Abstract
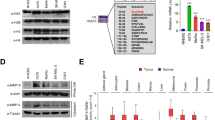
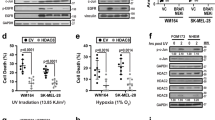
Cancer is a disease consisting of both genetic and epigenetic changes. Although increasing evidence demonstrates that tumour progression entails chromatin-mediated changes such as DNA methylation, the role of histone variants in cancer initiation and progression currently remains unclear. Histone variants replace conventional histones within the nucleosome and confer unique biological functions to chromatin1,2,3. Here we report that the histone variant macroH2A (mH2A) suppresses tumour progression of malignant melanoma. Loss of mH2A isoforms, histone variants generally associated with condensed chromatin and fine-tuning of developmental gene expression programs1,4,5,6, is positively correlated with increasing malignant phenotype of melanoma cells in culture and human tissue samples. Knockdown of mH2A isoforms in melanoma cells of low malignancy results in significantly increased proliferation and migration in vitro and growth and metastasis in vivo. Restored expression of mH2A isoforms rescues these malignant phenotypes in vitro and in vivo. We demonstrate that the tumour-promoting function of mH2A loss is mediated, at least in part, through direct transcriptional upregulation of CDK8. Suppression of CDK8, a colorectal cancer oncogene7,8, inhibits proliferation of melanoma cells, and knockdown of CDK8 in cells depleted of mH2A suppresses the proliferative advantage induced by mH2A loss. Moreover, a significant inverse correlation between mH2A and CDK8 expression levels exists in melanoma patient samples. Taken together, our results demonstrate that mH2A is a critical component of chromatin that suppresses the development of malignant melanoma, a highly intractable cutaneous neoplasm.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bernstein, E. & Hake, S. B. The nucleosome: a little variation goes a long way. Biochem. Cell Biol. 84, 505–517 (2006)
Sarma, K. & Reinberg, D. Histone variants meet their match. Nature Rev. Mol. Cell Biol. 6, 139–149 (2005)
Thambirajah, A. A., Li, A., Ishibashi, T. & Ausió, J. New developments in post-translational modifications and functions of histone H2A variants. Biochem. Cell Biol. 87, 7–17 (2009)
Changolkar, L. N. et al. Developmental changes in histone macroH2A1-mediated gene regulation. Mol. Cell. Biol. 27, 2758–2764 (2007)
Buschbeck, M. et al. The histone variant macroH2A is an epigenetic regulator of key developmental genes. Nature Struct. Mol. Biol. 16, 1074–1079 (2009)
Gamble, M. J., Frizzell, K. M., Yang, C., Krishnakumar, R. & Kraus, W. L. The histone variant macroH2A1 marks repressed autosomal chromatin, but protects a subset of its target genes from silencing. Genes Dev. 24, 21–32 (2010)
Firestein, R. et al. CDK8 is a colorectal cancer oncogene that regulates beta-catenin activity. Nature 455, 547–551 (2008)
Morris, E. J. et al. E2F1 represses beta-catenin transcription and is antagonized by both pRB and CDK8. Nature 455, 552–556 (2008)
Pehrson, J. R. & Fried, V. A. MacroH2A, a core histone containing a large nonhistone region. Science 257, 1398–1400 (1992)
Pehrson, J. R., Costanzi, C. & Dharia, C. Developmental and tissue expression patterns of histone macroH2A1 subtypes. J. Cell. Biochem. 65, 107–113 (1997)
Costanzi, C. & Pehrson, J. R. MacroH2A2, a new member of the MacroH2A core histone family. J. Biol. Chem. 276, 21776–21784 (2001)
Costanzi, C. & Pehrson, J. R. Histone macroH2A1 is concentrated in the inactive X chromosome of female mammals. Nature 393, 599–601 (1998)
Zhang, R. et al. Formation of MacroH2A-containing senescence-associated heterochromatin foci and senescence driven by ASF1a and HIRA. Dev. Cell 8, 19–30 (2005)
Bernstein, E. et al. A phosphorylated sub-population of the histone variant macroH2A1 is enriched during mitosis and excluded from the inactive X chromosome. Proc. Natl Acad. Sci. USA 105, 1533–1538 (2008)
Hua, S. et al. Genomic analysis of estrogen cascade reveals histone variant H2A.Z associated with breast cancer progression. Mol. Syst. Biol. 4, 188 (2008)
Sporn, J. C. et al. Histone macroH2A isoforms predict the risk of lung cancer recurrence. Oncogene 28, 3423–3428 (2009)
Hocker, T. L., Singh, M. K. & Tsao, H. Melanoma genetics and therapeutic approaches in the 21st century: moving from the benchside to the bedside. J. Invest. Dermatol. 128, 2575–2595 (2008)
Jemal, A. et al. Cancer Statistics, 2009. CA Cancer J. Clin. 59, 225–249 (2009)
Clark, W. H., Jr et al. Model predicting survival in stage I melanoma based on tumor progression. J. Natl Cancer Inst. 81, 1893–1904 (1989)
Fidler, I. J. Selection of successive tumor lines for metastasis. Nature 242, 148–149 (1973)
Satyamoorthy, K. et al. Melanoma cell lines from different stages of progression and their biological and molecular analyses. Melanoma Res. 7, S35–S42 (1997)
Plazas-Mayorca, M. D. et al. One-pot shotgun quantitative mass spectrometry characterization of histones. J. Proteome Res. 8, 5367–5374 (2009)
Yancovitz, M. et al. Detection of mutant BRAF alleles in the plasma of patients with metastatic melanoma. J. Mol. Diagn. 9, 178–183 (2007)
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002)
Qian, F., Vaux, D. L. & Weissman, I. L. Expression of the integrin α4β1 on melanoma cells can inhibit the invasive stage of metastasis formation. Cell 77, 335–347 (1994)
Ryu, B., Kim, D. S., Deluca, A. M. & Alani, R. M. Comprehensive expression profiling of tumor cell lines identifies molecular signatures of melanoma progression. PLoS ONE 2, e594 (2007)
Knuesel, M. T., Meyer, K. D., Bernecky, C. & Taatjes, D. J. The human CDK8 subcomplex is a molecular switch that controls Mediator coactivator function. Genes Dev. 23, 439–451 (2009)
Segura, M. F. et al. Aberrant miR-182 expression promotes melanoma metastasis by repressing FOXO3 and microphthalmia-associated transcription factor. Proc. Natl Acad. Sci. USA 106, 1814–1819 (2009)
Kumaki, Y., Oda, M. & Okano, M. QUMA: quantification tool for methylation analysis. Nucleic Acids Res. 36, W170–175 (2008)
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nature Protocols 4, 44–57 (2009)
Acknowledgements
We thank G. Hannon, S. Hake and M. Wirtz for reading this manuscript; J. Doucette for statistical support; the laboratories of S. Aaronson, J. Aguirre-Ghiso, D. Burstein and M. O’Connell for discussions and advice; M. Lebwohl, S. Mercer, J. Emer and G. Singer for dermatology and pathology support. We also thank N. Mall, L. Murray, S. Malu and S. Mungamuri for technical assistance; J. Pehrson, M. Narita, A. Aplin, H. Wei, A. Ting, S. Young Kim, M. Herlyn and J. Espinosa for reagents; T. Chu (Mount Sinai School of Medicine Microarray SRF) for data analysis; Q. Yu, the New York University Interdisciplinary Melanoma Cooperative Group and Mount Sinai Biorepository Cooperative for melanoma specimens. This work was supported by an American Skin Association Medical Student Grant to M.S.G., American Society for Mass Spectrometry Award, New Jersey Commission on Cancer Research Seed Grant, and National Science Foundation CBET-0941143 to B.A.G., National Institutes of Health CA109388 and the Sergei S. Zlinkoff Fund for Medical Education to D.P., New York University Cancer Institute Cancer Center Support Grant (5P30CA016087-27) and Marc Jacobs Campaign to I.O., NYSTEM IDEA C024291 and Harry L. Lloyd Charitable Trust to E.H., and an American Skin Association Research Scholar Award, Ellison Medical Foundation New Scholar Award, Tisch Cancer Institute Developmental Funds and NCI R21CA150117 to E.B.
Author information
Authors and Affiliations
Contributions
A.K., M.S.G., K.R. and E.B. conceived ideas and experimental design. A.K. generated shRNA and overexpression cell lines and performed all in vitro and in vivo studies, DNA methylation analysis, microarray studies, qPCR and ChIP. M.S.G. performed immunoblots, IHC and statistical analysis. L.K.C. performed IHC, immunoblots and in vivo studies, and K.R. performed MNase studies. C.V. performed ChIP, and P.O.E. and C.I.V. scored and evaluated all human and mouse tissues. S.M., M.F.S. and E.H. designed and performed mouse studies, G.L. and B.A.G. performed and analysed quantitative mass spectrometry studies, and D.P. and I.O. contributed critical reagents. E.B. wrote the manuscript with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-23 with legends and Supplementary Tables 1-4 and additional references. (PDF 5693 kb)
Supplementary Table 5
This excel file is appended with probe changes and intersections in microarray experiment #1. (XLS 228 kb)
Supplementary Table 6
This excel file is appended with probe changes and intersections in microarray experiment #2. (XLS 121 kb)
Rights and permissions
About this article
Cite this article
Kapoor, A., Goldberg, M., Cumberland, L. et al. The histone variant macroH2A suppresses melanoma progression through regulation of CDK8. Nature 468, 1105–1109 (2010). https://doi.org/10.1038/nature09590
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09590
This article is cited by
-
Hit discovery of potential CDK8 inhibitors and analysis of amino acid mutations for cancer therapy through computer-aided drug discovery
BMC Chemistry (2024)
-
Epigenetic control of cancer inflammation
Nature Cell Biology (2023)
-
MacroH2A1.1 as a crossroad between epigenetics, inflammation and metabolism of mesenchymal stromal cells in myelodysplastic syndromes
Cell Death & Disease (2023)
-
Cyclins and cyclin-dependent kinases: from biology to tumorigenesis and therapeutic opportunities
Journal of Cancer Research and Clinical Oncology (2023)
-
Current understanding of epigenetics role in melanoma treatment and resistance
Cancer Cell International (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.