Abstract
Atlantic cod (Gadus morhua) is a large, cold-adapted teleost that sustains long-standing commercial fisheries and incipient aquaculture1,2. Here we present the genome sequence of Atlantic cod, showing evidence for complex thermal adaptations in its haemoglobin gene cluster and an unusual immune architecture compared to other sequenced vertebrates. The genome assembly was obtained exclusively by 454 sequencing of shotgun and paired-end libraries, and automated annotation identified 22,154 genes. The major histocompatibility complex (MHC) II is a conserved feature of the adaptive immune system of jawed vertebrates3,4, but we show that Atlantic cod has lost the genes for MHC II, CD4 and invariant chain (Ii) that are essential for the function of this pathway. Nevertheless, Atlantic cod is not exceptionally susceptible to disease under natural conditions5. We find a highly expanded number of MHC I genes and a unique composition of its Toll-like receptor (TLR) families. This indicates how the Atlantic cod immune system has evolved compensatory mechanisms in both adaptive and innate immunity in the absence of MHC II. These observations affect fundamental assumptions about the evolution of the adaptive immune system and its components in vertebrates.
Similar content being viewed by others
Main
We sequenced the genome of a heterozygous male Atlantic cod (NEAC_001, Supplementary Notes 1 and 2), applying a whole-genome shotgun approach to 40× coverage (estimated genome size of 830 megabases (Mb), Supplementary Note 4 and Supplementary Fig. 2) using 454 technology (Supplementary Note 3). Two programs (Newbler6 and Celera7, Supplementary Notes 5 and 6) produced assemblies with short contigs, yet with scaffolds of comparable size to those of Sanger-sequenced teleost genomes (Supplementary Note 10 and Supplementary Fig. 8). Although fragmentation due to short tandem repeats is difficult to address (Supplementary Note 7), we resolved numerous gaps attributable to heterozygosity (Supplementary Note 8). The assemblies differ in scaffold and contig length (Table 1), although their scaffolds align to a large extent (Supplementary Note 9 and Supplementary Fig. 7). We obtained about one million single nucleotide polymorphisms (SNPs) by mapping 454 and Illumina reads from the sequenced individual to the Newbler assembly (Supplementary Note 11). Both assemblies cover more than 98% of the reads from an extensive transcriptome data set, indicating that the proteome is well represented (Supplementary Note 13). The assemblies are consistent with four independently assembled bacterial artificial chromosome (BAC) insert clones (Supplementary Note 14 and Supplementary Fig. 9), and with the expected insert size of paired BAC-end reads (Supplementary Note 15 and Supplementary Fig. 10).
A standard annotation approach based on protein evidence was complemented by a whole-genome alignment of the Atlantic cod with the stickleback (Gasterosteus aculeatus), after repeat-masking 25.4% of the Newbler assembly (Supplementary Note 16 and Supplementary Table 6). In this way, 17,920 out of 20,787 protein-coding stickleback genes were mapped onto reorganized scaffolds (Supplementary Note 17). Additional protein-coding genes, pseudogenes and non-coding RNAs were annotated using the standard Ensembl pipeline. These approaches resulted in a final gene set of 22,154 genes (Supplementary Table 7). Comparative analysis of gene ontology classes indicates that the major functional pathways are represented in the annotated gene set (Supplementary Note 18 and Supplementary Fig. 11). We anchored 332 Mb of the Newbler assembly to 23 linkage groups of an existing Atlantic cod linkage map using 924 SNPs8 (Supplementary Note 19 and Supplementary Table 8). These linkage groups have distinct orthology to chromosomes of other teleosts, on the basis of the number of co-occurring genes, showing that the whole-genome shotgun assembly reflects the expected chromosomal ancestry (Fig. 1, Supplementary Note 20 and Supplementary Table 9).
The co-occurrence of orthologous genes (with a minimum of 50% sequence identity over 50% of the alignment, sphere size indicates the numbers of syntenic genes) in 23 Atlantic cod linkage groups8 (x-axis) reveals synteny with the chromosomes of four teleosts (y-axis). Several genes located on the stickleback chromosome XIV, tetraodon chromosome 4 and medaka chromosome 12 indicate a lineage-specific chromosomal rearrangement in Atlantic cod.
Well-studied haemoglobin polymorphisms in Atlantic cod are indicative of functional molecular adaptation to thermal variation9,10,11,12. The genome contains nine α- and β-globin genes that are organized in two unlinked clusters, β5–α1–β1–α4 and β3–β4–α2–α3–β2 (refs 13, 14). We discovered an indel polymorphism of 73 base pairs (bp) in the intergenic promoter region of the α1–β1 globin pair (Fig. 2a and Supplementary Note 21). This promoter polymorphism occurs in highly significant linkage disequilibrium with two known polymorphic sites in the β1 gene, the Val55Met and Ala62Lys substitutions1, in eight Atlantic cod populations (Supplementary Note 22 and Supplementary Fig. 12). In fact, in the three most northern Atlantic populations and in both Baltic populations, the cod β1-globin gene predominantly occurs as a single homozygous genotype consisting of the long promoter and the Val 55–Ala 62 allele (Supplementary Table 10). By placing the two promoter variants in front of a luciferase reporter gene and transfecting the constructs into salmon kidney cells (Supplementary Note 23), we found that temperature and promoter type have a significant interaction effect (generalized linear model, F2, 36 = 7.85, P = 0.007, Fig. 2b) and that the long promoter has twofold higher transcriptional activity compared to the short promoter at 15 °C and 20 °C. Increased globin synthesis of the Val 55–Ala 62 allele would compensate for its lower oxygen affinity10,11 at high temperatures. Thus, the promoter polymorphism provides a molecular compensatory mechanism that helps to maintain the total oxygen-carrying capacity15. The tight linkage between the two types of polymorphism provides a compelling example of the coevolution of structural and regulatory adaptation, and highlights the relationship between temperature and functional molecular variation in the haemoglobin system16.
a, Schematic of the head-to-head organized α1 and β1 globin genes, the intergenic promoter region and transcription start sites (red arrows). A promoter polymorphism consisting of a 73-bp indel (red box) segregates in linkage disequilibrium with two amino-acid-substitution polymorphisms (vertical lines) at positions 55 and 62 in β1 globin that affect its oxygen-binding affinity. This linkage disequilibrium results in two predominant haplotypes, long–Val–Ala and short–Met–Lys. b, Normalized luciferase luminescence ratios in salmon kidney cells. Cells were transfected using the long promoter (black circles) or the short promoter (white circles) and incubated at 4 °C, 15 °C or 20 °C (n = 3 for each treatment level). Error bars show 95% confidence intervals.
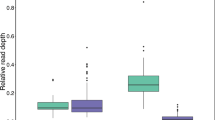
The Atlantic cod immune system has unusual properties that set it apart from that of other teleosts: high levels of IgM17, a minimal antibody response after pathogen exposure5,17,18 and abundant phagocytic neutrophils in the peripheral blood19,20. Despite speculation, the exact causes for these differences remain unknown5. We found that most genes involved in the vertebrate immune response are present in Atlantic cod (Supplementary Note 24, Supplementary Fig. 13 and Supplementary Table 11). Nevertheless, we did not find genes for the MHC II isoforms, their assembly and trafficking chaperone Ii21 and the MHC II-interacting protein CD4, which is essential for helper T-cell activation. By comparing a comprehensive set of vertebrate MHC II, CD4 and Ii sequences to the genome assemblies and all unassembled 454 and Illumina sequencing reads (a data set of about 49.5 gigabases), we detected a truncated pseudogene for CD4 (Supplementary Note 25), which is located in a region of conserved synteny (Supplementary Note 27 and Supplementary Fig. 18). No traces of MHC II and Ii were found in syntenous regions (Supplementary Note 27 and Supplementary Figs 16, 17, 19 and 20) and quantitative PCR (qPCR) targeting a conserved domain in MHC II did not amplify the target sequence (Supplementary Note 26 and Supplementary Fig. 15). The absence of MHC II and Ii, and the pseudogenic nature of CD4, show that Atlantic cod has lost the function of the classical pathway for adaptive immunity against bacterial and parasitic infections. Nevertheless, Atlantic cod deals adequately with its prevailing pathogen load in its natural ecological settings5. Previous transcriptional (complementary DNA) studies in Atlantic cod have indicated an expansion of the number of MHC I loci22,23. By targeting the conserved MHC I α3 domain in genomic DNA using qPCR, we quantified more accurately the number of loci belonging to the teleost U-lineage24 (Supplementary Note 28). Notably, Atlantic cod has about 100 classical MHC I loci, which is a highly expanded number compared to other teleosts (Fig. 3a). A phylogenetic analysis of teleost MHC I sequences supports the existence of two clades in cod (Fig. 3b and Supplementary Note 29). Within each clade, the mutation patterns show statistically significant signs of positive selection that are indicative of subfunctionalization. These findings indicate that loss of MHC II functionality has coincided with a more versatile usage of the cytosolic pathway of MHC I. Two different MHC I antigen-presentation pathways—the classical pathway and the alternative cross-presentation pathway—can initiate immune responses in mammals25. The cross-presentation pathway represents a structural and cellular modification of the MHC I machinery that allows activation of CD8+ T cells upon bacterial infection. The cytokine gene profile of Atlantic cod (Supplementary Table 11) supports the possibility of generating different subsets of CD8+ T cells that either provide direct protection or regulate other immune cells, and thus compensate for the loss of CD4+ T cells.
a, Copy-number estimates of the MHC I α3 domain. Estimates are based on qPCR ratios (see Supplementary Note 28) of the MHC I α3 domain and a single-copy reference gene. For Atlantic cod, β2-microglobulin and topoisomerase III-α (*) were used as reference genes; for human and stickleback, β2-microglobulin was used. The estimates for human and stickleback agree with the expected number of α3 domains found in both reference genomes (Supplementary Table 15). Black dots indicate 95% confidence intervals calculated by bootstrapping (n = 50,000). b, Phylogeny of amino-acid sequences of MHC I α1–α3 domains in teleosts. The Atlantic cod sequences are derived from cDNA and comprise classical U-lineage MHC I only. The other teleost sequences were obtained from Ensembl and NCBI, and contain classical and non-classical U-lineage MHC I. Alignments were visually inspected and corrected where necessary. Maximum likelihood (ML) values and Bayesian posterior probabilities (dots) support the main branches on the ML topology. Distance represents the number of substitutions per site (scale bar). The ratio of non-synonymous to synonymous variable sites (Ka/Ks), the average nucleotide diversity per site (π) and Tajima’s D (D) were calculated for the two main clades in Atlantic cod.
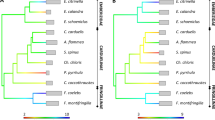
In addition to the MHC I expansion, we found an unusual composition of the highly conserved TLR families that have a fundamental role in the innate immune response and the initial detection of pathogens. Teleost TLR-encoding genes occur in well-supported phylogenetic clusters, most of which share functional properties with mammalian orthologues, although some are fish-specific26. The Atlantic cod TLR genes form monophyletic groups within the known teleost functional groups (Fig. 4, Supplementary Note 30 and Supplementary Fig. 22). Genes for several TLRs that recognize bacterial surface antigens (TLR1, TLR2 and TLR5) are, however, absent, leaving only the teleost-specific TLR14 and TLR18 as members of the TLR1 family in Atlantic cod. Moreover, several families of TLRs that recognize nucleic acids (TLR7, TLR8, TLR9 and TLR22) have markedly expanded, resulting in the highest number of TLRs found in a teleost so far. This TLR repertoire indicates that the Atlantic cod immune system relies relatively heavily on nucleic-acid-detecting TLRs to recognize bacterial pathogens. Notably, the gene expansion of TLR9 coincides with an expansion of interleukin-8 genes (IL-8, Supplementary Table 11). IL-8 is an important chemokine in the innate immune response and is directly induced by TLR9 in human neutrophils27. The corresponding expansions of IL-8 and TLR9 indicate that this signalling cascade is particularly important in Atlantic cod.
TLR protein sequences were selected on the basis of the conserved Toll-IL-1 receptor (TIR) domain for Atlantic cod, including known sequences from stickleback, zebrafish, tetraodon, fugu, medaka and human as references. TLR clades with (*) or without (−) Atlantic cod sequences are denoted according to human or teleost orthologues (summary tree topology, top left panel). Distance represents the average number of substitutions per site (scale bar). ML values and Bayesian posterior probabilities greater than 75/0.75 support the ML topology. Detailed topologies of TLR7 (blue), TLR8 (purple), TLR9 (green) and TLR22 (grey) show gene expansions for Atlantic cod (red). Multiple TLR copies within species are subdivided by letters, and follow Ensembl nomenclature for D. rerio.
The loss of MHC II function and lack of a CD4+ T-cell response represent a fundamental change in how the adaptive immune system is initiated and regulated in Atlantic cod. The marked expansion of MHC I genes and unusual TLR composition signify a shift of its immune system in handling microbial pathogens. An expanded MHC I repertoire in the presence of a non-polymorphic MHC II is found in an evolutionarily-distant vertebrate, the axolotl (Ambystoma mexicanum)28,29. These observations indicate that anomalous immune systems (possibly analogous to that of Atlantic cod) have evolved independently. Additionally, we did not recover evidence for expressed MHC II, CD4 and Ii in the transcriptomes of three other gadoids, indicating that the unusual immune system is a derived characteristic of the gadoid lineage (Supplementary Tables 18 and 19).
We have provided the first annotated genome of a species that supports extensive fisheries and is on the verge of becoming an important aquaculture species. This work provides a major foundation for addressing key issues related to the management of natural Atlantic cod populations, such as the concept of fisheries-induced evolution, which dictates that selective harvesting can change the evolutionary trajectory of major life-history traits of natural populations30. Moreover, our novel findings regarding the immune system will allow for more targeted vaccine development, aiding disease management and the process of domestication of Atlantic cod. These findings change fundamental assumptions regarding the evolution of the vertebrate immune system.
Methods Summary
Detailed methods on the sequencing and assembly of data from genomic and transcriptomic origins; annotation, synteny analyses, transfection experiments, bioinformatic analyses and phylogenetic analyses presented in this manuscript are described in the Supplementary Information.
Accession codes
Primary accessions
EMBL/GenBank/DDBJ
Data deposits
The unassembled sequencing reads and Newbler assembly have been deposited at ENA-EMBL under the accession numbers CAEA01000001–CAEA01554869. The annotation is available through Ensembl at http://www.ensembl.org/index.html. These and more resources are also available through http://codgenome.no.
References
Kurlansky, M. Cod: A Biography of the Fish that Changed the World. (Penguin, 1998)
Johansen, S. D. et al. Large-scale sequence analyses of Atlantic cod. New Biotechnol. 25, 263–271 (2009)
Flajnik, M. F. & Kasahara, M. Origin and evolution of the adaptive immune system: genetic events and selective pressures. Nature Rev. Genet. 11, 47–59 (2010)
Litman, G. W., Rast, J. P. & Fugmann, S. D. The origins of vertebrate adaptive immunity. Nature Rev. Immunol. 10, 543–553 (2010)
Pilstrom, L., Warr, G. W. & Stromberg, S. Why is the antibody response of Atlantic cod so poor? The search for a genetic explanation. Fish. Sci. 71, 961–971 (2005)
Miller, J. R., Koren, S. & Sutton, G. Assembly algorithms for next-generation sequencing data. Genomics 95, 315–327 (2010)
Miller, J. R. et al. Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 24, 2818–2824 (2008)
Hubert, S., Higgins, B., Borza, T. & Bowman, S. Development of a SNP resource and a genetic linkage map for Atlantic cod (Gadus morhua). BMC Genomics 11, 191 (2010)
Sick, K. Haemoglobin polymorphism in fishes. Nature 192, 894–896 (1961)
Brix, O., Thorkildsen, S. & Colosimo, A. Temperature acclimation modulates the oxygen binding properties of the Atlantic cod (Gadus morhua L.) genotypes HbI*1/1, HbI*1/2, and HbI*2/2 by changing the concentrations of their major hemoglobin components (results from growth studies at different temperatures). Comp. Biochem. Physiol. A 138, 241–251 (2004)
Andersen, O. et al. Haemoglobin polymorphisms affect the oxygen-binding properties in Atlantic cod populations. Proc. R. Soc. B 276, 833–841 (2009)
Petersen, M. F. & Steffensen, J. F. Preferred temperature of juvenile Atlantic cod Gadus morhua with different haemoglobin genotypes at normoxia and moderate hypoxia. J. Exp. Biol. 206, 359–364 (2003)
Borza, T., Stone, C., Gamperl, A. K. & Bowman, S. Atlantic cod (Gadus morhua) hemoglobin genes: multiplicity and polymorphism. BMC Genet. 10, 51 (2009)
Wetten, O. et al. Genomic organization and gene expression of the multiple globins in Atlantic cod: conservation of globin-flanking genes in chordates infers the origin of the vertebrate globin clusters. BMC Evol. Biol. 10, 315 (2010)
Gamperl, A. K., Busby, C. D., Hori, T. S. F., Afonso, L. O. B. & Hall, J. R. Hemoglobin genotype has minimal influence on the physiological response of juvenile Atlantic cod (Gadus morhua) to environmental challenges. Physiol. Biochem. Zool. 82, 483–494 (2009)
Weber, R. E. & Fago, A. Functional adaptation and its molecular basis in vertebrate hemoglobins, neuroglobins and cytoglobins. Respir. Physiol. Neurobiol. 144, 141–159 (2004)
Magnadottir, B., Gudmundsdottir, S., Gudmundsdottir, B. K. & Helgason, S. Natural antibodies of cod (Gadus morhua L.): specificity, activity and affinity. Comp. Biochem. Physiol. B 154, 309–316 (2009)
Solem, S. T. & Stenvik, J. Antibody repertoire development in teleosts—a review with emphasis on salmonids and Gadus morhua L. Dev. Comp. Immunol. 30, 57–76 (2006)
Rønneseth, A., Wergeland, H. I. & Pettersen, E. F. Neutrophils and B-cells in Atlantic cod (Gadus morhua L.). Fish Shellfish Immunol. 23, 493–503 (2007)
Øverland, H. S., Pettersen, E. F., Rønneseth, A. & Wergeland, H. I. Phagocytosis by B-cells and neutrophils in Atlantic salmon (Salmo salar L.) and Atlantic cod (Gadus morhua L.). Fish Shellfish Immunol. 28, 193–204 (2010)
Landsverk, O. J., Bakke, O. & Gregers, T. F. MHC II and the endocytic pathway: regulation by invariant chain. Scand. J. Immunol. 70, 184–193 (2009)
Persson, A. C., Stet, R. J. M. & Pilstrom, L. Characterization of MHC class I and β2-microglobulin sequences in Atlantic cod reveals an unusually high number of expressed class I genes. Immunogenetics 50, 49–59 (1999)
Miller, K. M., Kaukinen, K. H. & Schulze, A. D. Expansion and contraction of major histocompatibility complex genes: a teleostean example. Immunogenetics 53, 941–963 (2002)
Dijkstra, J. M. et al. A third broad lineage of major histocompatibility complex (MHC) class I in teleost fish; MHC class II linkage and processed genes. Immunogenetics 59, 305–321 (2007)
Amigorena, S. & Savina, A. Intracellular mechanisms of antigen cross presentation in dendritic cells. Curr. Opin. Immunol. 22, 109–117 (2010)
Rebl, A., Goldammer, T. & Seyfert, H. M. Toll-like receptor signaling in bony fish. Vet. Immunol. Immunopathol. 134, 139–150 (2010)
József, L., Khreiss, T., El Kebir, D. & Filep, J. G. Activation of TLR-9 induces IL-8 secretion through peroxynitrite signaling in human neutrophils. J. Immunol. 176, 1195–1202 (2006)
Tournefier, A. et al. Structure of MHC class I and class II cDNAs and possible immunodeficiency linked to class II expression in the Mexican axolotl. Immunol. Rev. 166, 259–277 (1998)
Kaufman, J., Volk, H. & Wallny, H. J. A. “Minimal essential Mhc” and an “unrecognized Mhc”: two extremes in selection for polymorphism. Immunol. Rev. 143, 63–88 (1995)
Stenseth, N. C. & Dunlop, E. S. Evolution: unnatural selection. Nature 457, 803–804 (2009)
Acknowledgements
This work was supported by a grant from the Research Council of Norway (FUGE program) to K.S.J. The authors wish to thank the following people and organizations: the 454 Life Science Sequencing Center (Branford, USA); the 454 and Illumina nodes of the Norwegian Sequencing Centre (University of Oslo); M. Egholm (formerly 454 Life Science); the Norwegian Metacenter for Computational Science (Notur) and the Norwegian Storage Infrastructure (Norstore); the Research Computing Services group, especially B. -H. Mevik, at the Center for Information Technology (University of Oslo); B. Walenz (Celera); the Canadian Cod Genomics and Broodstock Development Consortium; P. Olsvik, K. Lie and E. Holen at the Norwegian National Institute of Nutrition and Seafood Research (NIFES); J. Gaup and H. Bakke (CEES, University of Oslo); M. Kent (CIGENE, Norwegian University of Life Sciences); S. Bowman (Genome Atlantic); the FUGE bioinformatics platforms group, especially S. Grindhaug; I. Sandlie and O. B. Landsverk (Centre for Immune Regulation, University of Oslo); and Roche Norway.
Author information
Authors and Affiliations
Contributions
DNA and RNA isolation, library construction and sequencing: A.T.-K., M.S., M.H.S., T.B.R., M.M., M.E., B.S., A.J.N. and J.T. Sanger BAC (end-) sequencing: H.K. and R.R. Assembly: A.J.N., B.S., A.S. and A.L. Linkage map analyses: K.G.T. and B.S. SNP analyses: K.G.T., P.R.B., S.L. and A.J.N. Annotation: J.-H.V., B.A. and S.S. Repeat analyses: B.S. Synteny analyses: J.P. and B.S. Haemoglobin analyses: Ø.A., O.F.W., B.S. and T.G. Bioinformatics: A.J.N., B.S., A.S., T.B.R., J.P., C.P., C.N., R.B.E., R.W., J.K., K.L., A.L., I.J., M.M., K.M., P.R.B., K.G.T. and M.H.S. Immune analyses: U.G., M.M., M.H.S., M.E., B.S., B.O.K., T.M., K.L., S.D.J. and T.B.R. Interpretation of immune results: U.G., T.F.G., S.J., B.S. and K.S.J. 454 contributions: L.D. Revisions: Ø.A., T.M., S.D.J., F.N., I.J., S.J., N.C.S. and S.W.O. Project initiation: S.W.O., I.J., F.N., S.L., N.C.S. and K.S.J. Project coordination: S.J. Consortium leader: K.S.J. This manuscript is dedicated to the memory of L. Pilström and R. J. M. Stet. Their research inspired our work to understand further the Atlantic cod immune system.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, Supplementary Figures 1-22 with legends, Supplementary Tables 1-19 and additional references. (PDF 8386 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Star, B., Nederbragt, A., Jentoft, S. et al. The genome sequence of Atlantic cod reveals a unique immune system. Nature 477, 207–210 (2011). https://doi.org/10.1038/nature10342
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10342
This article is cited by
-
Copy number variation and elevated genetic diversity at immune trait loci in Atlantic and Pacific herring
BMC Genomics (2024)
-
Evolutionary trade-off between innate and acquired immune defences in birds
Frontiers in Zoology (2023)
-
A survey of the adaptive immune genes of the polka-dot batfish Ogcocephalus cubifrons
BMC Immunology (2023)
-
Comparative genomics of the T cell receptor μ locus in marsupials and monotremes
Immunogenetics (2023)
-
Evolutionary traits and functional roles of chemokines and their receptors in the male pregnancy of the Syngnathidae
Marine Life Science & Technology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.