Abstract
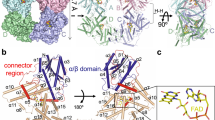
The cryptochrome/photolyase (CRY/PL) family of photoreceptors mediates adaptive responses to ultraviolet and blue light exposure in all kingdoms of life1,2,3,4,5. Whereas PLs function predominantly in DNA repair of cyclobutane pyrimidine dimers (CPDs) and 6-4 photolesions caused by ultraviolet radiation, CRYs transduce signals important for growth, development, magnetosensitivity and circadian clocks1,2,3,4,5. Despite these diverse functions, PLs/CRYs preserve a common structural fold, a dependence on flavin adenine dinucleotide (FAD) and an internal photoactivation mechanism3,6. However, members of the CRY/PL family differ in the substrates recognized (protein or DNA), photochemical reactions catalysed and involvement of an antenna cofactor. It is largely unknown how the animal CRYs that regulate circadian rhythms act on their substrates. CRYs contain a variable carboxy-terminal tail that appends the conserved PL homology domain (PHD) and is important for function7,8,9,10,11,12. Here, we report a 2.3-Å resolution crystal structure of Drosophila CRY with an intact C terminus. The C-terminal helix docks in the analogous groove that binds DNA substrates in PLs. Conserved Trp 536 juts into the CRY catalytic centre to mimic PL recognition of DNA photolesions. The FAD anionic semiquinone found in the crystals assumes a conformation to facilitate restructuring of the tail helix. These results help reconcile the diverse functions of the CRY/PL family by demonstrating how conserved protein architecture and photochemistry can be elaborated into a range of light-driven functions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cashmore, A. R. Cryptochromes: enabling plants and animals to determine circadian time. Cell 114, 537–543 (2003)
Sancar, A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 103, 2203–2238 (2003)
Partch, C. L. & Sancar, A. Photochemistry and photobiology of cryptochrome blue-light photopigments: the search for a photocycle. Photochem. Photobiol. 81, 1291–1304 (2005)
Gegear, R. J., Casselman, A., Waddell, S. & Reppert, S. M. Cryptochrome mediates light-dependent magnetosensitivity in Drosophila. Nature 454, 1014–1018 (2008)
Yoshii, T., Ahmad, M. & Helfrich-Forster, C. Cryptochrome mediates light-dependent magnetosensitivity of Drosophila’s circadian clock. PLoS Biol. 7, e1000086 (2009)
Hitomi, K. et al. Functional motifs in the (6-4) photolyase crystal structure make a comparative framework for DNA repair photolyases and clock cryptochromes. Proc. Natl Acad. Sci. USA 106, 6962–6967 (2009)
Busza, A., Emery-Le, M., Rosbash, M. & Emery, P. Roles of the two Drosophila CRYPTOCHROME structural domains in circadian photoreception. Science 304, 1503–1506 (2004)
Dissel, S. et al. A constitutively active cryptochrome in Drosophila melanogaster. Nature Neurosci. 7, 834–840 (2004)
Hemsley, M. J. et al. Linear motifs in the C-terminus of D. melanogaster cryptochrome. Biochem. Biophys. Res. Commun. 355, 531–537 (2007)
Rosato, E. et al. Light-dependent interaction between Drosophila CRY and the clock protein PER mediated by the carboxy terminus of CRY. Curr. Biol. 11, 909–917 (2001)
Partch, C. L., Clarkson, M. W., Ozgur, S., Lee, A. L. & Sancar, A. Role of structural plasticity in signal transduction by the cryptochrome blue-light photoreceptor. Biochemistry 44, 3795–3805 (2005)
van der Schalie, E. A., Conte, F. E., Marz, K. E. & Green, C. B. Structure/function analysis of Xenopus cryptochromes 1 and 2 reveals differential nuclear localization mechanisms and functional domains important for interaction with and repression of CLOCK-BMAL1. Mol. Cell. Biol. 27, 2120–2129 (2007)
Ozturk, N., Song, S. H., Selby, C. P. & Sancar, A. Animal type 1 cryptochromes. Analysis of the redox state of the flavin cofactor by site-directed mutagenesis. J. Biol. Chem. 283, 3256–3263 (2008)
Yuan, Q., Metterville, D., Briscoe, A. D. & Reppert, S. M. Insect cryptochromes: gene duplication and loss define diverse ways to construct insect circadian clocks. Mol. Biol. Evol. 24, 948–955 (2007)
Griffin, E. A., Jr, Staknis, D. & Weitz, C. J. Light-independent role of CRY1 and CRY2 in the mammalian circadian clock. Science 286, 768–771 (1999)
Koh, K., Zheng, X. Z. & Sehgal, A. JETLAG resets the Drosophila circadian clock by promoting light-induced degradation of TIMELESS. Science 312, 1809–1812 (2006)
Peschel, N., Chen, K. F., Szabo, G. & Stanewsky, R. Light-dependent interactions between the Drosophila circadian clock factors cryptochrome, jetlag, and timeless. Curr. Biol. 19, 241–247 (2009)
Ozturk, N., Selby, C. P., Annayev, Y., Zhong, D. & Sancar, A. Reaction mechanism of Drosophila cryptochrome. Proc. Natl Acad. Sci. USA 108, 516–521 (2011)
VanVickle-Chavez, S. J. & Van Gelder, R. N. Action spectrum of Drosophila cryptochrome. J. Biol. Chem. 282, 10561–10566 (2007)
Hoang, N. et al. Human and Drosophila cryptochromes are light activated by flavin photoreduction in living cells. PLoS Biol. 6, e160 (2008)
Berndt, A. et al. A novel photoreaction mechanism for the circadian blue light photoreceptor Drosophila cryptochrome. J. Biol. Chem. 282, 13011–13021 (2007)
Ceriani, M. F. et al. Light-dependent sequestration of TIMELESS by CRYPTOCHROME. Science 285, 553–556 (1999)
Maul, M. J. et al. Crystal structure and mechanism of a DNA (6-4) photolyase. Angew. Chem. Int. Edn Engl. 47, 10076–10080 (2008)
Kao, Y. T. et al. Ultrafast dynamics and anionic active states of the flavin cofactor in cryptochrome and photolyase. J. Am. Chem. Soc. 130, 7695–7701 (2008)
Gegear, R. J., Foley, L. E., Casselman, A. & Reppert, S. M. Animal cryptochromes mediate magnetoreception by an unconventional photochemical mechanism. Nature 463, 804–807 (2010)
Senda, T., Senda, M., Kimura, S. & Ishida, T. Redox control of protein conformation in flavoproteins. Antioxid. Redox Signal. 11, 1741–1766 (2009)
Rohr, A. K., Hersleth, H. P. & Andersson, K. K. Tracking flavin conformations in protein crystal structures with Raman spectroscopy and QM/MM calculations. Angew. Chem. Int. Ed. 49, 2324–2327 (2010)
Schleicher, E. et al. Electron nuclear double resonance differentiates complementary roles for active site histidines in (6-4) photolyase. J. Biol. Chem. 282, 4738–4747 (2007)
Tang, C. H. A., Hinteregger, E., Shang, Y. H. & Rosbash, M. Light-mediated TIM degradation within Drosophila pacemaker neurons (s-LNvs) is neither necessary nor sufficient for delay zone phase shifts. Neuron 66, 378–385 (2010)
Fogle, K. J., Parson, K. G., Dahm, N. A. & Holmes, T. C. CRYPTOCHROME is a blue-light sensor that regulates neuronal firing rate. Science 331, 1409–1413 (2011)
Bilwes, A. M., Quezada, C. M., Croal, L. R., Crane, B. R. & Simon, M. I. Nucleotide binding by the histidine kinase CheA. Nature Struct. Biol. 8, 353–360 (2001)
Acknowledgements
This study was supported by NIH Grant GM079679 to B.R.C. and GM054339 to M.W.Y. We thank the NE-CAT at the Advanced Photon Source of Argonne Laboratories for access to data collection facilities. We are indebted to C. Kemp for insect cell expression of dCRY and C. Manahan, X. Xu and W. Horne for their help with the ITC experiments.
Author information
Authors and Affiliations
Contributions
B.R.C., B.D.Z., A.T.V., D.T. and M.W.Y. designed the project. J.W. cloned, expressed and purified dCRY, B.D.Z. and A.T.V. purified and crystallized dCRY and collected diffraction data. B.D.Z. and B.R.C. determined the structure. D.T. and M.W.Y. performed CRY stability studies. A.T.V. performed ITC experiments. B.D.Z. and B.R.C. wrote the manuscript and all authors provided editorial input.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Figures 1-12 with legends, Supplementary Tables 1-2, Supplementary Materials and Methods and additional references (see contents for details). (PDF 1718 kb)
Rights and permissions
About this article
Cite this article
Zoltowski, B., Vaidya, A., Top, D. et al. Structure of full-length Drosophila cryptochrome. Nature 480, 396–399 (2011). https://doi.org/10.1038/nature10618
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10618
This article is cited by
-
Essential elements of radical pair magnetosensitivity in Drosophila
Nature (2023)
-
Effects of inter-radical interactions and scavenging radicals on magnetosensitivity: spin dynamics simulations of proposed radical pairs
European Biophysics Journal (2023)
-
Direct experimental observation of blue-light-induced conformational change and intermolecular interactions of cryptochrome
Communications Biology (2022)
-
CRY2 interacts with CIS1 to regulate thermosensory flowering via FLM alternative splicing
Nature Communications (2022)
-
Tuning flavin environment to detect and control light-induced conformational switching in Drosophila cryptochrome
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.