Abstract
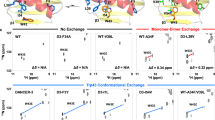
Proteins are inherently plastic molecules, whose function often critically depends on excursions between different molecular conformations (conformers)1,2,3. However, a rigorous understanding of the relation between a protein’s structure, dynamics and function remains elusive. This is because many of the conformers on its energy landscape are only transiently formed and marginally populated (less than a few per cent of the total number of molecules), so that they cannot be individually characterized by most biophysical tools. Here we study a lysozyme mutant from phage T4 that binds hydrophobic molecules4 and populates an excited state transiently (about 1 ms) to about 3% at 25 °C (ref. 5). We show that such binding occurs only via the ground state, and present the atomic-level model of the ‘invisible’, excited state obtained using a combined strategy of relaxation-dispersion NMR (ref. 6) and CS-Rosetta7 model building that rationalizes this observation. The model was tested using structure-based design calculations identifying point mutants predicted to stabilize the excited state relative to the ground state. In this way a pair of mutations were introduced, inverting the relative populations of the ground and excited states and altering function. Our results suggest a mechanism for the evolution of a protein’s function by changing the delicate balance between the states on its energy landscape. More generally, they show that our approach can generate and validate models of excited protein states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Karplus, M. & Kuriyan, J. Molecular dynamics and protein function. Proc. Natl Acad. Sci. USA 102, 6679–6685 (2005)
Boehr, D. D., McElheny, D., Dyson, H. J. & Wright, P. E. The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313, 1638–1642 (2006)
Henzler-Wildman, K. A. et al. Intrinsic motions along an enzymatic reaction trajectory. Nature 450, 838–844 (2007)
Eriksson, A. E., Baase, W. A., Wozniak, J. A. & Matthews, B. W. A cavity-containing mutant of T4 lysozyme is stabilized by buried benzene. Nature 355, 371–373 (1992)
Mulder, F. A. A., Mittermaier, A., Hon, B., Dahlquist, F. W. & Kay, L. E. Studying excited states of protein by NMR spectroscopy. Nature Struct. Biol. 8, 932–935 (2001)
Palmer, A. G., Kroenke, C. D. & Loria, J. P. NMR methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol. 339, 204–238 (2001)
Shen, Y. et al. Consistent blind protein structure generation from NMR chemical shift data. Proc. Natl Acad. Sci. USA 105, 4685–4690 (2008)
Baase, W. A., Liu, L., Tronrud, D. E. & Matthews, B. W. Lessons from the lysozyme of phage T4. Protein Sci. 19, 631–641 (2010)
Eriksson, A. E. et al. Response of a protein structure to cavity-creating mutations and its relation to the hydrophobic effect. Science 255, 178–183 (1992)
Korzhnev, D. M. & Kay, L. E. Probing invisible, low-populated states of protein molecules by relaxation dispersion NMR spectroscopy: an application to protein folding. Acc. Chem. Res. 41, 442–451 (2008)
Hansen, D. F., Vallurupalli, P. & Kay, L. E. Using relaxation dispersion NMR spectroscopy to determine structures of excited, invisible protein states. J. Biomol. NMR 41, 113–120 (2008)
Cavalli, A., Salvatella, X., Dobson, C. M. & Vendruscolo, M. Protein structure determination from NMR chemical shifts. Proc. Natl Acad. Sci. USA 104, 9615–9620 (2007)
Vallurupalli, P., Hansen, D. F. & Kay, L. E. Structures of invisible, excited protein states by relaxation dispersion NMR spectroscopy. Proc. Natl Acad. Sci. USA 105, 11766–11771 (2008)
Korzhnev, D. M., Religa, T. L., Banachewicz, W., Fersht, A. R. & Kay, L. E. A transient and low-populated protein-folding intermediate at atomic resolution. Science 329, 1312–1316 (2010)
Berjanskii, M. V. & Wishart, D. S. A simple method to predict protein flexibility using secondary chemical shifts. J. Am. Chem. Soc. 127, 14970–14971 (2005)
Wang, C., Bradley, P. & Baker, D. Protein-protein docking with backbone flexibility. J. Mol. Biol. 373, 503–519 (2007)
Fersht, A. Structure and Mechanism in Protein Science (Freeman and Company, 1999)
Montelione, G. T. & Wagner, G. 2D chemical-exchange NMR-spectroscopy by proton detected heteronuclear correlation. J. Am. Chem. Soc. 111, 3096–3098 (1989)
Farrow, N. A., Zhang, O., Forman-Kay, J. D. & Kay, L. E. A heteronuclear correlation experiment for simultaneous determination of 15N longitudinal decay and chemical exchange rates of systems in slow equilibrium. J. Biomol. NMR 4, 727–734 (1994)
Feher, V. A., Baldwin, E. P. & Dahlquist, F. W. Access of ligands to cavities within the core of a protein is rapid. Nature Struct. Biol. 3, 516–521 (1996)
Religa, T. L., Sprangers, R. & Kay, L. E. Dynamic regulation of archaeal proteasome gate opening as studied by TROSY NMR. Science 328, 98–102 (2010)
Hu, J. S., Grzesiek, S. & Bax, A. Two-dimensional NMR methods for determining χ1 angles of aromatic residues in proteins from three-bond JC’Cγ and JNCγ couplings. J. Am. Chem. Soc. 119, 1803–1804 (1997)
Nicholson, H., Tronrud, D. E., Becktel, W. J. & Matthews, B. W. Analysis of the effectiveness of proline substitutions and glycine replacements in increasing the stability of phage T4 lysozyme. Biopolymers 32, 1431–1441 (1992)
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009)
Jensen, R. A. Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol. 30, 409–425 (1976)
Walden, W. E. et al. Structure of dual function iron regulatory protein 1 complexed with ferritin IRE-RNA. Science 314, 1903–1908 (2006)
Tang, C., Schwieters, C. D. & Clore, G. M. Open-to-closed transition in apo maltose-binding protein observed by paramagnetic NMR. Nature 449, 1078–1082 (2007)
Liu, L. J., Baase, W. A. & Matthews, B. W. Halogenated benzenes bound within a non-polar cavity in T4 lysozyme provide examples of I···S and I···Se halogen-bonding. J. Mol. Biol. 385, 595–605 (2009)
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J. Biomol. NMR 44, 213–223 (2009)
Rodríguez, J. C., Jennings, P. A. & Melacini, G. Using chemical exchange to assign non-covalent protein complexes in slow exchange with the free state: enhanced resolution and efficient signal editing. J. Biomol. NMR 30, 155–161 (2004)
Lundström, P., Vallurupalli, P., Hansen, D. F. & Kay, L. E. Isotope labeling methods for studies of excited protein states by relaxation dispersion NMR spectroscopy. Nature Protocols 4, 1641–1648 (2009)
Vallurupalli, P., Hansen, D. F., Lundstrom, P. & Kay, L. E. CPMG relaxation dispersion NMR experiments measuring glycine 1Hα and 13Cα chemical shifts in the 'invisible' excited states of proteins. J. Biomol. NMR 45, 45–55 (2009)
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999)
Korzhnev, D. M. et al. Low-populated folding intermediates of Fyn SH3 characterized by relaxation dispersion NMR. Nature 430, 586–590 (2004)
Vallurupalli, P., Hansen, D. F., Stollar, E., Meirovitch, E. & Kay, L. E. Measurement of bond vector orientations in invisible excited states of proteins. Proc. Natl Acad. Sci. USA 104, 18473–18477 (2007)
Hansen, D. F., Vallurupalli, P., Lundstrom, P., Neudecker, P. & Kay, L. E. Probing chemical shifts of invisible states of proteins with relaxation dispersion NMR spectroscopy: how well can we do? J. Am. Chem. Soc. 130, 2667–2675 (2008)
Skrynnikov, N. R., Dahlquist, F. W. & Kay, L. E. Reconstructing NMR spectra of “invisible” excited protein states using HSQC and HMQC experiments. J. Am. Chem. Soc. 124, 12352–12360 (2002)
Bouvignies, G. et al. A simple method for measuring signs of 1HN chemical shift differences between ground and excited protein states. J. Biomol. NMR 47, 135–141 (2010)
Orekhov, V. Y., Korzhnev, D. M. & Kay, L. E. Double- and zero-quantum NMR relaxation dispersion experiments sampling millisecond time scale dynamics in proteins. J. Am. Chem. Soc. 126, 1886–1891 (2004)
Hu, J. S. & Bax, A. Determination of ϕ and χ1 angles in proteins from 13C-13C three-bond J couplings measured by three-dimensional heteronuclear NMR. How planar is the peptide bond? J. Am. Chem. Soc. 119, 6360–6368 (1997)
Delaglio, F. et al. NMRpipe – a multidimensional spectral processing system based on Unix pipes. J. Biomol. NMR 6, 277–293 (1995)
Goddard, T. D. & Kneller, D. G. SPARKY 3 (University of California, San Francisco, 2006)
Vranken, W. F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct. Funct. Bioinform. 59, 687–696 (2005)
Mulder, F. A., Hon, B., Muhandiram, D. R., Dahlquist, F. W. & Kay, L. E. Flexibility and ligand exchange in a buried cavity mutant of T4 lysozyme studied by multinuclear NMR. Biochemistry 39, 12614–12622 (2000)
McConnell, H. M. Reaction rates by nuclear magnetic resonance. J. Chem. Phys. 28, 430–431 (1958)
Sprangers, R., Gribun, A., Hwang, P. M., Houry, W. A. & Kay, L. E. Quantitative NMR spectroscopy of supramolecular complexes: dynamic side pores in ClpP are important for product release. Proc. Natl Acad. Sci. USA 102, 16678–16683 (2005)
Press, W. H., Flannery, B. P., Teukolsky, S. A. & Vetterling, W. T. Numerical Recipes in C. The Art of Scientific Computing 2nd edn (Cambridge Univ. Press, 1992)
Loken, C. et al. SciNet: Lessons learned from building a power-efficient top-20 system and data centre. J. Phys. Conf. Ser. 256, 012026 (2010)
The PyMOL Molecular Graphics System, Version 1.3 (Schrödinger, LLC, 2010)
Pettersen, E. F. et al. UCSF Chimera – a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Kellogg, E. H., Leaver-Fay, A. & Baker, D. Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins 79, 830–838 (2011)
Acknowledgements
We thank J. Forman-Kay for providing laboratory space, and J. Forman-Kay and T. Alber for discussions. G.B. acknowledges The European Molecular Biology Organization and The Canadian Institutes of Health Research (CIHR) for postdoctoral fellowships. L.E.K. holds a Canada Research Chair in Biochemistry. This work was supported by the CIHR and the Natural Sciences and Engineering Research Council of Canada, with computations performed on the GPC supercomputer at the SciNet HPC Consortium.
Author information
Authors and Affiliations
Contributions
G.B., P.V. and D.F.H. made samples. G.B., P.V., D.F.H. and L.E.K. designed and performed all NMR experiments. G.B., P.V., B.E.C., O.L. and R.M.V. performed structure calculations. G.B., P.V. and A.B. carried out initial crystallization trials. F.W.D, D.B., G.B., P.V. and L.E.K. designed experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-10 with legends, Supplementary Tables 1-9 and additional references. (PDF 4402 kb)
Rights and permissions
About this article
Cite this article
Bouvignies, G., Vallurupalli, P., Hansen, D. et al. Solution structure of a minor and transiently formed state of a T4 lysozyme mutant. Nature 477, 111–114 (2011). https://doi.org/10.1038/nature10349
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10349
This article is cited by
-
Beyond slow two-state protein conformational exchange using CEST: applications to three-state protein interconversion on the millisecond timescale
Journal of Biomolecular NMR (2024)
-
Application of High-Pressure Electron Paramagnetic Resonance (EPR) Spectroscopy in Protein Science
Applied Magnetic Resonance (2024)
-
A litmus test for classifying recognition mechanisms of transiently binding proteins
Nature Communications (2022)
-
Towards autonomous analysis of chemical exchange saturation transfer experiments using deep neural networks
Journal of Biomolecular NMR (2022)
-
FID-Net: A versatile deep neural network architecture for NMR spectral reconstruction and virtual decoupling
Journal of Biomolecular NMR (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.