Abstract
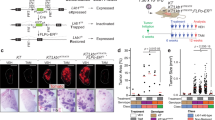
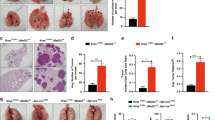
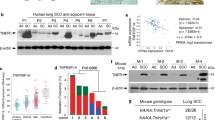
Despite the high prevalence and poor outcome of patients with metastatic lung cancer the mechanisms of tumour progression and metastasis remain largely uncharacterized. Here we modelled human lung adenocarcinoma, which frequently harbours activating point mutations in KRAS1 and inactivation of the p53 pathway2, using conditional alleles in mice3,4,5. Lentiviral-mediated somatic activation of oncogenic Kras and deletion of p53 in the lung epithelial cells of KrasLSL-G12D/+;p53flox/flox mice initiates lung adenocarcinoma development4. Although tumours are initiated synchronously by defined genetic alterations, only a subset becomes malignant, indicating that disease progression requires additional alterations. Identification of the lentiviral integration sites allowed us to distinguish metastatic from non-metastatic tumours and determine the gene expression alterations that distinguish these tumour types. Cross-species analysis identified the NK2-related homeobox transcription factor Nkx2-1 (also called Ttf-1 or Titf1) as a candidate suppressor of malignant progression. In this mouse model, Nkx2-1 negativity is pathognomonic of high-grade poorly differentiated tumours. Gain- and loss-of-function experiments in cells derived from metastatic and non-metastatic tumours demonstrated that Nkx2-1 controls tumour differentiation and limits metastatic potential in vivo. Interrogation of Nkx2-1-regulated genes, analysis of tumours at defined developmental stages, and functional complementation experiments indicate that Nkx2-1 constrains tumours in part by repressing the embryonically restricted chromatin regulator Hmga2. Whereas focal amplification of NKX2-1 in a fraction of human lung adenocarcinomas has focused attention on its oncogenic function6,7,8,9, our data specifically link Nkx2-1 downregulation to loss of differentiation, enhanced tumour seeding ability and increased metastatic proclivity. Thus, the oncogenic and suppressive functions of Nkx2-1 in the same tumour type substantiate its role as a dual function lineage factor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Rodenhuis, S. et al. Incidence and possible clinical significance of K-ras oncogene activation in adenocarcinoma of the human lung. Cancer Res. 48, 5738–5741 (1988)
Takahashi, T. et al. p53: a frequent target for genetic abnormalities in lung cancer. Science 246, 491–494 (1989)
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature Genet. 29, 418–425 (2001)
Jackson, E. L. et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res. 65, 10280–10288 (2005)
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Weir, B. A. et al. Characterizing the cancer genome in lung adenocarcinoma. Nature 450, 893–898 (2007)
Tanaka, H. et al. Lineage-specific dependency of lung adenocarcinomas on the lung development regulator TTF-1. Cancer Res. 67, 6007–6011 (2007)
Kendall, J. et al. Oncogenic cooperation and coamplification of developmental transcription factor genes in lung cancer. Proc. Natl Acad. Sci. USA 104, 16663–16668 (2007)
Kwei, K. A. et al. Genomic profiling identifies TITF1 as a lineage-specific oncogene amplified in lung cancer. Oncogene 27, 3635–3640 (2008)
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nature Protocols 4, 1064–1072 (2009)
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: A resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 52, 91–118 (2003)
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nature Med. 14, 822–827 (2008)
Nguyen, D. X. et al. WNT/TCF signaling through LEF1 and HOXB9 mediates lung adenocarcinoma metastasis. Cell 138, 51–62 (2009)
Kimura, S. et al. The T/ebp null mouse: thyroid-specific enhancer-binding protein is essential for the organogenesis of the thyroid, lung, ventral forebrain, and pituitary. Genes Dev. 10, 60–69 (1996)
Krude, H. et al. Choreoathetosis, hypothyroidism, and pulmonary alterations due to human NKX2-1 haploinsufficiency. J. Clin. Invest. 109, 475–480 (2002)
Maeda, Y., Dave, V. & Whitsett, J. A. Transcriptional control of lung morphogenesis. Physiol. Rev. 87, 219–244 (2007)
Barletta, J. A. et al. Clinical significance of TTF-1 protein expression and TTF-1 gene amplification in lung adenocarcinoma. J. Cell Mol. Med. 13, 1977–1986 (2009)
Berghmans, T. et al. Thyroid transcription factor 1—a new prognostic factor in lung cancer: a meta-analysis. Ann. Oncol. 17, 1673–1676 (2006)
Olive, K. P. et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 119, 847–860 (2004)
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nature Genet. 40, 499–507 (2008)
Nishino, J., Kim, I., Chada, K. & Morrison, S. J. Hmga2 promotes neural stem cell self-renewal in young but not old mice by reducing p16Ink4a and p19Arf expression. Cell 135, 227–239 (2008)
Yu, F. et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131, 1109–1123 (2007)
Li, O., Vasudevan, D., Davey, C. A. & Droge, P. High-level expression of DNA architectural factor HMGA2 and its association with nucleosomes in human embryonic stem cells. Genesis 44, 523–529 (2006)
Rommel, B. et al. HMGI-C, a member of the high mobility group family of proteins, is expressed in hematopoietic stem cells and in leukemic cells. Leuk. Lymphoma 26, 603–607 (1997)
Fusco, A. & Fedele, M. Roles of HMGA proteins in cancer. Nature Rev. Cancer 7, 899–910 (2007)
Meyer, B. et al. HMGA2 overexpression in non-small cell lung cancer. Mol. Carcinog. 46, 503–511 (2007)
Hristov, A. C. et al. HMGA2 protein expression correlates with lymph node metastasis and increased tumor grade in pancreatic ductal adenocarcinoma. Mod. Pathol. 22, 43–49 (2009)
Rogalla, P. et al. Expression of HMGI-C, a member of the high mobility group protein family, in a subset of breast cancers: relationship to histologic grade. Mol. Carcinog. 19, 153–156 (1997)
Di Cello, F. et al. HMGA2 participates in transformation in human lung cancer. Mol. Cancer Res. 6, 743–750 (2008)
Lee, Y. S. & Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 21, 1025–1030 (2007)
Tuveson, D. A. et al. Endogenous oncogenic K-ras(G12D) stimulates proliferation and widespread neoplastic and developmental defects. Cancer Cell 5, 375–387 (2004)
Trono, D. Lentiviral vectors: turning a deadly foe into a therapeutic agent. Gene Ther. 7, 20–23 (2000)
Wu, X., Li, Y., Crise, B. & Burgess, S. M. Transcription start regions in the human genome are favored targets for MLV integration. Science 300, 1749–1751 (2003)
Bengtsson, H., Ray, A., Spellman, P. & Speed, T. P. A single-sample method for normalizing and combining full-resolution copy numbers from multiple platforms, labs and analysis methods. Bioinformatics 25, 861–867 (2009)
Purdom, E. et al. FIRMA: a method for detection of alternative splicing from exon array data. Bioinformatics 24, 1707–1714 (2008)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
R Development Core Team . R: A language and environment for statistical computing, reference index version 2.6.1. 〈http://www.R-project.org 〉 (R Foundation for Statistical Computing, 2005)
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl Acad. Sci. USA 98, 5116–5121 (2001)
Liu, H. et al. AffyProbeMiner: a web resource for computing or retrieving accurately redefined Affymetrix probe sets. Bioinformatics 23, 2385–2390 (2007)
Assou, S. et al. A meta-analysis of human embryonic stem cells transcriptome integrated into a web-based expression atlas. Stem Cells 25, 961–973 (2007)
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009)
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010)
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006)
Kumar, M. S. et al. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc. Natl Acad. Sci. USA 105, 3903–3908 (2008)
Naviaux, R. K., Costanzi, E., Haas, M. & Verma, I. M. The pCL vector system: rapid production of helper-free, high-titer, recombinant retroviruses. J. Virol. 70, 5701–5705 (1996)
Dickins, R. A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nature Genet. 37, 1289–1295 (2005)
Luo, J. L. et al. Nuclear cytokine-activated IKKα controls prostate cancer metastasis by repressing Maspin. Nature 446, 690–694 (2007)
Morimoto-Tomita, M., Ohashi, Y., Matsubara, A., Tsuiji, M. & Irimura, T. Mouse colon carcinoma cells established for high incidence of experimental hepatic metastasis exhibit accelerated and anchorage-independent growth. Clin. Exp. Metastasis 22, 513–521 (2005)
Chiang, D. Y. et al. High-resolution mapping of copy-number alterations with massively parallel sequencing. Nature Methods 6, 99–103 (2009)
Acknowledgements
We thank A. Dooley, N. Dimitrova, T. Oliver and M. Ebert for providing reagents; M. Luo (Biology/Koch Institute BioMicro Core) for array processing; M. Kumar for experimental assistance; T. Staton, D. McFadden, A. Shaw and the entire T.J. laboratory for comments. M.M.W. was a Merck Fellow of the Damon Runyon Cancer Research Foundation and was funded by a Genentech Postdoctoral Fellowship. R.G.W.V. is supported by a Fellowship from the Dutch Cancer Society KWF. E.L.S. is supported by a training grant (T32-HL007627). D.Y.C. is supported by an Alfred P. Sloan Foundation Research Fellowship. D.M.F. is a Leukemia and Lymphoma Postdoctoral Fellow. This work was supported by National Institutes of Health grants U01-CA84306 (to T.J.) and K99-CA151968 (to M.M.W.), the Howard Hughes Medical Institute, the Ludwig Center for Molecular Oncology at MIT, and in part by the Cancer Center Support (core) grant P30-CA14051 from the National Cancer Institute. T.J. is the David H. Koch Professor of Biology and a Daniel K. Ludwig Scholar.
Author information
Authors and Affiliations
Contributions
M.M.W. and T.J. designed the study; M.M.W., T.L.D., C.K.-K. and E.L.S. performed experiments; R.G.W.V., M.M.W., C.A.W., D.D.H., S.H. and D.Y.C. conducted bioinformatic analyses; E.L.S. and R.T.B. provided pathology assistance; S.Y. and D.C. provided technical assistance; M.J.D. provided reagents; D.M.F. and M.M. gave conceptual advice; M.M.W. and T.J. wrote the paper with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Gene expression data was deposited in Gene Expression Omnibus (GSE26874).
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-13 with legends. (PDF 5817 kb)
Supplementary Table 1
This table contains the gene expression data for the TnonMet and advanced tumor-derived (TMet/Met) samples. Cell lines are listed and gene expression is shown. (XLS 10790 kb)
Supplementary Table 2
This table contains the Nkx2-1 regulated genes. Gene expression data for the TnonMet-control and TnonMet-shNkx2-1 cells is shown (XLS 6681 kb)
Rights and permissions
About this article
Cite this article
Winslow, M., Dayton, T., Verhaak, R. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011). https://doi.org/10.1038/nature09881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09881
This article is cited by
-
Transcriptome analysis reveals the essential role of NK2 homeobox 1/thyroid transcription factor 1 (NKX2-1/TTF-1) in gastric adenocarcinoma of fundic-gland type
Gastric Cancer (2023)
-
Oleic acid from cancer-associated fibroblast promotes cancer cell stemness by stearoyl-CoA desaturase under glucose-deficient condition
Cancer Cell International (2022)
-
Histone H3K9 methyltransferase SETDB1 augments invadopodia formation to promote tumor metastasis
Oncogene (2022)
-
NKX2-1 controls lung cancer progression by inducing DUSP6 to dampen ERK activity
Oncogene (2022)
-
Identification of a Novel circ_0010235/miR-1249-3p/HOXA13 Axis in Lung Adenocarcinoma
Biochemical Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.