Abstract
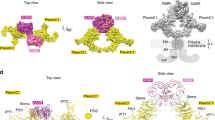
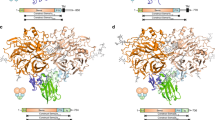
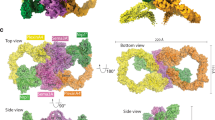
Semaphorins and their receptor plexins constitute a pleiotropic cell-signalling system that is used in a wide variety of biological processes, and both protein families have been implicated in numerous human diseases1,2,3,4. The binding of soluble or membrane-anchored semaphorins to the membrane-distal region of the plexin ectodomain activates plexin’s intrinsic GTPase-activating protein (GAP) at the cytoplasmic region, ultimately modulating cellular adhesion behaviour5. However, the structural mechanism underlying the receptor activation remains largely unknown. Here we report the crystal structures of the semaphorin 6A (Sema6A) receptor-binding fragment and the plexin A2 (PlxnA2) ligand-binding fragment in both their pre-signalling (that is, before binding) and signalling (after complex formation) states. Before binding, the Sema6A ectodomain was in the expected ‘face-to-face’ homodimer arrangement, similar to that adopted by Sema3A and Sema4D, whereas PlxnA2 was in an unexpected ‘head-on’ homodimer arrangement. In contrast, the structure of the Sema6A–PlxnA2 signalling complex revealed a 2:2 heterotetramer in which the two PlxnA2 monomers dissociated from one another and docked onto the top face of the Sema6A homodimer using the same interface as the head-on homodimer, indicating that plexins undergo ‘partner exchange’. Cell-based activity measurements using mutant ligands/receptors confirmed that the Sema6A face-to-face dimer arrangement is physiologically relevant and is maintained throughout signalling events. Thus, homodimer-to-heterodimer transitions of cell-surface plexin that result in a specific orientation of its molecular axis relative to the membrane may constitute the structural mechanism by which the ligand-binding ‘signal’ is transmitted to the cytoplasmic region, inducing GAP domain rearrangements and activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tamagnone, L. & Comoglio, P. M. To move or not to move? Semaphorin signalling in cell migration. EMBO Rep. 5, 356–361 (2004)
Zhou, Y., Gunput, R. A. & Pasterkamp, R. J. Semaphorin signaling: progress made and promises ahead. Trends Biochem. Sci. 33, 161–170 (2008)
Suzuki, K., Kumanogoh, A. & Kikutani, H. Semaphorins and their receptors in immune cell interactions. Nature Immunol. 9, 17–23 (2008)
Neufeld, G. & Kessler, O. The semaphorins: versatile regulators of tumour progression and tumour angiogenesis. Nature Rev. Cancer 8, 632–645 (2008)
Pasterkamp, R. J. R-Ras fills another GAP in semaphorin signalling. Trends Cell Biol. 15, 61–64 (2005)
Antipenko, A. et al. Structure of the semaphorin-3A receptor binding module. Neuron 39, 589–598 (2003)
Love, C. A. et al. The ligand-binding face of the semaphorins revealed by the high-resolution crystal structure of SEMA4D. Nature Struct. Biol. 10, 843–848 (2003)
Stamos, J., Lazarus, R. A., Yao, X., Kirchhofer, D. & Wiesmann, C. Crystal structure of the HGF β-chain in complex with the Sema domain of the Met receptor. EMBO J. 23, 2325–2335 (2004)
Suto, F. et al. Plexin-A4 mediates axon-repulsive activities of both secreted and transmembrane semaphorins and plays roles in nerve fiber guidance. J. Neurosci. 25, 3628–3637 (2005)
Renaud, J. et al. Plexin-A2 and its ligand, Sema6A, control nucleus-centrosome coupling in migrating granule cells. Nature Neurosci. 11, 440–449 (2008)
Merte, J. et al. A forward genetic screen in mice identifies Sema3A(K108N), which binds to neuropilin-1 but cannot signal. J. Neurosci. 30, 5767–5775 (2010)
Xu, X. M. et al. The transmembrane protein semaphorin 6A repels embryonic sympathetic axons. J. Neurosci. 20, 2638–2648 (2000)
Le Du, M. H., Stigbrand, T., Taussig, M. J., Menez, A. & Stura, E. A. Crystal structure of alkaline phosphatase from human placenta at 1.8 Å resolution. Implication for a substrate specificity. J. Biol. Chem. 276, 9158–9165 (2001)
Tong, Y. et al. Binding of Rac1, Rnd1, and RhoD to a novel Rho GTPase interaction motif destabilizes dimerization of the plexin-B1 effector domain. J. Biol. Chem. 282, 37215–37224 (2007)
Takahashi, T. & Strittmatter, S. M. Plexina1 autoinhibition by the plexin sema domain. Neuron 29, 429–439 (2001)
Chaudhry, C., Weston, M. C., Schuck, P., Rosenmund, C. & Mayer, M. L. Stability of ligand-binding domain dimer assembly controls kainate receptor desensitization. EMBO J. 28, 1518–1530 (2009)
Niemann, H. H. et al. Structure of the human receptor tyrosine kinase met in complex with the Listeria invasion protein InlB. Cell 130, 235–246 (2007)
Luo, B. H. & Springer, T. A. Integrin structures and conformational signaling. Curr. Opin. Cell Biol. 18, 579–586 (2006)
Oinuma, I., Ishikawa, Y., Katoh, H. & Negishi, M. The Semaphorin 4D receptor Plexin-B1 is a GTPase activating protein for R-Ras. Science 305, 862–865 (2004)
He, H., Yang, T., Terman, J. R. & Zhang, X. Crystal structure of the plexin A3 intracellular region reveals an autoinhibited conformation through active site sequestration. Proc. Natl Acad. Sci. USA 106, 15610–15615 (2009)
Tong, Y. et al. Structure and function of the intracellular region of the plexin-B1 transmembrane receptor. J. Biol. Chem. 284, 35962–35972 (2009)
Liu, H. et al. Structural basis of semaphorin-plexin recognition and viral mimicry from Sema7A and A39R complexes with plexinC1. Cell 142, 749–761 (2010)
Stanley, P. Chinese hamster ovary cell mutants with multiple glycosylation defects for production of glycoproteins with minimal carbohydrate heterogeneity. Mol. Cell. Biol. 9, 377–383 (1989)
Tabata, S. et al. A rapid screening method for cell lines producing singly-tagged recombinant proteins using the “TARGET tag” system. J. Proteomics 73, 1777–1785 (2010)
Toyofuku, T. et al. Repulsive and attractive semaphorins cooperate to direct the navigation of cardiac neural crest cells. Dev. Biol. 321, 251–262 (2008)
Goshima, Y. et al. A novel action of collapsin: collapsin-1 increases antero- and retrograde axoplasmic transport independently of growth cone collapse. J. Neurobiol. 33, 316–328 (1997)
Leahy, D. J., Dann, C. E., Longo, P., Perman, B. & Ramyar, K. X. A mammalian expression vector for expression and purification of secreted proteins for structural studies. Protein Expr. Purif. 20, 500–506 (2000)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Perrakis, A., Harkiolaki, M., Wilson, K. S. & Lamzin, V. S. ARP/wARP and molecular replacement. Acta Crystallogr. D 57, 1445–1450 (2001)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Lee, B. & Richards, F. M. The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55, 379–400 (1971)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000)
Schuck, P. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124 (2003)
Acknowledgements
We would like to thank Y. Yamada, N. Matsugaki and N. Igarashi of Photon Factory and Y. Kawano and N. Shimizu of SPring-8 BL-41XU for their help with the X-ray data collection; A. Rowe for discussions on the sedimentation equilibrium data analysis; C. Wu for performing the Sema6A–AP binding assay; K. Tamura-Kawakami and M. Nampo for their technical support; and M. Nakano for preparation of the manuscript. This work was supported in part by a ‘Target Proteins Research Program (TPRP)’ grant from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT).
Author information
Authors and Affiliations
Contributions
T.N. and J.T. conceived the project. No.Y. and E.M. expressed, purified and crystallized the proteins. Y.M., Na.Y. and T.T. performed cell biological assays. M.N. and S.U. performed analytical ultracentrifugation experiments. T.N. and No.Y. collected X-ray diffraction data. T.N. and J.T. determined and analysed the structures. T.N., S.U., Y.G., A.K. and J.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Results, additional references, Supplementary Tables 1-3 and Supplementary Figures 1-9 with legends. (PDF 1334 kb)
Rights and permissions
About this article
Cite this article
Nogi, T., Yasui, N., Mihara, E. et al. Structural basis for semaphorin signalling through the plexin receptor. Nature 467, 1123–1127 (2010). https://doi.org/10.1038/nature09473
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09473
This article is cited by
-
Cryo-EM structure of the PlexinC1/A39R complex reveals inter-domain interactions critical for ligand-induced activation
Nature Communications (2020)
-
Origin and evolution of plexins, semaphorins, and Met receptor tyrosine kinases
Scientific Reports (2019)
-
Diversity of oligomerization in Drosophila semaphorins suggests a mechanism of functional fine-tuning
Nature Communications (2019)
-
The extracellular SEMA domain attenuates intracellular apoptotic signaling of semaphorin 6A in lung cancer cells
Oncogenesis (2018)
-
Transcriptional changes in mesenteric and subcutaneous adipose tissue from Holstein cows in response to plane of dietary energy
Journal of Animal Science and Biotechnology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.