.E., C.S., D.M., C.S.L., P.F., N.T. and A.LS. the behavioural experiments, and S.W., A.V.S. and D.B. the electrophysiological experiments. E.S. conducted the survival analysis. M.J.S., E.E., S.W., A.V.S., C.S., D.B., D.M., R.T. and A.M.G. performed all data analyses and jointly drafted the manuscript with S.A.S., D.S., M.R.K., T.B., E.D.G. and T.M.B. All authors read and approved the final version. M.J.S., E.E. and S.W. contributed equally to this study. We thank H.-J. Kreienkamp, Hamburg, for providing the pan-Shank antibody ‘189.3’.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-15, Supplementary Methods, Supplementary References and Supplementary Tables. (PDF 3097 kb)
Rights and permissions
About this article
Cite this article
Schmeisser, M., Ey, E., Wegener, S. et al. Autistic-like behaviours and hyperactivity in mice lacking ProSAP1/Shank2. Nature 486, 256–260 (2012). https://doi.org/10.1038/nature11015
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11015
This article is cited by
-
Combined expansion and STED microscopy reveals altered fingerprints of postsynaptic nanostructure across brain regions in ASD-related SHANK3-deficiency
Molecular Psychiatry (2024)
-
The Shank/ProSAP N-Terminal (SPN) Domain of Shank3 Regulates Targeting to Postsynaptic Sites and Postsynaptic Signaling
Molecular Neurobiology (2024)
-
Expression profiles of the autism-related SHANK proteins in the human brain
BMC Biology (2023)
-
Shank3 deletion in PV neurons is associated with abnormal behaviors and neuronal functions that are rescued by increasing GABAergic signaling
Molecular Autism (2023)
-
Shank2 identifies a subset of glycinergic neurons involved in altered nociception in an autism model
Molecular Autism (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.