Abstract
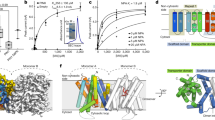
The phytohormone auxin acts as a prominent signal, providing, by its local accumulation or depletion in selected cells, a spatial and temporal reference for changes in the developmental program1,2,3,4,5,6,7. The distribution of auxin depends on both auxin metabolism (biosynthesis, conjugation and degradation)8,9,10 and cellular auxin transport11,12,13,14,15. We identified in silico a novel putative auxin transport facilitator family, called PIN-LIKES (PILS). Here we illustrate that PILS proteins are required for auxin-dependent regulation of plant growth by determining the cellular sensitivity to auxin. PILS proteins regulate intracellular auxin accumulation at the endoplasmic reticulum and thus auxin availability for nuclear auxin signalling. PILS activity affects the level of endogenous auxin indole-3-acetic acid (IAA), presumably via intracellular accumulation and metabolism. Our findings reveal that the transport machinery to compartmentalize auxin within the cell is of an unexpected molecular complexity and demonstrate this compartmentalization to be functionally important for a number of developmental processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Benková, E. et al. Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115, 591–602 (2003)
Friml, J. et al. Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426, 147–153 (2003)
Reinhardt, D. et al. Regulation of phyllotaxis by polar auxin transport. Nature 426, 255–260 (2003)
Leyser, O. Dynamic integration of auxin transport and signalling. Curr. Biol. 16, R424–R433 (2006)
Dubrovsky, J. G. et al. Auxin acts as a local morphogenetic trigger to specify lateral root founder cells. Proc. Natl Acad. Sci. USA 105, 8790–8794 (2008)
Sorefan, K. et al. A regulated auxin minimum is required for seed dispersal in Arabidopsis. Nature 459, 583–586 (2009)
Prasad, K. et al. Arabidopsis PLETHORA transcription factors control phyllotaxis. Curr. Biol. 21, 1123–1128 (2011)
Woodward, A. W. & Bartel, B. Auxin: regulation, action, and interaction. Ann. Bot. 95, 707–735 (2005)
Ikeda, Y. et al. Local auxin biosynthesis modulates gradient-directed planar polarity in Arabidopsis. Nature Cell Biol. 11, 731–738 (2009)
Zhao, Y. Auxin biosynthesis and its role in plant development. Annu. Rev. Plant Biol. 61, 49–64 (2010)
Bennett, M. J. et al. Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Science 273, 948–950 (1996)
Luschnig, C., Gaxiola, R. A., Grisafi, P. & Fink, G. R. EIR1, a root-specific protein involved in auxin transport, is required for gravitropism in Arabidopsis thaliana. Genes Dev. 12, 2175–2187 (1998)
Geisler, M. et al. Cellular efflux of auxin catalyzed by the Arabidopsis MDR/PGP transporter AtPGP1. Plant J. 44, 179–194 (2005)
Petrášek, J. et al. PIN proteins perform a rate-limiting function in cellular auxin efflux. Science 312, 914–918 (2006)
Zažímalová, E., Murphy, A. S., Yang, H., Hoyerová, K. & Hošek, P. Auxin transporters–why so many? Cold Spring Harb. Perspect. Biol. 2, a001552 (2010)
Mravec, J. et al. ER-localized PIN5 auxin transporter mediates subcellular homeostasis of phytohormone auxin. Nature 459, 1136–1140 (2009)
Ulmasov, T., Murfett, J., Hagen, G. & Guilfoyle, T. J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9, 1963–1971 (1997)
Lee, S. H. & Cho, H. T. PINOID positively regulates auxin efflux in Arabidopsis root hair cells and tobacco cells. Plant Cell 18, 1604–1616 (2006)
Nagata, T., Nemoto, Y. & Hasezawa, S. Tobacco BY-2 cell line as the “HeLa” cells in the cell biology of higher plants. Int. Rev. Cytol. 132, 1–30 (1992)
Marin, E. et al. miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 22, 1104–1117 (2010)
Langhans, M. et al. In vivo trafficking and localization of p24 proteins in plant cells. Traffic 9, 770–785 (2008)
Sauer, M., Paciorek, T., Benkova, E. & Friml, J. Immunocytochemical techniques for whole-mount in situ protein localization in plants. Nature Protocols 1, 98–103 (2006)
Delbarre, A., Muller, P., Imhoff, V. & Guern, J. Comparison of mechanisms controlling uptake and accumulation of 2,4-dichlorophenoxy acetic acid, naphthalene-1-acetic acid, and indole-3-acetic acid in suspension-cultured tobacco cells. Planta 198, 532–541 (1996)
Bailly, A. et al. Modulation of P-glycoproteins by auxin transport inhibitors is mediated by interaction with immunophilins. J. Biol. Chem. 283, 21817–21826 (2008)
Dobrev, P. I., Havlíček, L., Vágner, M., Malbeck, J. & Kamínek, M. Purification and determination of plant hormones auxin and abscisic acid using solid phase extraction and two-dimensional high performance liquid chromatography. J. Chromatogr. A 1075, 159–166 (2005)
Dobrev, P. I. & Kamínek, M. Fast and efficient separation of cytokinins from auxin and abscisic acid and their purification using mixed-mode solid-phase extraction. J. Chromatogr. A 950, 21–29 (2002)
Schultz, J., Milpetz, F., Bork, P. & Ponting, C. P. SMART, a simple modular architecture research tool: identification of signaling domains. Proc. Natl Acad. Sci. USA 95, 5857–5864 (1998)
Letunic, I., Doerks, T. & Bork, P. SMART 6: recent updates and new developments. Nucleic Acids Res. 37, 229–232 (2009)
Tusnády, G. E. & Simon, I. The HMMTOP transmembrane topology prediction server. Bioinformatics 17, 849–850 (2001)
Spyropoulos, I. C., Liakopoulos, T. D., Bagos, P. G. & Hamodrakas, S. J. TMRPres2D: high quality visual representation of transmembrane protein models. Bioinformatics 20, 3258–3260 (2004)
Proost, S. et al. PLAZA: a comparative genomics resource to study gene and genome evolution in plants. Plant Cell 21, 3718–3731 (2009)
Pěnčík, A. et al. Isolation of novel indole-3-acetic acid conjugates by immunoaffinity extraction. Talanta 80, 651–655 (2009)
Karimi, M., De Meyer, B. & Hilson, P. Modular cloning in plant cells. Trends Plant Sci. 10, 103–105 (2005)
Karimi, M., Inze, D. & Depicker, A. GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002)
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003)
Acknowledgements
We are grateful to C. Braeckman for plant transformation; W. Ardiles for sequencing support; L. Charrier for technical assistance; A. Maizel, N. Geldner and P. Pimpl for providing material; J.K.-V. group members for critical reading of the manuscript and the BOKU-VIBT Imaging Center for access and expertise. This work was supported by the Vienna Science and Technology Fund (WWTF) (to J.K.-V.), the Agency for Innovation by Science and Technology (IWT) (predoctoral fellowship to E.B.), the Odysseus program of the Research Foundation-Flanders (to J.F.), the Swiss National Funds (to M.G.), the Ministry of Education, Youth and Sports of the Czech Republic (LC06034) (to E.Z.), Grant Agency of the Czech Republic project P305/11/2476 (to J.P.) and P305/11/0797 (to E.Z).
Author information
Authors and Affiliations
Contributions
E.B. and J.K.V. conceived the project. E.B. carried out most of the experiments. M.K., P.I.B., E.Z. and J.P. performed auxin metabolite profile and auxin accumulation in BY-2. C.B. analysed auxin-dependent PILS expression and contributed to phenotype analysis. M.R.R. contributed to PILS cloning. J.R. and A.P. measured auxin content in Arabidopsis. B.W., J.Z. and M.G. performed auxin accumulation in yeast and protoplasts. Y.L. modified the oestradiol-inducible vector. E.B., M.K., J.R., E.Z., J.P., M.G., J.F. and J.K.V. discussed the experimental procedures. All authors analysed and discussed the data; E.B. and J.K.V. wrote the paper and all authors saw and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10 and Supplementary References. (PDF 3594 kb)
Rights and permissions
About this article
Cite this article
Barbez, E., Kubeš, M., Rolčík, J. et al. A novel putative auxin carrier family regulates intracellular auxin homeostasis in plants. Nature 485, 119–122 (2012). https://doi.org/10.1038/nature11001
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11001
This article is cited by
-
Epigenetic modifications regulate cultivar-specific root development and metabolic adaptation to nitrogen availability in wheat
Nature Communications (2023)
-
Full-length transcriptome and metabolism revealed the difference of callus formation of tea cutting under white, red and blue light
Plant Growth Regulation (2023)
-
Auxin- and pH-induced guttation in Phycomyces sporangiophores: relation between guttation and diminished elongation growth
Protoplasma (2023)
-
The Comparison of Temporal Transcriptome Changes Between Morning-Opening and Afternoon-Opening Iris Flowers Reveals the Candidate Genes Regulating Flower Opening and Closing
Journal of Plant Biology (2023)
-
Osa-miR1861e Modulates Sodium Toxicity Responses in Rice Immature Grains via OsGST and OsPILS7b
Journal of Plant Growth Regulation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.