Abstract
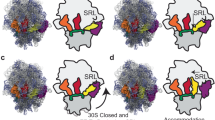
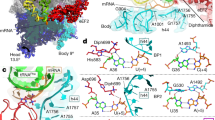
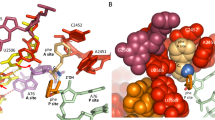
During protein synthesis, the ribosome accurately selects transfer RNAs (tRNAs) in accordance with the messenger RNA (mRNA) triplet in the decoding centre. tRNA selection is initiated by elongation factor Tu, which delivers tRNA to the aminoacyl tRNA-binding site (A site) and hydrolyses GTP upon establishing codon–anticodon interactions in the decoding centre1,2,3,4,5,6,7,8,9. At the following proofreading step the ribosome re-examines the tRNA and rejects it if it does not match the A codon2,3,10,11,12,13,14. It was suggested that universally conserved G530, A1492 and A1493 of 16S ribosomal RNA, critical for tRNA binding in the A site15,16,17, actively monitor cognate tRNA18, and that recognition of the correct codon–anticodon duplex induces an overall ribosome conformational change (domain closure)19. Here we propose an integrated mechanism for decoding based on six X-ray structures of the 70S ribosome determined at 3.1–3.4 Å resolution, modelling cognate or near-cognate states of the decoding centre at the proofreading step. We show that the 30S subunit undergoes an identical domain closure upon binding of either cognate or near-cognate tRNA. This conformational change of the 30S subunit forms a decoding centre that constrains the mRNA in such a way that the first two nucleotides of the A codon are limited to form Watson–Crick base pairs. When U·G and G·U mismatches, generally considered to form wobble base pairs, are at the first or second codon–anticodon position, the decoding centre forces this pair to adopt the geometry close to that of a canonical C·G pair. This by itself, or with distortions in the codon–anticodon mini-helix and the anticodon loop, causes the near-cognate tRNA to dissociate from the ribosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The atomic coordinates and structure factors for the determined crystal structures are deposited in the Protein Data Bank under accession numbers 3TVF and 3TVE (cognate tRNA2 Leu complex), 3UYE and 3UYD (near-cognate tRNA2 Leu complex), 3UZ3 and3UZ1 (near-cognate tRNA2 Leu complex with paromomycin),3UZ6 and 3UZ9 (cognate tRNATyr complex), 3UZG and 3UZF (near-cognate tRNATyr complex), and 3UZL and 3UZK (near-cognate tRNATyr complex with paromomycin).
References
Rodnina, M. V. & Wintermeyer, W. Fidelity of aminoacyl-tRNA selection on the ribosome: kinetic and structural mechanisms. Annu. Rev. Biochem. 70, 415–435 (2001)
Rodnina, M. V., Gromadski, K. B., Kothe, U. & Wieden, H. J. Recognition and selection of tRNA in translation. FEBS Lett. 579, 938–942 (2005)
Zaher, H. S. & Green, R. Fidelity at the molecular level: lessons from protein synthesis. Cell 136, 746–762 (2009)
Gromadski, K. B., Daviter, T. & Rodnina, M. V. A uniform response to mismatches in codon-anticodon complexes ensures ribosomal fidelity. Mol. Cell 21, 369–377 (2006)
Lee, T. H., Blanchard, S. C., Kim, H. D., Puglisi, J. D. & Chu, S. The role of fluctuations in tRNA selection by the ribosome. Proc. Natl Acad. Sci. USA 104, 13661–13665 (2007)
Li, W. et al. Recognition of aminoacyl-tRNA: a common molecular mechanism revealed by cryo-EM. EMBO J. 27, 3322–3331 (2008)
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009)
Schuette, J. C. et al. GTPase activation of elongation factor EF-Tu by the ribosome during decoding. EMBO J. 28, 755–765 (2009)
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010)
Ehrenberg, M., Kurland, C. G. & Ruusala, T. Counting cycles of EF-Tu to measure proofreading in translation. Biochimie 68, 261–273 (1986)
Voorhees, R. M., Weixlbaumer, A., Loakes, D., Kelley, A. C. & Ramakrishnan, V. Insights into substrate stabilization from snapshots of the peptidyl transferase center of the intact 70S ribosome. Nature Struct. Mol. Biol. 16, 528–533 (2009)
Jenner, L., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural rearrangements of the ribosome at the tRNA proofreading step. Nature Struct. Mol. Biol. 17, 1072–1078 (2010)
Geggier, P. et al. Conformational sampling of aminoacyl-tRNA during selection on the bacterial ribosome. J. Mol. Biol. 399, 576–595 (2010)
Whitford, P. C. et al. Accommodation of aminoacyl-tRNA into the ribosome involves reversible excursions along multiple pathways. RNA 16, 1196–1204 (2010)
Moazed, D. & Noller, H. F. Binding of tRNA to the ribosomal A and P sites protects two distinct sets of nucleotides in 16 S rRNA. J. Mol. Biol. 211, 135–145 (1990)
Powers, T. & Noller, H. F. Selective perturbation of G530 of 16 S rRNA by translational miscoding agents and a streptomycin-dependence mutation in protein S12. J. Mol. Biol. 235, 156–172 (1994)
Yoshizawa, S., Fourmy, D. & Puglisi, J. D. Recognition of the codon–anticodon helix by ribosomal RNA. Science 285, 1722–1725 (1999)
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001)
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
Jenner, L. B., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nature Struct. Mol. Biol. 17, 555–560 (2010)
BPS Database of RNA Base-pair Structures . http://bps.rutgers.edu/bps (2008)
Bebenek, K., Pedersen, L. C. & Kunkel, T. A. Replication infidelity via a mismatch with Watson–Crick geometry. Proc. Natl Acad. Sci. USA 108, 1862–1867 (2011)
Auffinger, P. & Westhof, E. An extended structural signature for the tRNA anticodon loop. RNA 7, 334–341 (2001)
Pape, T., Wintermeyer, W. & Rodnina, M. V. Conformational switch in the decoding region of 16S rRNA during aminoacyl-tRNA selection on the ribosome. Nature Struct. Biol. 7, 104–107 (2000)
Cochella, L. & Green, R. An active role for tRNA in decoding beyond codon:anticodon pairing. Science 308, 1178–1180 (2005)
Dale, T. & Uhlenbeck, O. C. Amino acid specificity in translation. Trends Biochem. Sci. 30, 659–665 (2005)
Schmeing, T. M., Voorhees, R. M., Kelley, A. C. & Ramakrishnan, V. How mutations in tRNA distant from the anticodon affect the fidelity of decoding. Nature Struct. Mol. Biol. 18, 432–436 (2011)
Mueller, M., Wang, M. & Schulze-Briese, C. Optimal fine φ-slicing for single-photon-counting pixel detectors. Acta Crystallogr. D 68, 42–56 (2012)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Praaenikar, J., Afonine, P. V., Guncar, G., Adams, P. D. & Turk, D. Averaged kick maps: less noise, more signal and probably less bias. Acta Crystallogr. D 65, 921–931 (2009)
Acknowledgements
We are grateful to C. Schulze-Briese and the staff at the Swiss Light Source (Switzerland) for help during synchrotron X-ray data collection. We thank S. Duclaud for ribosome preparation and the staff of the Structural Biology Department core facility at Institut de Génétique et de Biologie Moléculaire et Cellulaire, Université de Strasbourg. This work was supported by ANR BLAN07-3_190451 (to M.Y.), ANR-07-PCVI-0015-01 (to G.Y.), Fondation pour la Recherche Médicale en France (to N.D.) and by the European Commission SPINE2.
Author information
Authors and Affiliations
Contributions
N.D. and L.J. conducted experiments and performed analysis. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11, Supplementary Tables 1-3 and additional references. (PDF 5357 kb)
Supplementary Movie 1
This file contains an animation showing deformation of the codon-anticodon mini-helix and the A site tRNA anticodon loop upon transition from cognate to near-cognate states. The models are colored as follows: the mRNA codon is in yellow and the tRNA anticodon loop is in red. See the main text and Fig. 2d. (MOV 89 kb)
Supplementary Movie 2
This file contains an animation showing changes in the decoding pocket and its environment upon binding of aminoglycoside antibiotic paromomycin (PAR). The comparison is shown for the tRNATyr near-cognate state. The color code is the following: near-cognate tRNATyr, - red; mRNA, - yellow; PAR, - green; ‘530’ loop and helix 44 of 16S rRNA, - aquamarine; helix 69 of 23S rRNA, - grey. Refer to the main text and Fig. 2e. (MOV 164 kb)
Rights and permissions
About this article
Cite this article
Demeshkina, N., Jenner, L., Westhof, E. et al. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012). https://doi.org/10.1038/nature10913
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10913
This article is cited by
-
mRNA reading frame maintenance during eukaryotic ribosome translocation
Nature (2024)
-
A unified Watson-Crick geometry drives transcription of six-letter expanded DNA alphabets by E. coli RNA polymerase
Nature Communications (2023)
-
Geometric alignment of aminoacyl-tRNA relative to catalytic centers of the ribosome underpins accurate mRNA decoding
Nature Communications (2023)
-
Modulation of translational decoding by m6A modification of mRNA
Nature Communications (2023)
-
Non-AUG translation initiation in mammals
Genome Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.