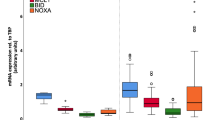
Abstract
Microtubules have pivotal roles in fundamental cellular processes and are targets of antitubulin chemotherapeutics1. Microtubule-targeted agents such as Taxol and vincristine are prescribed widely for various malignancies, including ovarian and breast adenocarcinomas, non-small-cell lung cancer, leukaemias and lymphomas1. These agents arrest cells in mitosis and subsequently induce cell death through poorly defined mechanisms2. The strategies that resistant tumour cells use to evade death induced by antitubulin agents are also unclear2. Here we show that the pro-survival protein MCL1 (ref. 3) is a crucial regulator of apoptosis triggered by antitubulin chemotherapeutics. During mitotic arrest, MCL1 protein levels decline markedly, through a post-translational mechanism, potentiating cell death. Phosphorylation of MCL1 directs its interaction with the tumour-suppressor protein FBW7, which is the substrate-binding component of a ubiquitin ligase complex. The polyubiquitylation of MCL1 then targets it for proteasomal degradation. The degradation of MCL1 was blocked in patient-derived tumour cells that lacked FBW7 or had loss-of-function mutations in FBW7, conferring resistance to antitubulin agents and promoting chemotherapeutic-induced polyploidy. Additionally, primary tumour samples were enriched for FBW7 inactivation and elevated MCL1 levels, underscoring the prominent roles of these proteins in oncogenesis. Our findings suggest that profiling the FBW7 and MCL1 status of tumours, in terms of protein levels, messenger RNA levels and genetic status, could be useful to predict the response of patients to antitubulin chemotherapeutics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 March 2011
Minor errors in the HTML and PDF were corrected on 09March 2011.
21 November 2011
Supplementary Fig. S21 contained a duplicated panel instead of the SB203580-treated cdc27 blot.
References
Jackson, J. R., Patrick, D. R., Dar, M. M. & Huang, P. S. Targeted anti-mitotic therapies: can we improve on tubulin agents? Nature Rev. Cancer 7, 107–117 (2007)
Rieder, C. L. & Maiato, H. Stuck in division or passing through: what happens when cells cannot satisfy the spindle assembly checkpoint. Dev. Cell 7, 637–651 (2004)
Youle, R. J. & Strasser, A. The BCL-2 protein family: opposing activities that mediate cell death. Nature Rev. Mol. Cell Biol. 9, 47–59 (2008)
Willis, S. N. et al. Apoptosis initiated when BH3 ligands engage multiple Bcl-2 homologs, not Bax or Bak. Science 315, 856–859 (2007)
Varfolomeev, E. & Vucic, D. (Un)expected roles of c-IAPs in apoptotic and NFκB signaling pathways. Cell Cycle 7, 1511–1521 (2008)
King, R. W. et al. A 20S complex containing CDC27 and CDC16 catalyzes the mitosis-specific conjugation of ubiquitin to cyclin B. Cell 81, 279–288 (1995)
Finley, D. Recognition and processing of ubiquitin–protein conjugates by the proteasome. Annu. Rev. Biochem. 78, 477–513 (2009)
Frescas, D. & Pagano, M. Deregulated proteolysis by the F-box proteins SKP2 and β-TrCP: tipping the scales of cancer. Nature Rev. Cancer 8, 438–449 (2008)
Welcker, M. & Clurman, B. E. FBW7 ubiquitin ligase: a tumour suppressor at the crossroads of cell division, growth and differentiation. Nature Rev. Cancer 8, 83–93 (2008)
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009)
Zhong, Q., Gao, W., Du, F. & Wang, X. Mule/ARF-BP1, a BH3-only E3 ubiquitin ligase, catalyzes the polyubiquitination of Mcl-1 and regulates apoptosis. Cell 121, 1085–1095 (2005)
Potapova, T. A. et al. The reversibility of mitotic exit in vertebrate cells. Nature 440, 954–958 (2006)
Huang, H. C., Shi, J., Orth, J. D. & Mitchison, T. J. Evidence that mitotic exit is a better cancer therapeutic target than spindle assembly. Cancer Cell 16, 347–358 (2009)
Domina, A. M., Vrana, J. A., Gregory, M. A., Hann, S. R. & Craig, R. W. MCL1 is phosphorylated in the PEST region and stabilized upon ERK activation in viable cells, and at additional sites with cytotoxic okadaic acid or Taxol. Oncogene 23, 5301–5315 (2004)
Gascoigne, K. E. & Taylor, S. S. Cancer cells display profound intra- and interline variation following prolonged exposure to antimitotic drugs. Cancer Cell 14, 111–122 (2008)
Finkin, S., Aylon, Y., Anzi, S., Oren, M. & Shaulian, E. Fbw7 regulates the activity of endoreduplication mediators and the p53 pathway to prevent drug-induced polyploidy. Oncogene 27, 4411–4421 (2008)
Terrano, D. T., Upreti, M. & Chambers, T. C. Cyclin-dependent kinase 1-mediated Bcl-xL/Bcl-2 phosphorylation acts as a functional link coupling mitotic arrest and apoptosis. Mol. Cell. Biol. 30, 640–656 (2010)
Ozalp, S. S., Yalcin, O. T., Tanir, M., Kabukcuoglu, S. & Etiz, E. Multidrug resistance gene-1 (Pgp) expression in epithelial ovarian malignancies. Eur. J. Gynaecol. Oncol. 23, 337–340 (2002)
Mesquita, B. et al. No significant role for β tubulin mutations and mismatch repair defects in ovarian cancer resistance to paclitaxel/cisplatin. BMC Cancer 5, 101 (2005)
Chen, L. et al. Differential targeting of prosurvival Bcl-2 proteins by their BH3-only ligands allows complementary apoptotic function. Mol. Cell 17, 393–403 (2005)
Acknowledgements
We thank P. Ekert for FDM cell lines, J. Stinson for sequencing assistance, S. Johnson and C. Santos for assistance with obtaining patient samples, P. Haverty for bioinformatics analysis, C. Grimaldi for cloning assistance, J. Dynek for TaqMan advice, D. French for tumour analysis, C. Quan and J. Tom for peptide synthesis, I. Zilberleyb and the Baculovirus Expression Group for cloning and protein production, S. Charuvu for generating MCL1 point mutants, A. Bruce for graphics assistance, the Genentech Cancer Genome Project Team, Z. Modrusan, R. Soriano and the microarray lab for ovarian tumour data sets, K. Newton for editorial assistance, W. Wei for sharing unpublished results, and A. Eldridge, D. Kirkpatrick, D. Vucic, E. Varfolomeev, T. Goncharov, A. Cochran, O. Huang, A. Huang, Y. Pereg, A. Loktev, D. Phillips, J. Wu, M. van Delft, D. Eaton, E. Shaulian, T. Hunter, S. Cory, J. Adams, A. Strasser, R. Deshaies and G. Evan for discussions. Work in the Huang laboratory is supported by the National Health and Medical Research Council (program grant #461221, IRIISS grant #361646 and a fellowship to D.C.S.H.), the Leukemia and Lymphoma Society (SCOR 7413), the National Institutes of Health (grants CA043540 and CA80188), the Australian Cancer Research Foundation, and an Australian Research Council Australian Postdoctoral fellowship to T.O. We apologize to our colleagues whose primary work could not be cited owing to space constraints.
Author information
Authors and Affiliations
Contributions
I.E.W., S.K., T.O., J.A.E., P.B.K., A.R.J., C.L., E.C.D., E.H., H.M. and K.G.L. designed and performed in vitro, cell-based and in vivo experiments. D.J.A. and M.J.C.L. designed and performed microscopy experiments. S.K., C.L., K.M.O., M.L.C. and M.E. made constructs. J.L. and J.K. performed bioinformatics analysis, K.P. and S.S. provided sequencing analysis. W.S. and J.R.L. designed and performed mass spectrometry experiments. I.E.W., S.K., C.L., T.O., W.S., D.J.A., M.J.C.L., K.G.L., E.C.D., H.M. and V.M.D. prepared the manuscript and figures. W.S., L.D.B., P.K.J., W.J.F., D.J.A., P.B.K., A.R.J., M.J.C.L., H.M., D.C.S.H. and I.E.W. contributed to the study design and data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-39 with legends, Supplementary Methods, additional references and Supplementary Tables 1-2. (PDF 6785 kb)
Supplementary Figure
This file contains a corrected version of Supplementary Figure S21 and legend. This file was uploaded on 21 November 2011. (PDF 294 kb)
Rights and permissions
About this article
Cite this article
Wertz, I., Kusam, S., Lam, C. et al. Sensitivity to antitubulin chemotherapeutics is regulated by MCL1 and FBW7. Nature 471, 110–114 (2011). https://doi.org/10.1038/nature09779
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09779
This article is cited by
-
FBXW7 regulates the sensitivity of imatinib in gastrointestinal stromal tumors by targeting MCL1
Gastric Cancer (2024)
-
Transient targeting of BIM-dependent adaptive MCL1 preservation enhances tumor response to molecular therapeutics in non-small cell lung cancer
Cell Death & Differentiation (2023)
-
MARCH5 regulates mitotic apoptosis through MCL1-dependent and independent mechanisms
Cell Death & Differentiation (2023)
-
FBXW7 tumor suppressor regulation by dualspecificity tyrosine-regulated kinase 2
Cell Death & Disease (2023)
-
Selective MCL-1 inhibitor ABBV-467 is efficacious in tumor models but is associated with cardiac troponin increases in patients
Communications Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.