Abstract
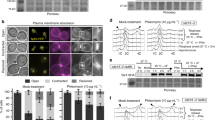
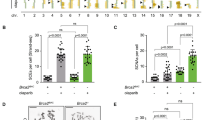
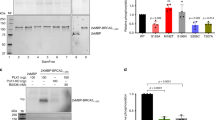
In somatic cells, Holliday junctions can be formed between sister chromatids during the recombinational repair of DNA breaks or after replication fork demise. A variety of processes act upon Holliday junctions to remove them from DNA, in events that are critical for proper chromosome segregation. In human cells, the BLM protein, inactivated in individuals with Bloom’s syndrome, acts in combination with topoisomerase IIIα, RMI1 and RMI2 (BTR complex) to promote the dissolution of double Holliday junctions1,2. Cells defective for BLM exhibit elevated levels of sister chromatid exchanges (SCEs) and patients with Bloom’s syndrome develop a broad spectrum of early-onset cancers caused by chromosome instability3. MUS81–EME1 (refs 4–7), SLX1–SLX4 (refs 8–11) and GEN1 (refs 12, 13) also process Holliday junctions but, in contrast to the BTR complex, do so by endonucleolytic cleavage. Here we deplete these nucleases from Bloom’s syndrome cells to analyse human cells compromised for the known Holliday junction dissolution/resolution pathways. We show that depletion of MUS81 and GEN1, or SLX4 and GEN1, from Bloom’s syndrome cells results in severe chromosome abnormalities, such that sister chromatids remain interlinked in a side-by-side arrangement and the chromosomes are elongated and segmented. Our results indicate that normally replicating human cells require Holliday junction processing activities to prevent sister chromatid entanglements and thereby ensure accurate chromosome condensation. This phenotype was not apparent when both MUS81 and SLX4 were depleted from Bloom’s syndrome cells, suggesting that GEN1 can compensate for their absence. Additionally, we show that depletion of MUS81 or SLX4 reduces the high frequency of SCEs in Bloom’s syndrome cells, indicating that MUS81 and SLX4 promote SCE formation, in events that may ultimately drive the chromosome instabilities that underpin early-onset cancers associated with Bloom’s syndrome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wu, L. & Hickson, I. D. The Bloom’s syndrome helicase suppresses crossing over during homologous recombination. Nature 426, 870–874 (2003)
Mankouri, H. W. & Hickson, I. D. The RecQ helicase-topoisomerase III-Rmi1 complex: a DNA structure-specific ‘dissolvasome’? Trends Biochem. Sci. 32, 538–546 (2007)
Bachrati, C. Z. & Hickson, I. D. RecQ helicases, suppressors of tumorigenesis and premature aging. Biochem. J. 374, 577–606 (2003)
Chen, X. B. et al. Human MUS81-associated endonuclease cleaves Holliday junctions in vitro. Mol. Cell 8, 1117–1127 (2001)
Ciccia, A., Constantinou, A. & West, S. C. Identification and characterization of the human MUS81/EME1 endonuclease. J. Biol. Chem. 278, 25172–25178 (2003)
Ciccia, A., McDonald, N. & West, S. C. Structural and functional relationships of the XPF/MUS81 family of proteins. Annu. Rev. Biochem. 77, 259–287 (2008)
Taylor, E. R. & McGowan, C. H. Cleavage mechanism of human MUS81–EME1 acting on Holliday-junction structures. Proc. Natl Acad. Sci. USA 105, 3757–3762 (2008)
Andersen, S. L. et al. Drosophila MUS312 and the vertebrate ortholog BTBD12 interact with DNA structure-specific endonucleases in DNA repair and recombination. Mol. Cell 35, 128–135 (2009)
Fekairi, S. et al. Human SLX4 is a Holliday junction resolvase subunit that binds multiple DNA repair/recombination endonucleases. Cell 138, 78–89 (2009)
Munoz, I. M. et al. Coordination of structure-specific nucleases by human SLX4/BTBD12 is required for DNA repair. Mol. Cell 35, 116–127 (2009)
Svendsen, J. M. et al. Mammalian BTBD12/SLX4 assembles a Holliday junction resolvase and is required for DNA repair. Cell 138, 63–77 (2009)
Ip, S. C. Y. et al. Identification of Holliday junction resolvases from humans and yeast. Nature 456, 357–361 (2008)
Rass, U. et al. Mechanism of Holliday junction resolution by the human GEN1 protein. Genes Dev. 24, 1559–1569 (2010)
Osman, F., Dixon, J., Doe, C. L. & Whitby, M. C. Generating crossovers by resolution of nicked Holliday junctions: a role of Mus81–Eme1 in meiosis. Mol. Cell 12, 761–774 (2003)
Gaillard, P.-H. L., Noguchi, E., Shanahan, P. & Russell, P. The endogenous Mus81–Eme1 complex resolves Holliday junctions by a nick and couternick mechanism. Mol. Cell 12, 747–759 (2003)
Blanco, M. G., Matos, J., Rass, U., Ip, S. C. Y. & West, S. C. Functional overlap between the structure-specific nucleases Yen1 and Mus81-Mms4 for DNA damage repair in S. cerevisiae. DNA Repair (Amst.) 9, 394–402 (2010)
Tay, Y. D. & Wu, L. Overlapping roles for Yen1 and Mus81 in cellular Holliday junction processing. J. Biol. Chem. 285, 11427–11432 (2010)
Ho, C. K., Mazón, G., Lam, A. F. & Symington, L. S. Mus81 and Yen1 promote reciprocal exchange during mitotic recombination to maintain genome integrity in budding yeast. Mol. Cell 40, 988–1000 (2011)
Breger, K. S., Smith, L., Turker, M. S. & Thayer, M. J. Ionizing radiation induces frequent translocations with delayed replication and condensation. Cancer Res. 64, 8231–8238 (2004)
Hearst, J., Kauffman, L. & McClain, W. A simple mechanism for the avoidance of entanglement during chromosome replication. Trends Genet. 14, 244–247 (1998)
Chan, K. L., North, P. S. & Hickson, I. D. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. EMBO J. 26, 3397–3409 (2007)
Chan, K. L., Palmai-Pallag, T., Ying, S. M. & Hickson, I. D. Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nature Cell Biol. 11, 753–760 (2009)
Naim, V. & Rosselli, F. The FANC pathway and BLM collaborate during mitosis to prevent micro-nucleation and chromosome abnormalities. Nature Cell Biol. 11, 761–768 (2009)
Baumann, C., Korner, R., Hofmann, K. & Nigg, E. A. PICH, a centromere-associated SNF2 family ATPase, is regulated by Plk1 and required for the spindle checkpoint. Cell 128, 101–114 (2007)
Wu, L., Davies, S. L., Levitt, N. C. & Hickson, I. D. Potential role for the BLM helicase in recombinational repair via a conserved interaction with RAD51. J. Biol. Chem. 276, 19375–19381 (2001)
Ellis, N. A., Proytcheva, M., Sanz, M. M., Ye, T.-Z. & German, J. Transfection of BLM into cultured Bloom syndrome cells reduced the sister-chromatid exchange rate toward normal. Am. J. Hum. Genet. 65, 1368–1374 (1999)
Gaymes, T. J. et al. Increased error-prone non homologous DNA end-joining – a proposed mechanism of chromosomal instability in Bloom’s syndrome. Oncogene 21, 2525–2533 (2002)
Bender, C. F. et al. Cancer predisposition and hematopoietic failure in Rad50(S/S) mice. Genes Dev. 16, 2237–2251 (2002)
Bayani, J. & Squire, J. A. Sister chromatid exchange. Curr. Protoc. Cell Biol. 22 7. (2005)
Alsop, A. E., Teschendorff, A. E. & Edwards, P. A. W. Distribution of breakpoints on chromosome 18 in breast, colorectal, and pancreatic carcinoma cell lines. Cancer Genet. Cytogenet. 164, 97–109 (2006)
Acknowledgements
We thank I. Hickson for providing the Bloom’s syndrome cell lines and advice, P. Edwards for help and providing facilities for chromosome painting, S. Horswell for the statistical analysis, S. Ip for the GEN1 antibody, M.G. Blanco for assistance with SCE scoring and our laboratory colleagues for their encouragement and suggestions. We further thank K. Cimprich, the Cimprich laboratory members, and C. Wang, W. Johnson and A. Straight. This work was supported by Cancer Research UK, the Louis-Jeantet Foundation, the European Research Council, the Swiss Bridge Foundation and the Breast Cancer Campaign. S.N. was supported by a studentship from the UK Medical Research Council.
Author information
Authors and Affiliations
Contributions
T.W. and S.C.W. designed the project that was undertaken entirely by T.W. Expertise for the chromosome paints was provided by S.N. The manuscript was written by S.C.W. with help from T.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
The file contains Supplementary Figures 1-9 with legends. (PDF 3149 kb)
Rights and permissions
About this article
Cite this article
Wechsler, T., Newman, S. & West, S. Aberrant chromosome morphology in human cells defective for Holliday junction resolution. Nature 471, 642–646 (2011). https://doi.org/10.1038/nature09790
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09790
This article is cited by
-
Structural insights into sequence-dependent Holliday junction resolution by the chloroplast resolvase MOC1
Nature Communications (2020)
-
The MRE11 nuclease promotes homologous recombination not only in DNA double-strand break resection but also in post-resection in human TK6 cells
Genome Instability & Disease (2020)
-
Nuclear and cytoplasmic WDR-23 isoforms mediate differential effects on GEN-1 and SKN-1 substrates
Scientific Reports (2019)
-
Keeping ribosomal DNA intact: a repeating challenge
Chromosome Research (2019)
-
Unresolved recombination intermediates lead to ultra-fine anaphase bridges, chromosome breaks and aberrations
Nature Cell Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.