Abstract
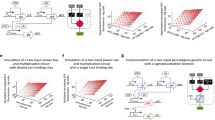
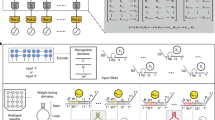
The impressive capabilities of the mammalian brain—ranging from perception, pattern recognition and memory formation to decision making and motor activity control—have inspired their re-creation in a wide range of artificial intelligence systems for applications such as face recognition, anomaly detection, medical diagnosis and robotic vehicle control1. Yet before neuron-based brains evolved, complex biomolecular circuits provided individual cells with the ‘intelligent’ behaviour required for survival2. However, the study of how molecules can ‘think’ has not produced an equal variety of computational models and applications of artificial chemical systems. Although biomolecular systems have been hypothesized to carry out neural-network-like computations in vivo3,2,4 and the synthesis of artificial chemical analogues has been proposed theoretically5,6,7,8,9, experimental work10,11,12,13 has so far fallen short of fully implementing even a single neuron. Here, building on the richness of DNA computing14 and strand displacement circuitry15, we show how molecular systems can exhibit autonomous brain-like behaviours. Using a simple DNA gate architecture16 that allows experimental scale-up of multilayer digital circuits17, we systematically transform arbitrary linear threshold circuits18 (an artificial neural network model) into DNA strand displacement cascades that function as small neural networks. Our approach even allows us to implement a Hopfield associative memory19 with four fully connected artificial neurons that, after training in silico, remembers four single-stranded DNA patterns and recalls the most similar one when presented with an incomplete pattern. Our results suggest that DNA strand displacement cascades could be used to endow autonomous chemical systems with the capability of recognizing patterns of molecular events, making decisions and responding to the environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rojas, R. Neural Networks: A Systematic Introduction (Springer, 1996)
Bray, D. Protein molecules as computational elements in living cells. Nature 376, 307–312 (1995)
Mjolsness, E., Sharp, D. H. & Reinitz, J. A connectionist model of development. J. Theor. Biol. 152, 429–453 (1991)
Buchler, N. E., Gerland, U. & Hwa, T. On schemes of combinatorial transcription logic. Proc. Natl Acad. Sci. USA 100, 5136–5141 (2003)
Hjelmfelt, A., Weinberger, E. D. & Ross, J. Chemical implementation of neural networks and Turing machines. Proc. Natl Acad. Sci. USA 88, 10983–10987 (1991)
Baum, E. B. Building an associative memory vastly larger than the brain. Science 268, 583–585 (1995)
Mills, A. P., Yurke, B. & Platzman, P. M. Article for analog vector algebra computation. Biosystems 52, 175–180 (1999)
Kim, J., Hopfield, J. J. & Winfree, E. in Advances in Neural Information Processing Systems Vol. 17 (eds Saul, L. K., Weiss, Y. & Bottou, L. ) 681–688 (MIT Press, 2004)
Zhang, D. Y. & Seelig, G. in DNA Computing and Molecular Programming (eds Sakakibara, Y. & Mi, Y. ) 176–186 (Lecture Notes in Computer Science, Vol. 6518, Springer, 2011)
Laplante, J. P., Pemberton, M., Hjelmfelt, A. & Ross, J. Experiments on pattern recognition by chemical kinetics. J. Phys. Chem. 99, 10063–10065 (1995)
Mills, A. P., Jr, Turberfield, M., Turberfield, A. J., Yurke, B. & Platzman, P. M. Experimental aspects of DNA neural network computation. Soft Comput. 5, 10–18 (2001)
Lim, H. W. et al. In vitro molecular pattern classification via DNA-based weighted-sum operation. Biosystems 100, 1–7 (2010)
Kim, J. & Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 7, 465 (2011)
Adleman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994)
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nature Chem. 3, 103–113 (2011)
Qian, L. & Winfree, E. A simple DNA gate motif for synthesizing large-scale circuits. J. R. Soc. Interface 10.1098/rsif.2010.0729 (published online 4 February 2011)
Qian, L. & Winfree, E. Scaling up digital circuit computation with DNA strand displacement cascades. Science 332, 1196–1201 (2011)
Muroga, S. Threshold Logic and its Applications Vol. 18 (Wiley-Interscience, 1971).
Hopfield, J. J. Neural networks and physical systems with emergent collective computational abilities. Proc. Natl Acad. Sci. USA 79, 2554–2558 (1982)
McCulloch, W. S. & Pitts, W. A logical calculus of the ideas immanent in nervous activity. Bull. Math. Biol. 5, 115–133 (1943)
Kautz, W. H. The realization of symmetric switching functions with linear-input logical elements. IRE Trans. Electron. Comput. EC-10, 371–378 (1961)
Wegener, I. The complexity of the parity function in unbounded fan-in, unbounded depth circuits. Theor. Comput. Sci. 85, 155–170 (1991)
Hajnal, A., Maass, W., Pudlák, P., Szegedy, M. & Turan, G. Threshold circuits of bounded depth. J. Comput. Syst. Sci. 46, 129–154 (1993)
Müller, D. E. in Symp. on the Application of Switching Theory (eds Aiken, H. & Main, W. F. ) 289–297 (Stanford Univ. Press, 1963)
Lederman, H., Macdonald, J., Stefanovic, D. & Stojanovic, M. N. Deoxyribozyme-based three-input logic gates and construction of a molecular full adder. Biochemistry 45, 1194–1199 (2006)
Amari, S. I. Characteristics of sparsely encoded associative memory. Neural Netw. 2, 451–457 (1989)
Soloveichik, D., Seelig, G. & Winfree, E. DNA as a universal substrate for chemical kinetics. Proc. Natl Acad. Sci. USA 107, 5393–5398 (2010)
Fernando, C. T. et al. Molecular circuits for associative learning in single-celled organisms. J. R. Soc. Interface 6, 463–469 (2009)
Pei, R., Matamoros, E., Liu, M., Stefanovic, D. & Stojanovic, M. N. Training a molecular automaton to play a game. Nature Nanotechnol. 5, 773–777 (2010)
Rosenfeld, N. et al. MicroRNAs accurately identify cancer tissue origin. Nature Biotechnol. 26, 462–469 (2008)
Simmel, F. C. Towards biomedical applications for nucleic acid nanodevices. Nanomedicine 2, 817–830 (2007)
Acknowledgements
We thank P. Rothemund, P. Yin, D. Woods, D. Soloveichik and N. Dabby for comments on the manuscript. We also thank R. Murray for the use of experimental facilities. This work was supported by the NSF (grant nos 0728703 and 0832824 (The Molecular Programming Project)) and by HFSP award no. RGY0074/2006-C.
Author information
Authors and Affiliations
Contributions
L.Q. designed the system, performed the experiments and analysed the data; L.Q. and E.W. performed the in silico training and wrote the manuscript; E.W. guided the project and discussed the design and the data; and J.B. initiated and guided the project, and discussed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1-28 with legends, Supplementary Methods, Supplementary Text and Data comprising: 1 The seesaw DNA gate motif; 2 Four types of seesaw gates; 3 Four transformation rules; 4 Circuit design lessons learned from experiments; 5 In silico training of dual-rail monotone Hopfield associative memories; 6 A four-neuron dual-rail monotone Hopfield associative memory; 7 Modeling and simulations; 8 Sequences, Supplementary Tables 1-7 and additional references. (PDF 19348 kb)
Rights and permissions
About this article
Cite this article
Qian, L., Winfree, E. & Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 475, 368–372 (2011). https://doi.org/10.1038/nature10262
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10262
This article is cited by
-
Synthetic molecular switches driven by DNA-modifying enzymes
Nature Communications (2024)
-
Three-input logic gate based on DNA strand displacement reaction
Scientific Reports (2023)
-
Digital circuits and neural networks based on acid-base chemistry implemented by robotic fluid handling
Nature Communications (2023)
-
Design and Simulation of an Autonomous Molecular Mechanism Using Spatially Localized DNA Computation
Interdisciplinary Sciences: Computational Life Sciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.