Abstract
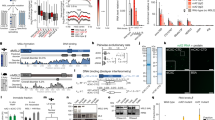
The evolution of sex chromosomes has resulted in numerous species in which females inherit two X chromosomes but males have a single X, thus requiring dosage compensation. MSL (Male-specific lethal) complex increases transcription on the single X chromosome of Drosophila males to equalize expression of X-linked genes between the sexes1. The biochemical mechanisms used for dosage compensation must function over a wide dynamic range of transcription levels and differential expression patterns. It has been proposed2 that the MSL complex regulates transcriptional elongation to control dosage compensation, a model subsequently supported by mapping of the MSL complex and MSL-dependent histone 4 lysine 16 acetylation to the bodies of X-linked genes in males, with a bias towards 3′ ends3,4,5,6,7. However, experimental analysis of MSL function at the mechanistic level has been challenging owing to the small magnitude of the chromosome-wide effect and the lack of an in vitro system for biochemical analysis. Here we use global run-on sequencing (GRO-seq)8 to examine the specific effect of the MSL complex on RNA Polymerase II (RNAP II) on a genome-wide level. Results indicate that the MSL complex enhances transcription by facilitating the progression of RNAP II across the bodies of active X-linked genes. Improving transcriptional output downstream of typical gene-specific controls may explain how dosage compensation can be imposed on the diverse set of genes along an entire chromosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gelbart, M. E. & Kuroda, M. I. Drosophila dosage compensation: a complex voyage to the X chromosome. Development 136, 1399–1410 (2009)
Lucchesi, J. C. Dosage compensation in flies and worms: the ups and downs of X-chromosome regulation. Curr. Opin. Genet. Dev. 8, 179–184 (1998)
Smith, E. R., Allis, C. D. & Lucchesi, J. C. Linking global histone acetylation to the transcription enhancement of X-chromosomal genes in Drosophila males. J. Biol. Chem. 276, 31483–31486 (2001)
Alekseyenko, A. A., Larschan, E., Lai, W. R., Park, P. J. & Kuroda, M. I. High-resolution ChIP-chip analysis reveals that the Drosophila MSL complex selectively identifies active genes on the male X chromosome. Genes Dev. 20, 848–857 (2006)
Gilfillan, G. D. et al. Chromosome-wide gene-specific targeting of the Drosophila dosage compensation complex. Genes Dev. 20, 858–870 (2006)
Kind, J. et al. Genome-wide analysis reveals MOF as a key regulator of dosage compensation and gene expression in Drosophila. Cell 133, 813–828 (2008)
Gelbart, M. E., Larschan, E., Peng, S., Park, P. J. & Kuroda, M. I. Drosophila MSL complex globally acetylates H4K16 on the male X chromosome for dosage compensation. Nature Struct. Mol. Biol. 16, 825–832 (2009)
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008)
Hamada, F. N., Park, P. J., Gordadze, P. R. & Kuroda, M. I. Global regulation of X chromosomal genes by the MSL complex in Drosophila melanogaster. Genes Dev. 19, 2289–2294 (2005)
Zhang, Y. et al. Expression in aneuploid Drosophila S2 cells. PLoS Biol. 8, e1000320 (2010)
Muse, G. W. et al. RNA polymerase is poised for activation across the genome. Nature Genet. 39, 1507–1511 (2007)
Zeitlinger, J. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nature Genet. 39, 1512–1516 (2007)
Belote, J. M. & Lucchesi, J. C. Male-specific lethal mutations of Drosophila melanogaster. Genetics 96, 165–186 (1980)
Bai, X., Alekseyenko, A. A. & Kuroda, M. I. Sequence-specific targeting of MSL complex regulates transcription of the roX RNA genes. EMBO J. 23, 2853–2861 (2004)
Meller, V. H. & Rattner, B. P. The roX genes encode redundant male-specific lethal transcripts required for targeting of the MSL complex. EMBO J. 21, 1084–1091 (2002)
Hilfiker, A., Hilfiker-Kleiner, D., Pannuti, A. & Lucchesi, J. C. mof, a putative acetyl transferase gene related to the Tip60 and MOZ human genes and to the SAS genes of yeast, is required for dosage compensation in Drosophila. EMBO J. 16, 2054–2060 (1997)
Wang, Z. et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nature Genet. 40, 897–903 (2008)
Kaplan, C. D., Laprade, L. & Winston, F. Transcription elongation factors repress transcription initiation from cryptic sites. Science 301, 1096–1099 (2003)
Mavrich, T. N. et al. Nucleosome organization in the Drosophila genome. Nature 453, 358–362 (2008)
Mavrich, T. N. et al. A barrier nucleosome model for statistical positioning of nucleosomes throughout the yeast genome. Genome Res. 18, 1073–1083 (2008)
Belotserkovskaya, R. et al. FACT facilitates transcription-dependent nucleosome alteration. Science 301, 1090–1093 (2003)
Robinson, P. J. et al. 30 nm chromatin fibre decompaction requires both H4–K16 acetylation and linker histone eviction. J. Mol. Biol. 381, 816–825 (2008)
Shogren-Knaak, M. et al. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006)
Park, Y., Kelley, R. L., Oh, H., Kuroda, M. I. & Meller, V. H. Extent of chromatin spreading determined by roX RNA recruitment of MSL proteins. Science 298, 1620–1623 (2002)
Oh, H., Park, Y. & Kuroda, M. I. Local spreading of MSL complexes from roX genes on the Drosophila X chromosome. Genes Dev. 17, 1334–1339 (2003)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Nechaev, S. et al. Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327, 335–338 (2010)
Rozowsky, J. et al. PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls. Nature Biotechnol. 27, 66–75 (2009)
Acknowledgements
We thank F. M. Winston, S. Buratowski, A. Alekseyenko, M. Gelbart, C. Wang and A. Gortchakov for comments on the manuscript, and are grateful to N. Gehlenborg for graphic design expertise. This work was supported by the following NIH grants: GM45744 (M.I.K.), GM082798 (P.J.P.) and HG4845 (J.T.L.). E.L. was supported by a Charles A. King Trust fellowship from the Medical Foundation.
Author information
Authors and Affiliations
Contributions
E.L. performed the experiments and E.P.B. and P.V.K. performed the computational analyses. P.J.P. advised on the computational analyses and the manuscript preparation. L.J.C., J.T.L. and M.I.K. advised on experimental protocols and/or design. E.L. and M.I.K. prepared the manuscript.
Corresponding authors
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-10 with legends, Supplementary Tables 1-3 and additional references. (PDF 13927 kb)
Rights and permissions
About this article
Cite this article
Larschan, E., Bishop, E., Kharchenko, P. et al. X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila. Nature 471, 115–118 (2011). https://doi.org/10.1038/nature09757
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09757
This article is cited by
-
H4K16ac activates the transcription of transposable elements and contributes to their cis-regulatory function
Nature Structural & Molecular Biology (2023)
-
The zinc finger protein CLAMP promotes long-range chromatin interactions that mediate dosage compensation of the Drosophila male X-chromosome
Epigenetics & Chromatin (2021)
-
Interaction of Male Specific Lethal complex and genomic imbalance on global gene expression in Drosophila
Scientific Reports (2021)
-
MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
Nature Communications (2019)
-
Non-canonical Drosophila X chromosome dosage compensation and repressive topologically associated domains
Epigenetics & Chromatin (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.