Abstract
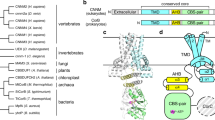
Gram-negative bacteria, such as Escherichia coli, expel toxic chemicals through tripartite efflux pumps that span both the inner and outer membrane. The three parts are an inner membrane, substrate-binding transporter; a membrane fusion protein; and an outer-membrane-anchored channel. The fusion protein connects the transporter to the channel within the periplasmic space. A crystallographic model of this tripartite efflux complex has been unavailable because co-crystallization of the various components of the system has proven to be extremely difficult. We previously described the crystal structures of both the inner membrane transporter CusA1 and the membrane fusion protein CusB2 of the CusCBA efflux system3,4 of E. coli. Here we report the co-crystal structure of the CusBA efflux complex, showing that the transporter (or pump) CusA, which is present as a trimer, interacts with six CusB protomers and that the periplasmic domain of CusA is involved in these interactions. The six CusB molecules seem to form a continuous channel. The affinity of the CusA and CusB interaction was found to be in the micromolar range. Finally, we have predicted a three-dimensional structure for the trimeric CusC outer membrane channel and developed a model of the tripartite efflux assemblage. This CusC3–CusB6–CusA3 model shows a 750-kilodalton efflux complex that spans the entire bacterial cell envelope and exports Cu i and Ag i ions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Atomic coordinates and structure factors for the CusBA structure have been deposited with the Protein Data Bank (http://www.pdb.org) under accession code 3NE5.
References
Long, F. et al. Crystal structures of the CusA heavy-metal efflux pump suggest methionine-mediated metal transport mechanism. Nature 467, 484–488 (2010)
Su, C.-C. et al. Crystal structure of the membrane fusion protein CusB from Escherichia coli. J. Mol. Biol. 393, 342–355 (2009)
Franke, S., Grass, G. & Nies, D. H. The product of the ybdE gene of the Escherichia coli chromosome is involved in detoxification of silver ions. Microbiology 147, 965–972 (2001)
Franke, S., Grass, G., Rensing, C. & Nies, D. H. Molecular analysis of the copper-transporting efflux system CusCFBA of Escherichia coli. J. Bacteriol. 185, 3804–3812 (2003)
Tseng, T. T. et al. The RND permease superfamily: an ancient, ubiquitous and diverse family that includes human disease and development proteins. J. Mol. Microbiol. Biotechnol. 1, 107–125 (1999)
Nies, D. H. Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol. Rev. 27, 313–339 (2003)
Murakami, S., Nakashima, R., Yamashita, E. & Yamaguchi, A. Crystal structure of bacterial multidrug efflux transporter AcrB. Nature 419, 587–593 (2002)
Yu, E. W., McDermott, G., Zgruskaya, H. I., Nikaido, H. & Koshland, D. E., Jr Structural basis of multiple drug binding capacity of the AcrB multidrug efflux pump. Science 300, 976–980 (2003)
Murakami, S., Nakashima, R., Yamashita, E., Matsumoto, T. & Yamaguchi, A. Crystal structures of a multidrug transporter reveal a functionally rotating mechanism. Nature 443, 173–179 (2006)
Seeger, M. A. et al. Structural asymmetry of AcrB trimer suggests a peristaltic pump mechanism. Science 313, 1295–1298 (2006)
Sennhauser, G., Amstutz, P., Briand, C., Storchengegger, O. & Grütter, M. G. Drug export pathway of multidrug exporter AcrB revealed by DARPin inhibitors. PLoS Biol. 5, e7 (2007)
Yu, E. W., Aires, J. R., McDermott, G. & Nikaido, H. A periplasmic-drug binding site of the AcrB multidrug efflux pump: a crystallographic and site-directed mutagenesis study. J. Bacteriol. 187, 6804–6815 (2005)
Sennhauser, G., Bukowska, M. A., Briand, C. & Grütter, M. G. Crystal structure of the multidrug exporter MexB from Pseudomonas aeruginosa. J. Mol. Biol. 389, 134–145 (2009)
Su, C.-C. et al. Conformation of the AcrB multidrug efflux pump in mutants of the putative proton relay pathway. J. Bacteriol. 188, 7290–7296 (2006)
Higgins, M. K., Bokma, E., Koronakis, E., Hughes, C. & Koronakis, V. Structure of the periplasmic component of a bacterial drug efflux pump. Proc. Natl Acad. Sci. USA 101, 9994–9999 (2004)
Akama, H. et al. Crystal structure of the membrane fusion protein, MexA, of the multidrug transporter in Pseudomonas aeruginosa. J. Biol. Chem. 279, 25939–25942 (2004)
Mikolosko, J., Bobyk, K., Zgurskaya, H. I. & Ghosh, P. Conformational flexibility in the multidrug efflux system protein AcrA. Structure 14, 577–587 (2006)
Symmons, M., Bokma, E., Koronakis, E., Hughes, C. & Koronakis, V. The assembled structure of a complete tripartite bacterial multidrug efflux pump. Proc. Natl Acad. Sci. USA 106, 7173–7178 (2009)
Koronakis, V., Sharff, A., Koronakis, E., Luisi, B. & Hughes, C. Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature 405, 914–919 (2000)
Akama, H. et al. Crystal structure of the drug discharge outer membrane protein, OprM, of Pseudomonas aeruginosa. J. Biol. Chem. 279, 52816–52819 (2004)
Xue, Y. et al. Cu i recognition via cation-π and methionine interactions in CusF. Nature Chem. Biol. 4, 107–109 (2008)
Loftin, I. R., Franke, S., Blackburn, N. J. & McEvoy, M. M. Unusual Cu i/ Ag i coordination of Escherichia coli CusF as revealed by atomic resolution crystallography and X-ray absorption spectroscopy. Protein Sci. 16, 2287–2293 (2007)
Bagai, I., Liu, W., Rensing, C., Blackburn, N. & McEvoy, M. M. Substrate-linked conformational change in the periplasmic component of Cu i/ Ag i efflux system. J. Biol. Chem. 282, 35695–35702 (2007)
Gerken, H. & Misra, R. Genetic evidence for functional interactions between TolC and AcrA proteins of a major antibiotic efflux pump of Escherichia coli. Mol. Microbiol. 54, 620–631 (2004)
Bagai, I., Rensing, C., Blackburn, N. J. & McEvoy, M. M. Direct metal transfer between periplasmic proteins identifies a bacterial copper chaperone. Biochemistry 47, 11408–11414 (2008)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Terwilliger, T. C. Maximum-likelihood density modification using pattern recognition of structural motifs. Acta Crystallogr. D 57, 1755–1762 (2001)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nature Protocols 5, 725–738 (2010)
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005)
Feller, S. E. & MacKerell, A. D., Jr An improved empirical potential energy for molecular simulations of phospholipids. J. Phys. Chem. B 104, 7510–7515 (2000)
Acknowledgements
This work is supported by National Institutes of Health grants R01GM074027 (E.W.Y.), R01GM086431 (E.W.Y.), R01GM081680 (R.L.J.) and R01GM072014 (R.L.J.). This work is based on research conducted at the Northeastern Collaborative Access Team beamlines of the Advanced Photon Source, supported by award RR-15301 from the National Center for Research Resources at the National Institutes of Health. Use of the Advanced Photon Source is supported by the US Department of Energy, Office of Basic Energy Sciences, under contract DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
C.-C.S., F.L. and E.W.Y. designed the research. C.-C.S. and F.L. performed experiments. M.T.Z. and R.L.J. performed the docking. C.-C.S., F.L., K.R.R. and E.W.Y. performed the model building and refinement. C.-C.S., F.L., R.L.J. and E.W.Y. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References, Supplementary Table 1 and Supplementary Figures 1-4 with legends. (PDF 3220 kb)
Rights and permissions
About this article
Cite this article
Su, CC., Long, F., Zimmermann, M. et al. Crystal structure of the CusBA heavy-metal efflux complex of Escherichia coli. Nature 470, 558–562 (2011). https://doi.org/10.1038/nature09743
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09743
This article is cited by
-
Insights into the defensive mechanism of bioleaching microorganisms under extreme environmental copper stress
Reviews in Environmental Science and Bio/Technology (2023)
-
Unique underlying principles shaping copper homeostasis networks
JBIC Journal of Biological Inorganic Chemistry (2022)
-
Antibiotic export by MexB multidrug efflux transporter is allosterically controlled by a MexA-OprM chaperone-like complex
Nature Communications (2020)
-
Characterization of the copper resistance mechanism and bioremediation potential of an Acinetobacter calcoaceticus strain isolated from copper mine sludge
Environmental Science and Pollution Research (2020)
-
In situ structure and assembly of the multidrug efflux pump AcrAB-TolC
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.