Abstract
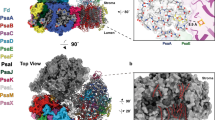
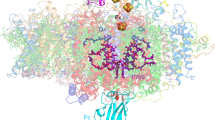
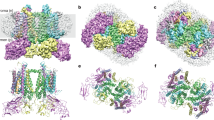
Photosystem II is the site of photosynthetic water oxidation and contains 20 subunits with a total molecular mass of 350 kDa. The structure of photosystem II has been reported at resolutions from 3.8 to 2.9 Å. These resolutions have provided much information on the arrangement of protein subunits and cofactors but are insufficient to reveal the detailed structure of the catalytic centre of water splitting. Here we report the crystal structure of photosystem II at a resolution of 1.9 Å. From our electron density map, we located all of the metal atoms of the Mn4CaO5 cluster, together with all of their ligands. We found that five oxygen atoms served as oxo bridges linking the five metal atoms, and that four water molecules were bound to the Mn4CaO5 cluster; some of them may therefore serve as substrates for dioxygen formation. We identified more than 1,300 water molecules in each photosystem II monomer. Some of them formed extensive hydrogen-bonding networks that may serve as channels for protons, water or oxygen molecules. The determination of the high-resolution structure of photosystem II will allow us to analyse and understand its functions in great detail.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kok, B., Forbush, B. & McGloin, M. Cooperation of charges in photosynthetic oxygen evolution. I. A linear four step mechanism. Photochem. Photobiol. 11, 457–475 (1970)
Joliot, P. Period-four oscillations of the flash-induced oxygen formation in photosynthesis. Photosynth. Res. 76, 65–72 (2003)
Zouni, A. et al. Crystal structure of photosystem II from Synechococcus elongatus at 3.8 Å resolution. Nature 409, 739–743 (2001)
Ferreira, K. N., Iverson, T. M., Maghlaoui, K., Barber, J. & Iwata, S. Architecture of the photosynthetic oxygen-evolving center. Science 303, 1831–1838 (2004)
Guskov, A. et al. Cyanobacterial photosystem II at 2.9 Å resolution and role of quinones, lipids, channels and chloride. Nature Struct. Mol. Biol. 16, 334–342 (2009)
Kamiya, N. & Shen, J.-R. Crystal structure of oxygen-evolving photosystem II from Thermosynechococcus vulcanus at 3.7-Å resolution. Proc. Natl Acad. Sci. USA 100, 98–103 (2003)
Kawakami, K., Iwai, M., Ikeuchi, M., Kamiya, N. & Shen, J.-R. Location of PsbY in oxygen-evolving photosystem II revealed by mutagenesis and X-ray crystallography. FEBS Lett. 581, 4983–4987 (2007)
Broser, M. et al. Crystal structure of monomeric photosystem II from Thermosynechococcus elongatus at 3.6 Å resolution. J. Biol. Chem. 285, 26255–26262 (2010)
De Paula, J. C., Beck, W. F. & Brudvig, G. W. Magnetic properties of manganese in the photosynthetic O2-evolving complex. 2. Evidence for a manganese tetramer. J. Am. Chem. Soc. 108, 4002–4009 (1986)
Carrell, G. & Tyryshkin, A. M. &. Dismukes, G. C. An evaluation of structural models for the photosynthetic water-oxidizing complex derived from spectroscopic and X-ray diffraction signatures. J. Biol. Inorg. Chem. 7, 2–22 (2002)
Vincent, J. B. & Christou, G. A molecular ‘double-pivot’ mechanism for water oxidation. Inorg. Chim. Acta 136, L41–L43 (1987)
Peloquin, J. M. & Britt, R. D. EPR/ENDOR characterization of the physical and electronic structure of the OEC Mn cluster. Biochim. Biophys. Acta 1503, 96–111 (2001)
Robblee, J. H., Cince, R. M. & Yachandra, V. K. X-ray spectroscopy-based structure of the Mn cluster and mechanism of photosynthetic oxygen evolution. Biochim. Biophys. Acta 1503, 7–23 (2001)
Zein, S. et al. Focusing the view on nature’s water-splitting catalyst. Phil. Trans. R. Soc. B 363, 1167–1177 (2008)
Nixon, P. J. & Diner, B. Analysis of water-oxidation mutants constructed in the cyanobacterium Synechocystis sp. PCC 6803. Biochem. Soc. Trans. 22, 338–343 (1994)
Chu, H.-A., Nguyne, A. P. & Debus, R. J. Amino acid residues that influence the binding of manganese or calcium to Photosystem II. 1. The lumenal inter-helical domains of the D1 polypeptide. Biochemistry 34, 5839–5858 (1995)
Hwang, H. J., Dilbeck, P., Debus, R. J. & Burnap, R. L. Mutation of arginine 357 of the CP43 protein of photosystem II severely impairs the catalytic S-state cycle of the H2O oxidation complex. Biochemistry 46, 11987–11997 (2007)
Debus, R. J. Protein ligation of the photosynthetic oxygen-evolving center. Coord. Chem. Rev. 252, 244–258 (2008)
Service, R. J., Hillier, W. & Debus, R. J. Evidence from FTIR difference spectroscopy of an extensive network of hydrogen bonds near the oxygen-evolving Mn4Ca cluster of photosystem II involving D1-Glu65, D2-Glu312, and D1-Glu329. Biochemistry 49, 6655–6669 (2010)
Murray, J. W. & Barber, J. Structural characteristics of channels and pathways in photosystem II including the identification of an oxygen channel. J. Struct. Biol. 159, 228–237 (2007)
Ho, F. M. & Styring, S. Access channels and methanol binding site to the CaMn4 cluster in Photosystem II based on solvent accessibility simulation, with implications for substrate water access. Biochim. Biophys. Acta 1777, 140–153 (2008)
Zhang, C. Low-barrier hydrogen bond plays key role in active photosystem II—A new model for photosynthetic water oxidation. Biochim. Biophys. Acta 1767, 493–499 (2007)
Hoganson, C. W. & Babcock, G. T. A metalloradical mechanism for the generation of oxygen from water in photosynthesis. Science 277, 1953–1956 (1997)
Tommos, C. & Babcock, G. T. Proton and hydrogen currents in photosynthetic water oxidation. Biochim. Biophys. Acta 1458, 199–219 (2000)
Hays, A.-M. A., Vassiliev, I. R., Golbeck, J. H. & Debus, R. J. Role of D1-His190 in the proton-coupled oxidation of tyrosine YZ in manganese-depleted photosystem II. Biochemistry 38, 11851–11865 (1999)
Murray, J. W. et al. X-ray crystallography identifies two chloride binding sites in the oxygen evolving centre of photosystem II. Energy Environ. Sci. 1, 161–166 (2008)
Kawakami, K., Umena, Y., Kamiya, N. & Shen, J.-R. Location of chloride and its possible functions in oxygen-evolving photosystem II revealed by X-ray crystallography. Proc. Natl Acad. Sci. USA 106, 8567–8572 (2009)
Burrel, J. W. K., Jackman, L. M. & Weedon, B. L. C. Stereochemistry and synthesis of phytol, geraniol, and nerol. Proc. Chem. Soc. 1959, 263–264 (1959)
Crabbe, P., Djerassi, C., Eisenbraun, E. J. & Liu, S. Optical rotatory dispersion studies. XXIX. Absolute configuration of phytol. Proc. Chem. Soc. 1959, 264–265 (1959)
Vasil’ev, S., Orth, P., Zouni, A., Owens, T. G. & Bruce, D. Excited-state dynamics in photosystem II: Insights from the x-ray crystal structure. Proc. Natl Acad. Sci. USA 98, 8602–8607 (2001)
Shen, J.-R. & Inoue, Y. Binding and functional properties of two new extrinsic components, cytochrome c-550 and a 12 kDa protein, in cyanobacterial photosystem II. Biochemistry 32, 1825–1832 (1993)
Shen, J.-R. & Kamiya, N. Crystallization and the crystal properties of the oxygen-evolving photosystem II from Synechococcus vulcanus . Biochemistry 39, 14739–14744 (2000)
Yano, J. et al. X-ray damage to the Mn4Ca complex in single crystals of photosystem II: a case study for metalloprotein crystallography. Proc. Natl Acad. Sci. USA 102, 12047–12052 (2005)
Cruickshank, D. W. J. Remarks about protein structure precision. Acta Crystallogr. D 55, 583–601 (1999)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
Otwinowski, Z. & Minor, M. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Collaborative Computational Project, Number 4 . The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Daopin, S., Davies, D. R., Schlunegger, M. P. & Grütter, M. G. Comparison of two crystal structures of TGF-β2: the accuracy of refined protein structures. Acta Crystallogr. D 50, 85–92 (1994)
Acknowledgements
The X-ray diffraction data was taken at beamlines BL44XU, BL41XU and BL38B1 at SPring-8. We thank E. Yamashita, N. Shimizu, S. Baba and N. Mizuno for their help in using the beamlines. J.-R.S. thanks Y. Inoue for his support in the initiation of this work. This work was supported by a Grant-in-Aid for Scientific Research on Priority Areas (Structures of Biological Macromolecular Assemblies), a Grant-in-Aid for Creative Scientific Research, a GCOE programme on Pico-biology at the University of Hyogo, a Grant-in-Aid for Scientific Research (C), from the Ministry of Education, Culture, Sports, Science and Technology of Japan, and a research grant from the Yamada Science foundation.
Author information
Authors and Affiliations
Contributions
K.K. performed, and J.-R.S. supervised, the purification and crystallization of PSII. K.K., Y.U. and J.-R.S. performed X-ray diffraction experiments. Y.U. analysed the structure, and N.K. supervised the structure analysis and refinement process. J.-R.S. and N.K. jointly wrote the paper, and all of the authors joined the discussion of the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5, Supplementary Figures 1-6 with legends, Supplementary Data and Results, and additional references. (PDF 1710 kb)
Rights and permissions
About this article
Cite this article
Umena, Y., Kawakami, K., Shen, JR. et al. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011). https://doi.org/10.1038/nature09913
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09913
This article is cited by
-
Electric-field-assisted proton coupling enhanced oxygen evolution reaction
Nature Communications (2024)
-
The photosystem-II repair cycle: updates and open questions
Planta (2024)
-
Reinvestigation on primary processes of PSII-dimer from Thermosynechococcus vulcanus by femtosecond pump-probe spectroscopy
Photosynthesis Research (2024)
-
The photosynthetic oxygen evolution does not exclude the important role and contribution of bicarbonate photolysis
Acta Geochimica (2024)
-
Zinc Provisioned Enhancement of Manganese Use Efficiency Results in Differential Biomass and Grain Production in Two Rice Cultivars Grown in Clay Loam Soil
Journal of Soil Science and Plant Nutrition (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.