Abstract
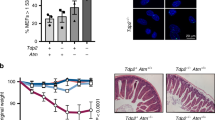
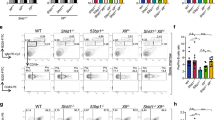
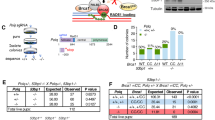
Misrepair of DNA double-strand breaks produced by the V(D)J recombinase (the RAG1/RAG2 proteins) at immunoglobulin (Ig) and T cell receptor (Tcr) loci has been implicated in pathogenesis of lymphoid malignancies in humans1 and in mice2,3,4,5,6,7. Defects in DNA damage response factors such as ataxia telangiectasia mutated (ATM) protein and combined deficiencies in classical non-homologous end joining and p53 predispose to RAG-initiated genomic rearrangements and lymphomagenesis2,3,4,5,6,7,8,9,10,11. Although we showed previously that RAG1/RAG2 shepherd the broken DNA ends to classical non-homologous end joining for proper repair12,13, roles for the RAG proteins in preserving genomic stability remain poorly defined. Here we show that the RAG2 carboxy (C) terminus, although dispensable for recombination14,15, is critical for maintaining genomic stability. Thymocytes from ‘core’ Rag2 homozygotes (Rag2c/c mice) show dramatic disruption of Tcrα/δ locus integrity. Furthermore, all Rag2c/c p53−/− mice, unlike Rag1c/c p53−/− and p53−/− animals, rapidly develop thymic lymphomas bearing complex chromosomal translocations, amplifications and deletions involving the Tcrα/δ and Igh loci. We also find these features in lymphomas from Atm−/− mice. We show that, like ATM-deficiency3, core RAG2 severely destabilizes the RAG post-cleavage complex. These results reveal a novel genome guardian role for RAG2 and suggest that similar ‘end release/end persistence’ mechanisms underlie genomic instability and lymphomagenesis in Rag2c/c p53−/− and Atm−/− mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kuppers, R. & Dalla-Favera, R. Mechanisms of chromosomal translocations in B cell lymphomas. Oncogene 20, 5580–5594 (2001)
Callen, E. et al. ATM prevents the persistence and propagation of chromosome breaks in lymphocytes. Cell 130, 63–75 (2007)
Bredemeyer, A. L. et al. ATM stabilizes DNA double-strand-break complexes during V(D)J recombination. Nature 442, 466–470 (2006)
Zhu, C. et al. Unrepaired DNA breaks in p53-deficient cells lead to oncogenic gene amplification subsequent to translocations. Cell 109, 811–821 (2002)
Gao, Y. et al. Interplay of p53 and DNA-repair protein XRCC4 in tumorigenesis, genomic stability and development. Nature 404, 897–900 (2000)
Difilippantonio, M. J. et al. DNA repair protein Ku80 suppresses chromosomal aberrations and malignant transformation. Nature 404, 510–514 (2000)
Zha, S. et al. ATM-deficient thymic lymphoma is associated with aberrant tcrd rearrangement and gene amplification. J. Exp. Med. 207, 1369–1380 (2010)
Callen, E. et al. Chimeric IgH-TCRα/δ translocations in T lymphocytes mediated by RAG. Cell Cycle 8, 2408–2412 (2009)
Matei, I. R. et al. ATM deficiency disrupts Tcra locus integrity and the maturation of CD4+CD8+ thymocytes. Blood 109, 1887–1896 (2007)
Liyanage, M. et al. Abnormal rearrangement within the α/δ T-cell receptor locus in lymphomas from Atm-deficient mice. Blood 96, 1940–1946 (2000)
Barlow, C. et al. Atm-deficient mice: a paradigm of ataxia telangiectasia. Cell 86, 159–171 (1996)
Corneo, B. et al. Rag mutations reveal robust alternative end joining. Nature 449, 483–486 (2007)
Lee, G. S., Neiditch, M. B., Salus, S. S. & Roth, D. B. RAG proteins shepherd double-strand breaks to a specific pathway, suppressing error-prone repair, but RAG nicking initiates homologous recombination. Cell 117, 171–184 (2004)
Jones, J. M. & Simkus, C. The roles of the RAG1 and RAG2 “non-core” regions in V(D)J recombination and lymphocyte development. Arch. Immunol. Ther. Exp. (Warsz.) 57, 105–116 (2009)
Liang, H. E. et al. The “dispensable” portion of RAG2 is necessary for efficient V-to-DJ rearrangement during B and T cell development. Immunity 17, 639–651 (2002)
Qiu, J. X., Kale, S. B., Yarnell Schultz, H. & Roth, D. B. Separation-of-function mutants reveal critical roles for RAG2 in both the cleavage and joining steps of V(D)J recombination. Mol. Cell 7, 77–87 (2001)
Steen, S. B., Han, J.-O., Mundy, C., Oettinger, M. A. & Roth, D. B. Roles of the “dispensable” portions of RAG-1 and RAG-2 in V(D)J recombination. Mol. Cell. Biol. 19, 3010–3017 (1999)
Curry, J. D. & Schlissel, M. S. RAG2’s non-core domain contributes to the ordered regulation of V(D)J recombination. Nucleic Acids Res. 36, 5750–5762 (2008)
Talukder, S. R., Dudley, D. D., Alt, F. W., Takahama, Y. & Akamatsu, Y. Increased frequency of aberrant V(D)J recombination products in core RAG-expressing mice. Nucleic Acids Res. 32, 4539–4549 (2004)
Jacks, T. et al. Tumor spectrum analysis in p53-mutant mice. Curr. Biol. 4, 1–7 (1994)
Liao, M. J. et al. No requirement for V(D)J recombination in p53-deficient thymic lymphoma. Mol. Cell. Biol. 18, 3495–3501 (1998)
Forster, A., Hobart, M., Hengartner, H. & Rabbitts, T. H. An immunoglobulin heavy-chain gene is altered in two T-cell clones. Nature 286, 897–899 (1980)
Haines, B. B. et al. Block of T cell development in P53-deficient mice accelerates development of lymphomas with characteristic RAG-dependent cytogenetic alterations. Cancer Cell 9, 109–120 (2006)
Dudley, D. D. et al. Impaired V(D)J recombination and lymphocyte development in core RAG1-expressing mice. J. Exp. Med. 198, 1439–1450 (2003)
Difilippantonio, S. et al. 53BP1 facilitates long-range DNA end-joining during V(D)J recombination. Nature 456, 529–533 (2008)
Arnal, S. M., Holub, A. J., Salus, S. S. & Roth, D. B. Non-consensus heptamer sequences destabilize the RAG post-cleavage complex, making ends available to alternative DNA repair pathways. Nucleic Acids Res. 38, 2944–2954 (2010)
Helmink, B. A. et al. MRN complex function in the repair of chromosomal Rag-mediated DNA double-strand breaks. J. Exp. Med. 206, 669–679 (2009)
Deriano, L., Stracker, T. H., Baker, A., Petrini, J. H. & Roth, D. B. Roles for NBS1 in alternative nonhomologous end-joining of V(D)J recombination intermediates. Mol. Cell 34, 13–25 (2009)
Simsek, D. & Jasin, M. Alternative end-joining is suppressed by the canonical NHEJ component Xrcc4-ligase IV during chromosomal translocation formation. Nature Struct. Mol. Biol. 17, 410–416 (2010)
Li, Z., Dordai, D. I., Lee, J. & Desiderio, S. A conserved degradation signal regulates RAG-2 accumulation during cell division and links V(D)J recombination to the cell cycle. Immunity 5, 575–589 (1996)
Theunissen, J. W. & Petrini, J. H. Methods for studying the cellular response to DNA damage: influence of the Mre11 complex on chromosome metabolism. Methods Enzymol. 409, 251–284 (2006)
Multani, A. S. et al. Caspase-dependent apoptosis induced by telomere cleavage and TRF2 loss. Neoplasia 2, 339–345 (2000)
Pathak, S. Chromosome banding techniques. J. Reprod. Med. 17, 25–28 (1976)
Hewitt, S. L. et al. RAG-1 and ATM coordinate monoallelic recombination and nuclear positioning of immunoglobulin loci. Nature Immunol. 10, 655–664 (2009)
Skok, J. A. et al. Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nature Immunol. 8, 378–387 (2007)
Yang, Y. H. et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 30, e15 (2002)
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004)
Aguirre, A. J. et al. High-resolution characterization of the pancreatic adenocarcinoma genome. Proc. Natl Acad. Sci. USA 101, 9067–9072 (2004)
R. development Core Team. R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing. (2006)
Acknowledgements
We thank M. Schlissel for the gift of core Rag2 mice, F. Alt for the gift of core Rag1 mice and S. Hewitt for the Igh BAC probes. D.B.R. was supported by National Institutes of Health Roadmap Initiative in Nanomedicine through a Nanomedicine Development Center award (1PN2EY018244), a National Institutes of Health grant CA104588 and the Irene Diamond Fund. L.D. is a Fellow of The Leukemia and Lymphoma Society. A.V.A. was supported in part by grant 1UL1RR029893 from the National Center for Research Resources, National Institutes of Health. J.A.S. was supported by a National Institutes of Health grant R01GM086852, a National Institutes of Health Challenge grant NCI R01CA145746-01, a Leukemia and Lymphoma Scholar Award and a Wellcome trust project grant 085096.
Author information
Authors and Affiliations
Contributions
L.D. and D.B.R. conceived the study and co-wrote the manuscript. L.D. designed the experiments. L.D., J.C., M.C. and A.M. performed the experiments. Y.C. provided assistance with the mouse colonies. A.V.A. performed the aCGH data analysis. J.A.S. and S.C. provided technical and conceptual support. J.C. and J.A.S revised the manuscript. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-13 with legends, Supplementary Table 1 and an additional reference. (PDF 12155 kb)
Rights and permissions
About this article
Cite this article
Deriano, L., Chaumeil, J., Coussens, M. et al. The RAG2 C terminus suppresses genomic instability and lymphomagenesis. Nature 471, 119–123 (2011). https://doi.org/10.1038/nature09755
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09755
This article is cited by
-
PAXX promotes KU accumulation at DNA breaks and is essential for end-joining in XLF-deficient mice
Nature Communications (2017)
-
RAG2 and XLF/Cernunnos interplay reveals a novel role for the RAG complex in DNA repair
Nature Communications (2016)
-
Mechanisms of human lymphoid chromosomal translocations
Nature Reviews Cancer (2016)
-
Neuroblastoma: oncogenic mechanisms and therapeutic exploitation of necroptosis
Cell Death & Disease (2015)
-
Histone methylation and V(D)J recombination
International Journal of Hematology (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.