Abstract
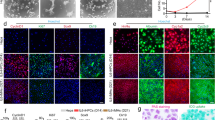
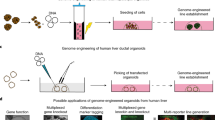
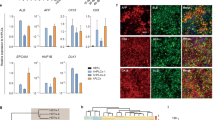
The location and timing of cellular differentiation must be stringently controlled for proper organ formation. Normally, hepatocytes differentiate from hepatic progenitor cells to form the liver during development1,2. However, previous studies have shown that the hepatic program can also be activated in non-hepatic lineage cells after exposure to particular stimuli or fusion with hepatocytes3,4,5,6,7,8,9. These unexpected findings suggest that factors critical to hepatocyte differentiation exist and become activated to induce hepatocyte-specific properties in different cell types. Here, by screening the effects of twelve candidate factors, we identify three specific combinations of two transcription factors, comprising Hnf4α plus Foxa1, Foxa2 or Foxa3, that can convert mouse embryonic and adult fibroblasts into cells that closely resemble hepatocytes in vitro. The induced hepatocyte-like (iHep) cells have multiple hepatocyte-specific features and reconstitute damaged hepatic tissues after transplantation. The generation of iHep cells may provide insights into the molecular nature of hepatocyte differentiation and potential therapies for liver diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zaret, K. S. Regulatory phases of early liver development: paradigms of organogenesis. Nature Rev. Genet. 3, 499–512 (2002)
Zaret, K. S. & Grompe, M. Generation and regeneration of cells of the liver and pancreas. Science 322, 1490–1494 (2008)
Scarpelli, D. G. & Rao, M. S. Differentiation of regenerating pancreatic cells into hepatocyte-like cells. Proc. Natl Acad. Sci. USA 78, 2577–2581 (1981)
Reddy, J. K. et al. Induction and origin of hepatocytes in rat pancreas. J. Cell Biol. 98, 2082–2090 (1984)
Wang, X., Al-Dhalimy, M., Lagasse, E., Finegold, M. & Grompe, M. Liver repopulation and correction of metabolic liver disease by transplanted adult mouse pancreatic cells. Am. J. Pathol. 158, 571–579 (2001)
Lee, K. D. et al. In vitro hepatic differentiation of human mesenchymal stem cells. Hepatology 40, 1275–1284 (2004)
Banas, A. et al. Adipose tissue-derived mesenchymal stem cells as a source of human hepatocytes. Hepatology 46, 219–228 (2007)
Wang, X. et al. Cell fusion is the principal source of bone-marrow-derived hepatocytes. Nature 422, 897–901 (2003)
Vassilopoulos, G., Wang, P. R. & Russell, D. W. Transplanted bone marrow regenerates liver by cell fusion. Nature 422, 901–904 (2003)
Overturf, K. et al. Hepatocytes corrected by gene therapy are selected in vivo in a murine model of hereditary tyrosinaemia type I. Nature Genet. 12, 266–273 (1996)
Suzuki, A. et al. Flow cytometric isolation and clonal identification of self-renewing bipotent hepatic progenitor cells in adult mouse liver. Hepatology 48, 1964–1978 (2008)
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001)
Alvarez-Dolado, M. et al. Fusion of bone-marrow-derived cells with Purkinje neurons, cardiomyocytes and hepatocytes. Nature 425, 968–973 (2003)
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987–1000 (1987)
Xie, H., Ye, M., Feng, R. & Graf, T. Stepwise reprogramming of B cells into macrophages. Cell 117, 663–676 (2004)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Zhou, Q., Brown, J., Kanarek, A., Rajagopal, J. & Melton, D. A. In vivo reprogramming of adult pancreatic exocrine cells to β-cells. Nature 455, 627–632 (2008)
Feng, R. et al. PU.1 and C/EBPα/β convert fibroblasts into macrophage-like cells. Proc. Natl Acad. Sci. USA 105, 6057–6062 (2008)
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010)
Ieda, M. et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142, 375–386 (2010)
Szabo, E. et al. Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 468, 521–526 (2010)
Seglen, P. O. Hepatocyte suspensions and cultures as tools in experimental carcinogenesis. J. Toxicol. Environ. Health 5, 551–560 (1979)
Kaneko, S., Onodera, M., Fujiki, Y., Nagasawa, T. & Nakauchi, H. Simplified retroviral vector GCsap with murine stem cell virus long terminal repeat allows high and continued expression of enhanced green fluorescent protein by human hematopoietic progenitors engrafted in nonobese diabetic/severe combined immunodeficient mice. Hum. Gene Ther. 12, 35–44 (2001)
Suzuki, A., Nakauchi, H. & Taniguchi, H. Glucagon-like peptide 1 (1–37) converts intestinal epithelial cells into insulin-producing cells. Proc. Natl Acad. Sci. USA 100, 5034–5039 (2003)
Suzuki, A., Iwama, A., Miyashita, H., Nakauchi, H. & Taniguchi, H. Role for growth factors and extracellular matrix in controlling differentiation of prospectively isolated hepatic stem cells. Development 130, 2513–2524 (2003)
Suzuki, A. et al. Flow cytometric separation and enrichment of hepatic progenitor cells in the developing mouse liver. Hepatology 32, 1230–1239 (2000)
Suzuki, A. et al. Clonal identification and characterization of self-renewing pluripotent stem cells in the developing liver. J. Cell Biol. 156, 173–184 (2002)
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003)
Quackenbush, J. Microarray data normalization and transformation. Nature Genet. 32, 496–501 (2002)
Acknowledgements
We thank A. Iwama, H. Miyoshi and R. M. Tanguay for sharing reagents, F. Costantini for providing the R26RYFP mice and E. Gunshima, A. Kaneyuki and H. Kuboyama for excellent technical assistance. This work was supported in part by the Program for Improvement of the Research Environment for Young Researchers from the Special Coordination Funds for Promoting Science and Technology commissioned by the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan, a Grant-in-Aid for Scientific Research from the MEXT of Japan and the Precursory Research for Embryonic Science and Technology Program of the Japan Science and Technology Agency.
Author information
Authors and Affiliations
Contributions
A.S. designed the study and wrote the paper. S.S. and A.S. performed experiments and analysed and interpreted the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References, Supplementary Figures 1-23 with legends and Supplementary Table 1. (PDF 30029 kb)
Rights and permissions
About this article
Cite this article
Sekiya, S., Suzuki, A. Direct conversion of mouse fibroblasts to hepatocyte-like cells by defined factors. Nature 475, 390–393 (2011). https://doi.org/10.1038/nature10263
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10263
This article is cited by
-
Suppression of CEBPδ recovers exhaustion in anti-metastatic immune cells
Scientific Reports (2023)
-
Liver cell therapies: cellular sources and grafting strategies
Frontiers of Medicine (2023)
-
Hepatocyte generation in liver homeostasis, repair, and regeneration
Cell Regeneration (2022)
-
Orthogonally induced differentiation of stem cells for the programmatic patterning of vascularized organoids and bioprinted tissues
Nature Biomedical Engineering (2022)
-
IL6 supports long-term expansion of hepatocytes in vitro
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.