Abstract
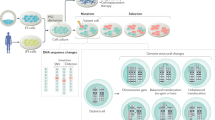
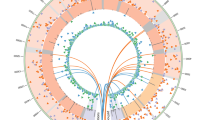
The mechanisms underlying the low efficiency of reprogramming somatic cells into induced pluripotent stem (iPS) cells are poorly understood. There is a clear need to study whether the reprogramming process itself compromises genomic integrity and, through this, the efficiency of iPS cell establishment. Using a high-resolution single nucleotide polymorphism array, we compared copy number variations (CNVs) of different passages of human iPS cells with their fibroblast cell origins and with human embryonic stem (ES) cells. Here we show that significantly more CNVs are present in early-passage human iPS cells than intermediate passage human iPS cells, fibroblasts or human ES cells. Most CNVs are formed de novo and generate genetic mosaicism in early-passage human iPS cells. Most of these novel CNVs rendered the affected cells at a selective disadvantage. Remarkably, expansion of human iPS cells in culture selects rapidly against mutated cells, driving the lines towards a genetic state resembling human ES cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Affymetrix SNP array 6.0 data from each individual sample have been deposited with the NCBI Gene Expression Omnibus(http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE26173.
References
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009)
Kaji, K. et al. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 458, 771–775 (2009)
Yu, J. et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 324, 797–801 (2009)
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010)
Kim, D. et al. Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 4, 472–476 (2009)
Utikal, J. et al. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature 460, 1145–1148 (2009)
Marión, R. M. et al. A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 460, 1149–1153 (2009)
Li, H. et al. The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature 460, 1136–1139 (2009)
Kawamura, T. et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 460, 1140–1144 (2009)
Hong, H. et al. Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature 460, 1132–1135 (2009)
Närvä, E. et al. High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity. Nature Biotechnol. 28, 371–377 (2010)
Chan, E. M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nature Biotechnol. 27, 1033–1037 (2009)
McCarroll, S. A. et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nature Genet. 40, 1166–1174 (2008)
Conrad, D. F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010)
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004)
Iafrate, A. J. et al. Detection of large-scale variation in the human genome. Nature Genet. 36, 949–951 (2004)
Hagenkord, J. M. et al. Array-based karyotyping for prognostic assessment in chronic lymphocytic leukemia: performance comparison of Affymetrix 10K2.0, 250K Nsp, and SNP6.0 arrays. J. Mol. Diagn. 12, 184–196 (2010)
Eiselleova, L. et al. A complex role for FGF-2 in self-renewal, survival, and adhesion of human embryonic stem cells. Stem Cells 27, 1847–1857 (2009)
Brill, L. M. et al. Phosphoproteomic analysis of human embryonic stem cells. Cell Stem Cell 5, 204–213 (2009)
Wang, L. et al. Self-renewal of human embryonic stem cells requires insulin-like growth factor-1 receptor and ERBB2 receptor signaling. Blood 110, 4111–4119 (2007)
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006)
Melton, C., Judson, R. L. & Blelloch, R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 463, 621–626 (2010)
Scott, G. K. et al. Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J. Biol. Chem. 282, 1479–1486 (2007)
Maimets, T., Neganova, I., Armstrong, L. & Lako, M. Activation of p53 by nutlin leads to rapid differentiation of human embryonic stem cells. Oncogene 27, 5277–5287 (2008)
Johnson, S. M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005)
Arlt, M. F. et al. Replication stress induces genome-wide copy number changes in human cells that resemble polymorphic and pathogenic variants. Am. J. Hum. Genet. 84, 339–350 (2009)
Durkin, S. G. et al. Replication stress induces tumor-like microdeletions in FHIT/FRA3B. Proc. Natl Acad. Sci. USA 105, 246–251 (2008)
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009)
Gorgoulis, V. G. et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434, 907–913 (2005)
Bartkova, J. et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434, 864–870 (2005)
Casper, A. M., Nghiem, P., Arlt, M. F. & Glover, T. W. ATR regulates fragile site stability. Cell 111, 779–789 (2002)
Arlt, M. F., Durkin, S. G., Ragland, R. L. & Glover, T. W. Common fragile sites as targets for chromosome rearrangements. DNA Repair (Amst.) 5, 1126–1135 (2006)
Esteban, M. A. et al. Vitamin C enhances the generation of mouse and human induced pluripotent stem cells. Cell Stem Cell 6, 71–79 (2010)
Kulkarni, A., Zschenker, O., Reynolds, G., Miller, D. & Murnane, J. P. Effect of telomere proximity on telomere position effect, chromosome healing, and sensitivity to DNA double-strand breaks in a human tumor cell line. Mol. Cell. Biol. 30, 578–589 (2010)
Zschenker, O. et al. Increased sensitivity of subtelomeric regions to DNA double-strand breaks in a human cancer cell line. DNA Repair (Amst.) 8, 886–900 (2009)
Linardopoulou, E. V. et al. Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 437, 94–100 (2005)
Riethman, H., Ambrosini, A. & Paul, S. Human subtelomere structure and variation. Chromosome Res. 13, 505–515 (2005)
Mayshar, Y. et al. Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 7, 521–531 (2010)
Laurent, L. C. et al. Dynamic changes in the copy number of pluripotency and cell proliferation genes in human ESCs and iPSCs during reprogramming and time in culture. Cell Stem Cell 8, 106–118 (2011)
Baker, D. E. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nature Biotechnol. 25, 207–215 (2007)
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nature Rev. Genet. 10, 551–564 (2009)
Mikkola, M. et al. Distinct differentiation characteristics of individual human embryonic stem cell lines. BMC Dev. Biol. 6, 40 (2006)
Deb-Rinker, P., Ly, D., Jezierski, A., Sikorska, M. & Walker, P. R. Sequential DNA methylation of the Nanog and Oct-4 upstream regions in human NT2 cells during neuronal differentiation. J. Biol. Chem. 280, 6257–6260 (2005)
Horn, C. et al. Splinkerette PCR for more efficient characterization of gene trap events. Nature Genet. 39, 933–934 (2007)
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454 (2006)
Ory, D. S., Neugeboren, B. A. & Mulligan, R. C. A stable human-derived packaging cell line for production of high titer retrovirus/vesicular stomatitis virus G pseudotypes. Proc. Natl Acad. Sci. USA 93, 11400–11406 (1996)
Kroon, E. et al. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo. Nature Biotechnol. 26, 443–452 (2008)
Acknowledgements
This work was supported by funding for the ESTOOLS consortium under the Sixth RFP of the EU (S.M.H., T.O., R.L. and O.B.), the Academy of Finland (T.O. and R.A.), the Finnish Cancer Organizations (R.L.), the Research Funds of the Helsinki University Hospital (T.O.), the Sigrid Jusélius Foundation (T.O.), the Stem Cell Network Canada, GL2 and CIHR (A.N.). S.M.H. is a recipient of a McEwen Post-doctoral Fellowship from the McEwen Centre for Regenerative Medicine. N.N.B. is the recipient of a New Investigator Award from the Ontario Institute for Cancer Research, through the Ontario Ministry of Research and Innovation. A.N. is Tier 1 Canada Research Chair in Stem Cells and Regeneration. We are grateful to I. Rogers, M. Faiz and K. Nagy for critically reading and editing the manuscript, and J. Ustinov, H. Sariola and J. Palgi for skilled assistance in the characterization of human iPS cells. We thank N. Rahkonen for sample processing, as well as the Finnish Microarray and Sequencing Centre (http://www.btk.fi/), and the Institute of Human Genetics, University of Bonn, for array processing.
Author information
Authors and Affiliations
Contributions
S.M.H. coordinated and performed most of the experiments in this project, analysed and interpreted the data, and prepared the manuscript. N.N.B. analysed and interpreted the data and prepared an early version of the manuscript. S.V. provided essential experimental assistance and performed some experiments in this study. R.W.C. performed, and advised on the planning of, FISH experiments. R.A. and E.N. analysed and interpreted the data and coordinated the sample processing of the arrays. O.B. and M.P. provided analysed data and samples from Illumina arrays for validation. M.S., S.N., R.H., R.T., M.M., C.O. and K.L. contributed experimentally. D.P.B-J. and K.A. provided essential experimental material and support. R.L. directed the SNP array data generation and analysis. T.O. directed, coordinated and supervised the first stage of the project and contributed to the preparation of the manuscript. A.N. prepared the manuscript, supervised the second stage of the project and helped to interpret the data. K.A. and R.L. contributed equally as co-authors. A.N. and T.O. contributed equally as senior authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-9 with Legends, Supplementary Tables 1-14 and additional references. (PDF 13911 kb)
Rights and permissions
About this article
Cite this article
Hussein, S., Batada, N., Vuoristo, S. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011). https://doi.org/10.1038/nature09871
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09871
This article is cited by
-
Xeno-free generation of human induced pluripotent stem cells from donor-matched fibroblasts isolated from dermal and oral tissues
Stem Cell Research & Therapy (2023)
-
Stem cell-based strategies and challenges for production of cultivated meat
Nature Food (2023)
-
Replication stress causes delayed mitotic entry and chromosome 12 fragility at the ANKS1B large neuronal gene in human induced pluripotent stem cells
Chromosome Research (2023)
-
Passage number affects differentiation of sensory neurons from human induced pluripotent stem cells
Scientific Reports (2022)
-
Building gut from scratch — progress and update of intestinal tissue engineering
Nature Reviews Gastroenterology & Hepatology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.