Abstract
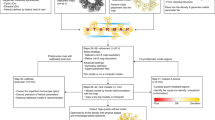
Molecular replacement1,2,3,4 procedures, which search for placements of a starting model within the crystallographic unit cell that best account for the measured diffraction amplitudes, followed by automatic chain tracing methods5,6,7,8, have allowed the rapid solution of large numbers of protein crystal structures. Despite extensive work9,10,11,12,13,14, molecular replacement or the subsequent rebuilding usually fail with more divergent starting models based on remote homologues with less than 30% sequence identity. Here we show that this limitation can be substantially reduced by combining algorithms for protein structure modelling with those developed for crystallographic structure determination. An approach integrating Rosetta structure modelling with Autobuild chain tracing yielded high-resolution structures for 8 of 13 X-ray diffraction data sets that could not be solved in the laboratories of expert crystallographers and that remained unsolved after application of an extensive array of alternative approaches. We estimate that the new method should allow rapid structure determination without experimental phase information for over half the cases where current methods fail, given diffraction data sets of better than 3.2 Å resolution, four or fewer copies in the asymmetric unit, and the availability of structures of homologous proteins with >20% sequence identity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rossmann, M. G. The Molecular Replacement Method (Gordon & Breach, 1972)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Brünger, A. T. et al. Crystallography & NMR system: a new software system for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997)
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protocols 3, 1171–1179 (2008)
Terwilliger, T. C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D 64, 61–69 (2008)
DePristo, M. A., de Bakker, P. I. W., Johnson, R. J. K. & Blundell, T. L. Crystallographic refinement by knowledge-based exploration of complex energy landscapes. Structure 13, 1311–1319 (2005)
Cowtan, K. The Buccaneer software for automated model building. Acta Crystallogr. D 62, 1002–1011 (2006)
Schwarzenbacher, R., Godzik, A., Grzechnik, S. K. & Jaroszewski, L. The importance of alignment accuracy for molecular replacement. Acta Crystallogr. D 60, 1229–1236 (2004)
Rodríguez, D. D. et al. Crystallographic ab initio protein structure solution below atomic resolution. Nature Methods 6, 651–653 (2009)
Suhre, K. & Sanejouand, Y. H. On the potential of normal-mode analysis for solving difficult molecular-replacement problems. Acta Crystallogr. D 60, 796–799 (2004)
Qian, B. et al. High-resolution structure prediction and the crystallographic phase problem. Nature 450, 259–264 (2007)
Schröder, G., Levitt, M. & Brünger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Brünger, A. T., Kuriyan, J. & Karplus, M. Crystallographic R factor refinement by molecular dynamics. Science 235, 458–460 (1987)
Raman, S. et al. NMR structure determination for larger proteins using backbone-only data. Science 327, 1014–1018 (2010)
Das, R. & Baker, D. Macromolecular modeling with Rosetta. Annu. Rev. Biochem. 77, 363–382 (2008)
Brünger, A. T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992)
Söding, J. Protein homology detection by HMM–HMM comparison. Bioinformatics 21, 951–960 (2005)
Schröder, G. F., Brunger, A. T. & Levitt, M. Combining efficient conformational sampling with a deformable elastic network model facilitates structure refinement at low resolution. Structure 15, 1630–1641 (2007)
Brünger, A. T. Extension of molecular replacement: a new search strategy based on Patterson correlation refinement. Acta Crystallogr. A 46, 46–57 (1990)
Brünger, A. T., Karplus, M. & Petsko, G. A. Crystallographic refinement by simulated annealing: application to crambin. Acta Crystallogr. A 45, 50–61 (1989)
Read, R. J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42, 140–149 (1986)
Vitkup, D., Melamud, E., Moult, J. & Sander, C. Completeness in structural genomics. Nature Struct. Biol. 8, 559–566 (2001)
Canutescu, A. & Dunbrack, R. Cyclic coordinate descent: a new algorithm for loop closure in protein modeling. Protein Sci. 12, 963–972 (2003)
DiMaio, F., Tyka, M. D., Baker, M. L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using Rosetta. J. Mol. Biol. 392, 181–190 (2009)
André, I., Bradley, P., Wang, C. & Baker, D. Prediction of the structure of symmetrical protein assemblies. Proc. Natl Acad. Sci. USA 104, 17656–17661 (2007)
Weis, W. I., Brünger, A. T., Skehel, J. J. & Wiley, D. D. Refinement of the influenza virus hemagglutinin by simulated annealing. J. Mol. Biol. 212, 737–761 (1990)
Abe, H., Braun, W., Noguti, T. & Go¯, N. Rapid calculation of first and second derivatives of conformational energy with respect to dihedral angles for proteins general recurrent equations. Comput. Chem. 8, 239–247 (1984)
Eswar, N. et al. Comparative protein structure modeling with MODELLER. Curr. Protoc. Bioinform. (Suppl.) 15, 5.6 10.1002/0471250953.bi0506s15. (2006)
Acknowledgements
R.J.R., T.C.T. and D.B. thank the NIH (5R01GM092802), the Wellcome Trust (R.J.R.), and HHMI (D.B.) for funding this research. F.D. acknowledges the NIH (P41RR002250) and HHMI. D.F. and A.A. acknowledge support from the Israel Science Foundation. G.O. thanks DK Molecular Enzymology (FWF-project W901) and the Austrian Science Fund (FWF-project P19858). The work of A.W. was supported by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. H.I. acknowledges support from the academy of Finland (1131413). S.M.V. was supported by a grant from the Protein Structure Initiative of National Institute of General Medical Sciences (U54 GM074958). The work of P.R.P. at Argonne National Laboratory was supported by the US Department of Energy’s Office of Science, Biological and Environmental Research GTL programme under contract DE-AC02-06CH11357. We thank all members of the JCSG for their general contributions to the protein production and structural work. The JCSG is supported by the NIH, National Institutes of General Medical Sciences, Protein Structure Initiative (U54 GM094586 and GM074898).
Author information
Authors and Affiliations
Contributions
F.D., T.C.T., R.J.R. and D.B. developed the methods described in the manuscript; F.D., T.C.T., R.J.R., A.W. and D.B wrote the paper. A.W., G.O., U.W., E.V., A.A., D.F., H.L.A., D.D., S.M.V., H.I. and P.R.P. provided the data and refined one or more structures to completion.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Text, additional references, Supplementary Tables 1-5 and Supplementary Figures 1-3 with legends. (PDF 1670 kb)
Rights and permissions
About this article
Cite this article
DiMaio, F., Terwilliger, T., Read, R. et al. Improved molecular replacement by density- and energy-guided protein structure optimization. Nature 473, 540–543 (2011). https://doi.org/10.1038/nature09964
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09964
This article is cited by
-
The [4Fe-4S] cluster of sulfurtransferase TtuA desulfurizes TtuB during tRNA modification in Thermus thermophilus
Communications Biology (2020)
-
Structural insights into the mechanism of rhodopsin phosphodiesterase
Nature Communications (2020)
-
Machine learning-based surrogate modeling for data-driven optimization: a comparison of subset selection for regression techniques
Optimization Letters (2020)
-
Crystal structure of heliorhodopsin
Nature (2019)
-
Opportunistic complexes of E. coli L-asparaginases with citrate anions
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.