Abstract
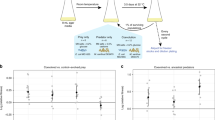
Almost all species are subject to continuous attack by parasites and pathogens. Because parasites and pathogens tend to have shorter generation times1,2 and often experience stronger selection due to interaction than their victims do3,4, it is frequently argued that they should evolve more rapidly and thus maintain an advantage in the evolutionary race between defence and counter-defence1,5. This prediction generates an apparent paradox: how do victim species survive and even thrive in the face of a continuous onslaught of more rapidly evolving enemies5? One potential explanation is that defence is physiologically, mechanically or behaviourally easier than attack, so that evolution is less constrained for victims than for parasites or pathogens6. Another possible explanation is that parasites and pathogens have enemies themselves and that victim species persist because parasites and pathogens are regulated from the top down and thus generally have only modest demographic impacts on victim populations7,8. Here we explore a third possibility: that victim species are not as evolutionarily impotent as conventional wisdom holds, but instead have unique evolutionary advantages that help to level the playing field. We use quantitative genetic analysis and individual-based simulations to show that victims can achieve such an advantage when coevolution involves multiple traits in both the host and the parasite.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hafner, M. S. et al. Disparate rates of molecular evolution in cospeciating hosts and parasites. Science 265, 1087–1090 (1994)
Page, R. D. M., Lee, P. L. M., Becher, S. A., Griffiths, R. & Clayton, D. H. A different tempo of mitochondrial DNA evolution in birds and their parasitic lice. Mol. Phylogenet. Evol. 9, 276–293 (1998)
Dowton, M. & Austin, A. D. Increased genetic diversity in mitochondrial genes is correlated with the evolution of parasitism in the hymenoptera. J. Mol. Evol. 41, 958–965 (1995)
King, K. C., Jokela, J. & Lively, C. M. Trematode parasites infect or die in snail hosts. Biol. Lett. 7, 265–268 (2011)
Hamilton, W. D., Axelrod, R. & Tanese, R. Sexual reproduction as an adaptation to resist parasites (a review). Proc. Natl Acad. Sci. USA 87, 3566–3573 (1990)
Thompson, J. N. Constraints on arms races in coevolution. Trends Ecol. Evol. 1, 105–107 (1986)
Costamagna, A. C. & Landis, D. A. Predators exert top-down control of soybean aphid across a gradient of agricultural management systems. Ecol. Appl. 16, 1619–1628 (2006)
Gomez, J. M. & Zamora, R. Top-down effects in a tritrophic system—parasitoids enhance plant fitness. Ecology 75, 1023–1030 (1994)
Abrams, P. A. The evolution of predator–prey interactions: theory and evidence. Annu. Rev. Ecol. Syst. 31, 79–105 (2000)
Frank, S. A. Evolution of host–parasite diversity. Evolution 47, 1721–1732 (1993)
Hochberg, M. E. Hide or fight? The competitive evolution of concealment and encapsulation in parasitoid–host associations. Oikos 80, 342–352 (1997)
Agrawal, A. F. & Lively, C. M. Modelling infection as a two step process combining gene-for-gene and matching allele genetics. Proc. R. Soc. Lond. B 270, 323–334 (2003)
Vermeij, G. J. Unsuccessful predation and evolution. Am. Nat. 120, 701–720 (1982)
Berenbaum, M. R., Zangerl, A. R. & Nitao, J. K. Constraints on chemical coevolution—wild parsnips and the parsnip webworm. Evolution 40, 1215–1228 (1986)
Jones, S. R. M. The occurrence and mechanisms of innate immunity against parasites in fish. Dev. Comp. Immunol. 25, 841–852 (2001)
Doebeli, M. & Ispolatov, I. Complexity and diversity. Science 328, 494–497 (2010)
Toju, H. et al. Climatic gradients of arms race coevolution. Am. Nat. 177, 562–573 (2011)
Nuismer, S. L. & Ridenhour, B. J. The contribution of parasitism to selection on floral traits in Heuchera grossulariifolia. J. Evol. Biol. 21, 958–965 (2008)
Rasmann, S., Erwin, A. C., Halitschke, R. & Agrawal, A. A. Direct and indirect root defences of milkweed (Asclepias syriaca): trophic cascades, trade-offs and novel methods for studying subterranean herbivory. J. Ecol. 99, 16–25 (2011)
Lande, R. Quantitative genetic analysis of multivariate evolution, applied to brain:body size allometry. Evolution 33, 402–416 (1979)
Lande, R. & Arnold, S. J. The measurement of selection on correlated characters. Evolution 37, 1210–1226 (1983)
Kirkpatrick, M. Patterns of quantitative genetic variation in multiple dimensions. Genetica 136, 271–284 (2009)
Arnold, S. J., Burger, R., Hohenlohe, P. A., Ajie, B. C. & Jones, A. G. Understanding the evolution and stability of the G-matrix. Evolution 62, 2451–2461 (2008)
Jones, A. G., Arnold, S. J. & Burger, R. Stability of the G-matrix in a population experiencing pleiotropic mutation, stabilizing selection, and genetic drift. Evolution 57, 1747–1760 (2003)
Jones, A. G., Arnold, S. J. & Burger, R. Evolution and stability of the G-matrix on a landscape with a moving optimum. Evolution 58, 1639–1654 (2004)
Nuismer, S. L., Doebeli, M. & Browning, D. The coevolutionary dynamics of antagonistic interactions mediated by quantitative traits with evolving variances. Evolution 59, 2073–2082 (2005)
Zangerl, A. R. & Berenbaum, M. R. Phenotype matching in wild parsnip and parsnip webworms: causes and consequences. Evolution 57, 806–815 (2003)
Zalucki, M. P., Brower, L. P. & Alonso-Mejia, A. Detrimental effects of latex and cardiac glycosides on survival and growth of first-instar monarch butterfly larvae Danaus plexippus feeding on the sandhill milkweed Asclepias humistrata. Ecol. Entomol. 26, 212–224 (2001)
Hairston, N. G., Smith, F. E. & Slobodkin, L. B. Community structure, population control, and competition. Am. Nat. 94, 421–425 (1960)
Acknowledgements
We thank A. R. Ives and F. Débarre for comments. R.T.G. and D.C.J. are postdoctoral fellows at the National Institute for Mathematical and Biological Synthesis (NIMBioS). NIMBioS is sponsored by the National Science Foundation (NSF), the US Department of Homeland Security and the US Department of Agriculture through NSF award no. EF-0832858, with support from The University of Tennessee, Knoxville. Additional support for this research was provided by NSF grants DMS 0540392 and DEB 1118947 to S.L.N.
Author information
Authors and Affiliations
Contributions
R.T.G. and S.L.N. conceived the study and derived the analytical results. R.T.G. coded and ran numerical simulations and analysed output data. R.T.G. and D.C.J. derived proofs 1–3. R.T.G. and S.L.N. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References, Supplementary Data 1-3, Supplementary Box 1, Supplementary Tables 1-6 and Supplementary Figures 1-5. (PDF 1956 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Gilman, R., Nuismer, S. & Jhwueng, DC. Coevolution in multidimensional trait space favours escape from parasites and pathogens. Nature 483, 328–330 (2012). https://doi.org/10.1038/nature10853
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10853
This article is cited by
-
Complex eco-evolutionary dynamics induced by the coevolution of predator–prey movement strategies
Evolutionary Ecology (2022)
-
How long do Red Queen dynamics survive under genetic drift? A comparative analysis of evolutionary and eco-evolutionary models
BMC Evolutionary Biology (2020)
-
Third-party mutualists have contrasting effects on host invasion under the enemy-release and biotic-resistance hypotheses
Evolutionary Ecology (2017)
-
Effects of plant evolution on nutrient cycling couple aboveground and belowground processes
Theoretical Ecology (2017)
-
Fixed prey cue preferences among Dusky Pigmy Rattlesnakes (Sistrurus miliarius barbouri) raised on different long-term diets
Evolutionary Ecology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.