Abstract
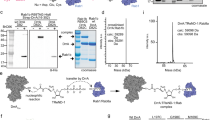
Legionella pneumophila actively modulates host vesicle trafficking pathways to facilitate its intracellular replication with effectors translocated by the Dot/Icm type IV secretion system (T4SS)1. The SidM/DrrA protein functions by locking the small GTPase Rab1 into an active form by its guanine nucleotide exchange factor (GEF) and AMPylation activity2,3,4. Here we demonstrate that the L. pneumophila protein SidD preferably deAMPylates Rab1. We found that the deAMPylation activity of SidD could suppress the toxicity of SidM to yeast and is required to release Rab1 from bacterial phagosomes efficiently. A molecular mechanism for the temporal control of Rab1 activity in different phases of L. pneumophila infection is thus established. These observations indicate that AMPylation-mediated signal transduction is a reversible process regulated by specific enzymes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ensminger, A. W. & Isberg, R. R. Legionella pneumophila Dot/Icm translocated substrates: a sum of parts. Curr. Opin. Microbiol. 12, 67–73 (2009)
Machner, M. P. & Isberg, R. R. Targeting of host Rab GTPase function by the intravacuolar pathogen Legionella pneumophila . Dev. Cell 11, 47–56 (2006)
Murata, T. et al. The Legionella pneumophila effector protein DrrA is a Rab1 guanine nucleotide-exchange factor. Nature Cell Biol. 8, 971–977 (2006)
Muller, M. P. et al. The Legionella effector protein DrrA AMPylates the membrane traffic regulator Rab1b. Science 329, 946–949 (2010)
Isberg, R. R., O'Connor, T. J. & Heidtman, M. The Legionella pneumophila replication vacuole: making a cosy niche inside host cells. Nature Rev. Microbiol. 7, 13–24 (2009)
Kubori, T., Shinzawa, N., Kanuka, H. & Nagai, H. Legionella metaeffector exploits host proteasome to temporally regulate cognate effector. PLoS Pathog. 6, e1001216 (2010)
Ingmundson, A., Delprato, A., Lambright, D. G. & Roy, C. R. Legionella pneumophila proteins that regulate Rab1 membrane cycling. Nature 450, 365–369 (2007)
Worby, C. A. et al. The fic domain: regulation of cell signaling by adenylylation. Mol. Cell 34, 93–103 (2009)
Kinch, L. N., Yarbrough, M. L., Orth, K. & Grishin, N. V. Fido, a novel AMPylation domain common to Fic, Doc, and AvrB. PLoS ONE 4, e5818 (2009)
Pan, X., Luhrmann, A., Satoh, A., Laskowski-Arce, M. A. & Roy, C. R. Ankyrin repeat proteins comprise a diverse family of bacterial type IV effectors. Science 320, 1651–1654 (2008)
Belyi, Y., Tabakova, I., Stahl, M. & Aktories, K. Lgt: a family of cytotoxic glucosyltransferases produced by Legionella pneumophila . J. Bacteriol. 190, 3026–3035 (2008)
Shen, X. et al. Targeting eEF1A by a Legionella pneumophila effector leads to inhibition of protein synthesis and induction of host stress response. Cell. Microbiol. 11, 911–926 (2009)
Luo, Z. Q. & Isberg, R. R. Multiple substrates of the Legionella pneumophila Dot/Icm system identified by interbacterial protein transfer. Proc. Natl Acad. Sci. USA 101, 841–846 (2004)
Chien, M. et al. The genomic sequence of the accidental pathogen Legionella pneumophila . Science 305, 1966–1968 (2004)
Roy, C. R. & Mukherjee, S. Bacterial FIC proteins AMP up infection. Sci. Signal. 2, pe14 (2009)
Brombacher, E. et al. Rab1 guanine nucleotide exchange factor SidM is a major phosphatidylinositol 4-phosphate-binding effector protein of Legionella pneumophila . J. Biol. Chem. 284, 4846–4856 (2009)
Soding, J. Protein homology detection by HMM–HMM comparison. Bioinformatics 21, 951–960 (2005)
Rantanen, M. K., Lehtio, L., Rajagopal, L., Rubens, C. E. & Goldman, A. Structure of Streptococcus agalactiae serine/threonine phosphatase. The subdomain conformation is coupled to the binding of a third metal ion. FEBS J. 274, 3128–3137 (2007)
Schlicker, C. et al. Structural analysis of the PP2C phosphatase tPphA from Thermosynechococcus elongatus: a flexible flap subdomain controls access to the catalytic site. J. Mol. Biol. 376, 570–581 (2008)
Seabra, M. C. & Wasmeier, C. Controlling the location and activation of Rab GTPases. Curr. Opin. Cell Biol. 16, 451–457 (2004)
Woolery, A. R., Luong, P., Broberg, C. A. & Orth, K. AMPylation: something old is new again. Front. Microbiol 1, 113 (2010)
Yarbrough, M. L. et al. AMPylation of Rho GTPases by Vibrio VopS disrupts effector binding and downstream signaling. Science 323, 269–272 (2009)
Zhu, W. et al. Comprehensive identification of protein substrates of the Dot/Icm type IV transporter of Legionella pneumophila . PLoS ONE 6, e17638 (2011)
Berger, K. H. & Isberg, R. R. Two distinct defects in intracellular growth complemented by a single genetic locus in Legionella pneumophila . Mol. Microbiol. 7, 7–19 (1993)
Conover, G. M., Derre, I., Vogel, J. P. & Isberg, R. R. The Legionella pneumophila LidA protein: a translocated substrate of the Dot/Icm system associated with maintenance of bacterial integrity. Mol. Microbiol. 48, 305–321 (2003)
Liu, Y., Gao, P., Banga, S. & Luo, Z. Q. An in vivo gene deletion system for determining temporal requirement of bacterial virulence factors. Proc. Natl Acad. Sci. USA 105, 9385–9390 (2008)
Fan, H. Y., Cheng, K. K. & Klein, H. L. Mutations in the RNA polymerase II transcription machinery suppress the hyperrecombination mutant hpr1Δ of Saccharomyces cerevisiae . Genetics 142, 749–759 (1996)
Gietz, R. D., Schiestl, R. H., Willems, A. R. & Woods, R. A. Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11, 355–360 (1995)
Xu, L. et al. Inhibition of host vacuolar H+-ATPase activity by a Legionella pneumophila effector. PLoS Pathog. 6, e1000822 (2010)
Acknowledgements
We thank R. Isberg for the antibody against SidM and A. Aronson and A. Tao for critical reading of the manuscript and for discussions. This work was supported by NIH-NIAID grants R01AI069344, K02AI085403 and R21AI092043 (Z.-Q.L).
Author information
Authors and Affiliations
Contributions
Y.T. and Z.-Q.L. conceived the project. Y.T. performed the experiments. Y.T. and Z.-Q.L analysed the data. Z.-Q.L wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-5 with legends and Supplementary Table 1. (PDF 1986 kb)
Rights and permissions
About this article
Cite this article
Tan, Y., Luo, ZQ. Legionella pneumophila SidD is a deAMPylase that modifies Rab1. Nature 475, 506–509 (2011). https://doi.org/10.1038/nature10307
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10307
This article is cited by
-
Translocated Legionella pneumophila small RNAs mimic eukaryotic microRNAs targeting the host immune response
Nature Communications (2022)
-
Regulation of the small GTPase Rab1 function by a bacterial glucosyltransferase
Cell Discovery (2018)
-
Legionella and Coxiella effectors: strength in diversity and activity
Nature Reviews Microbiology (2017)
-
A unique deubiquitinase that deconjugates phosphoribosyl-linked protein ubiquitination
Cell Research (2017)
-
Genomic analysis of 38 Legionella species identifies large and diverse effector repertoires
Nature Genetics (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.