Abstract
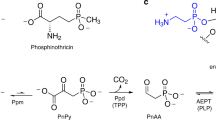
Phosphorus is an essential element for all known forms of life. In living systems, phosphorus is an integral component of nucleic acids, carbohydrates and phospholipids, where it is incorporated as a derivative of phosphate. However, most Gram-negative bacteria have the capability to use phosphonates as a nutritional source of phosphorus under conditions of phosphate starvation1. In these organisms, methylphosphonate is converted to phosphate and methane. In a formal sense, this transformation is a hydrolytic cleavage of a carbon–phosphorus (C–P) bond, but a general enzymatic mechanism for the activation and conversion of alkylphosphonates to phosphate and an alkane has not been elucidated despite much effort for more than two decades. The actual mechanism for C–P bond cleavage is likely to be a radical-based transformation2. In Escherichia coli, the catalytic machinery for the C–P lyase reaction has been localized to the phn gene cluster1. This operon consists of the 14 genes phnC, phnD, …, phnP. Genetic and biochemical experiments have demonstrated that the genes phnG, phnH, …, phnM encode proteins that are essential for the conversion of phosphonates to phosphate and that the proteins encoded by the other genes in the operon have auxiliary functions1,3,4,5,6. There are no functional annotations for any of the seven proteins considered essential for C–P bond cleavage. Here we show that methylphosphonate reacts with MgATP to form α-d-ribose-1-methylphosphonate-5-triphosphate (RPnTP) and adenine. The triphosphate moiety of RPnTP is hydrolysed to pyrophosphate and α-d-ribose-1-methylphosphonate-5-phosphate (PRPn). The C–P bond of PRPn is subsequently cleaved in a radical-based reaction producing α-d-ribose-1,2-cyclic-phosphate-5-phosphate and methane in the presence of S-adenosyl-l-methionine. Substantial quantities of phosphonates are produced worldwide for industrial processes, detergents, herbicides and pharmaceuticals7,8,9. Our elucidation of the chemical steps for the biodegradation of alkylphosphonates shows how these compounds can be metabolized and recycled to phosphate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Metcalf, W. W. & Wanner, B. L. Evidence for a fourteen-gene, phnC to phnP locus for phosphonate metabolism in Escherichia coli. Gene 129, 27–32 (1993)
Ahn, Y., Ye, Q., Cho, H., Walsh, C. T. & Floss, H. G. Stereochemistry of carbon-phosphorus cleavage in ethylphosphonate catalyzed by C-P lyase from Escherichia coli. J. Am. Chem. Soc. 114, 7953–7954 (1992)
Metcalf, W. W. & Wanner, B. L. Mutational analysis of an Escherichia coli fourteen-gene operon for phosphonate degradation, using TnphoA’ elements. J. Bacteriol. 175, 3430–3442 (1993)
Hove-Jensen, B., Rosenkrantz, T. J., Haldimann, A. & Wanner, B. L. Escherichia coli phnN, encoding ribose 1,5-bisphosphokinase activity (phosphoribosyl diphosphate forming): dual role in phosphonate degradation and NAD biosynthesis pathways. J. Bacteriol. 185, 2793–2801 (2003)
Errey, J. C. & Blanchard, J. S. Functional annotation and kinetic characterization of PhnO from Salmonella enterica. Biochemistry 45, 3033–3039 (2006)
Hove-Jensen, B., McSorley, F. R. & Zechel, D. L. Physiological role of phnP-specified phosphoribosyl cyclic phosphodiesterase in catabolism of organophosphoric acids by the carbon-phosphorus lyase pathway. J. Am. Chem. Soc. 133, 3617–3624 (2011)
Ternan, N. G., McGrath, J. W., McMullan, G. & Quinn, J. P. Organophosphonates: occurrence, synthesis and biodegradation by microorganisms. World J. Microbiol. Biotechnol. 14, 635–647 (1998)
Kononova, S. V. & Nesmeyanova, M. A. Phosphonates and their degradation by microorganisms. Biochemistry (Mosc.) 67, 184–195 (2002)
White, A. K. & Metcalf, W. W. Microbial metabolism of reduced phosphorus compounds. Annu. Rev. Microbiol. 61, 379–400 (2007)
Avila, L. Z., Draths, K. M. & Frost, J. W. Metabolites associated with organophosphonate C-P bond cleavage: chemical synthesis and microbial degradation of [32P]-ethylphosphonic acid. Bioorg. Med. Chem. Lett. 1, 51–54 (1991)
Frost, J. W., Loo, S., Cordeiro, M. L. & Li, D. Radical-based dephosphorylation and organophosphonate biodegradation. J. Am. Chem. Soc. 109, 2166–2171 (1987)
Parker, G. F., Higgins, T. P., Hawkes, T. & Robson, R. L. Rhizobium (Sinorhizobium) meliloti phn genes: characterization and identification of their protein products. J. Bacteriol. 181, 389–395 (1999)
Kamat, S. S. et al. Catalytic mechanism and three-dimensional structure of adenine deaminase. Biochemistry 50, 1917–1927 (2011)
Maynes, J. T., Yuan, R. G. & Snyder, F. F. Identification, expression and characterization of the Escherichia coli guanine deaminase. J. Bacteriol. 182, 4658–4660 (2000)
Hall, R. S. et al. Three dimensional structure and catalytic mechanism of cytosine deaminase. Biochemistry 50, 5077–5085 (2011)
Krenitsky, T. A., Neil, S. M., Elion, G. B. & Hitchings, G. H. A comparison of the specificities of xanthine oxidase and aldehyde oxidase. Arch. Biochem. Biophys. 150, 585–599 (1972)
La . Vallie, E. R., McCoy, J. M., Smith, D. B. & Riggs, P . Enzymatic and chemical cleavage of fusion proteins. Curr. Protocols Mol. Biol. Unit 16.4B. (1994)
Jochimsen, B. et al. Five phosphonate operon gene products as components of a multi-enzyme complex of the carbon-phosphorus lyase pathway. Proc. Natl Acad. Sci. USA 108, 11393–11398 (2011)
Seibert, C. M. &. Raushel, F. M. Structural and catalytic diversity within the amidohydrolase superfamily. Biochemistry 44, 6383–6391 (2005)
Frey, P. A., Hegeman, A. D. & Ruzicka, F. J. The radical SAM superfamily. Crit. Rev. Biochem. Mol. Biol. 43, 63–88 (2008)
Chatterjee, A. et al. Reconstitution of ThiC in thiamine pyrimidine biosynthesis expands the radical SAM superfamily. Nature Chem. Biol. 4, 758–765 (2008)
McGlynn, S. E. et al. Identification and characterization of a novel member of the radical AdoMet enzyme superfamily and implications for the biosynthesis of the Hmd hydrogenase active site cofactor. J. Bacteriol. 192, 595–598 (2010)
Zhang, Y. et al. Diphthamide biosynthesis requires an organic radical generated by iron-sulphur enzyme. Nature 465, 891–896 (2010)
Parast, C. V., Wong, K. K., Lewisch, S. A. & Kozarich, J. W. Hydrogen exchange of the glycyl radical of pyruvate formate-lyase is catalyzed by cysteine 419. Biochemistry 34, 2393–2399 (1995)
Buis, J. M. & Broderick, J. B. Pyruvate formate-lyase activating enzyme: elucidation of a novel mechanism for glycyl radical formation. Arch. Biochem. Biophys. 433, 288–296 (2005)
Thauer, R. K. & Shima, S. Methane as fuel for anaerobic microorganisms. Ann. NY Acad. Sci. 1125, 158–170 (2008)
Acknowledgements
We thank D. Barondeau and his laboratory for use of the anaerobic chamber and for advice on the assembly of Fe–S clusters in radical SAM enzymes. We also thank C. Xu for help with some of the 31P NMR spectra and S. Burrows for help with cloning phnI and phnH. This work was supported in part by the National Institutes of Health (GM93342, GM71790) and the Robert A. Welch Foundation (A-840). The cryoprobe for the NMR spectrometer was purchased with funds from the National Science Foundation (0840464)
Author information
Authors and Affiliations
Contributions
S.S.K., H.J.W. and F.M.R. designed the experiments. S.S.K. and H.J.W. performed the experiments. S.S.K., H.J.W. and F.M.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-24 with legends, Supplementary Table 1, Supplementary Text and additional references. (PDF 1937 kb)
Rights and permissions
About this article
Cite this article
Kamat, S., Williams, H. & Raushel, F. Intermediates in the transformation of phosphonates to phosphate by bacteria. Nature 480, 570–573 (2011). https://doi.org/10.1038/nature10622
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10622
This article is cited by
-
The facilitating role of phycospheric heterotrophic bacteria in cyanobacterial phosphonate availability and Microcystis bloom maintenance
Microbiome (2023)
-
Methylphosphonate-driven methane formation and its link to primary production in the oligotrophic North Atlantic
Nature Communications (2023)
-
Structural remodelling of the carbon–phosphorus lyase machinery by a dual ABC ATPase
Nature Communications (2023)
-
Integration of Microbial Transformation Mechanism of Polyphosphate Accumulation and Sulfur Cycle in Subtropical Marine Mangrove Ecosystems with Spartina alterniflora Invasion
Microbial Ecology (2023)
-
Global and seasonal variation of marine phosphonate metabolism
The ISME Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.