Abstract
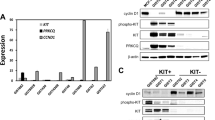
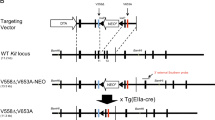
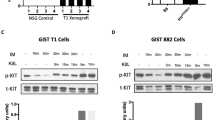
Gastrointestinal stromal tumour (GIST) is the most common human sarcoma and is primarily defined by activating mutations in the KIT or PDGFRA receptor tyrosine kinases1,2. KIT is highly expressed in interstitial cells of Cajal (ICCs)—the presumed cell of origin for GIST—as well as in haematopoietic stem cells, melanocytes, mast cells and germ cells2,3. Yet, families harbouring germline activating KIT mutations and mice with knock-in Kit mutations almost exclusively develop ICC hyperplasia and GIST4,5,6,7, suggesting that the cellular context is important for KIT to mediate oncogenesis. Here we show that the ETS family member ETV1 is highly expressed in the subtypes of ICCs sensitive to oncogenic KIT mediated transformation8, and is required for their development. In addition, ETV1 is universally highly expressed in GISTs and is required for growth of imatinib-sensitive and resistant GIST cell lines. Transcriptome profiling and global analyses of ETV1-binding sites suggest that ETV1 is a master regulator of an ICC-GIST-specific transcription network mainly through enhancer binding. The ETV1 transcriptional program is further regulated by activated KIT, which prolongs ETV1 protein stability and cooperates with ETV1 to promote tumorigenesis. We propose that GIST arises from ICCs with high levels of endogenous ETV1 expression that, when coupled with an activating KIT mutation, drives an oncogenic ETS transcriptional program. This differs from other ETS-dependent tumours such as prostate cancer, melanoma and Ewing sarcoma where genomic translocation or amplification drives aberrant ETS expression9,10,11. It also represents a novel mechanism of oncogenic transcription factor activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Heinrich, M. C. et al. PDGFRA activating mutations in gastrointestinal stromal tumors. Science 299, 708–710 (2003)
Hirota, S. et al. Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 279, 577–580 (1998)
Kindblom, L. G., Remotti, H. E., Aldenborg, F. & Meis-Kindblom, J. M. Gastrointestinal pacemaker cell tumor (GIPACT): gastrointestinal stromal tumors show phenotypic characteristics of the interstitial cells of Cajal. Am. J. Pathol. 152, 1259–1269 (1998)
Antonescu, C. R. Gastrointestinal stromal tumor (GIST) pathogenesis, familial GIST, and animal models. Semin. Diagn. Pathol. 23, 63–69 (2006)
Nakai, N. et al. A mouse model of a human multiple GIST family with KIT-Asp820Tyr mutation generated by a knock-in strategy. J. Pathol. 214, 302–311 (2008)
Rubin, B. P. et al. A knock-in mouse model of gastrointestinal stromal tumor harboring kit K641E. Cancer Res 65, 6631–6639 (2005)
Sommer, G. et al. Gastrointestinal stromal tumors in a mouse model by targeted mutation of the Kit receptor tyrosine kinase. Proc. Natl Acad. Sci. USA 100, 6706–6711 (2003)
Kwon, J. G. et al. Changes in the structure and function of ICC networks in ICC hyperplasia and gastrointestinal stromal tumors. Gastroenterology 136, 630–639 (2009)
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005)
Mertens, F. et al. Translocation-related sarcomas. Semin. Oncol. 36, 312–323 (2009)
Jane-Valbuena, J. et al. An oncogenic role for ETV1 in melanoma. Cancer Res 70, 2075–2084 (2010)
Nielsen, T. O. et al. Molecular characterisation of soft tissue tumours: a gene expression study. Lancet 359, 1301–1307 (2002)
Segal, N. H. et al. Classification and subtype prediction of adult soft tissue sarcoma by functional genomics. Am. J. Pathol. 163, 691–700 (2003)
Tomlins, S. A. et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448, 595–599 (2007)
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010)
Ward, S. M. & Sanders, K. M. Physiology and pathophysiology of the interstitial cell of Cajal: from bench to bedside. I. Functional development and plasticity of interstitial cells of Cajal networks. Am. J. Physiol. Gastrointest. Liver Physiol. 281, G602–G611 (2001)
Chen, H. et al. Differential gene expression in functional classes of interstitial cells of Cajal in murine small intestine. Physiol. Genomics 31, 492–509 (2007)
Arber, S., Ladle, D. R., Lin, J. H., Frank, E. & Jessell, T. M. ETS gene Er81 controls the formation of functional connections between group Ia sensory afferents and motor neurons. Cell 101, 485–498 (2000)
Gomez-Pinilla, P. J. et al. Ano1 is a selective marker of interstitial cells of Cajal in the human and mouse gastrointestinal tract. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G1370–G1381 (2009)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009)
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009)
Miettinen, M. & Lasota, J. Gastrointestinal stromal tumors: review on morphology, molecular pathology, prognosis, and differential diagnosis. Arch. Pathol. Lab. Med. 130, 1466–1478 (2006)
Andersen, R. J. et al. Regression of castrate-recurrent prostate cancer by a small-molecule inhibitor of the amino-terminus domain of the androgen receptor. Cancer Cell 17, 535–546 (2010)
Moellering, R. E. et al. Direct inhibition of the NOTCH transcription factor complex. Nature 462, 182–188 (2009)
Bauer, S., Yu, L. K., Demetri, G. D. & Fletcher, J. A. Heat shock protein 90 inhibition in imatinib-resistant gastrointestinal stromal tumor. Cancer Res. 66, 9153–9161 (2006)
Yamaguchi, U. et al. Distinct gene expression-defined classes of gastrointestinal stromal tumor. J. Clin. Oncol. 26, 4100–4108 (2008)
Ostrowski, J. et al. Functional features of gene expression profiles differentiating gastrointestinal stromal tumours according to KIT mutations and expression. BMC Cancer 9, 413 (2009)
de Jonge, H. J. et al. Evidence based selection of housekeeping genes. PLoS ONE 2, e898 (2007)
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005)
Goldberg, A. D. et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140, 678–691 (2010)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Bailey, T. L., Williams, N., Misleh, C. & Li, W. W. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006)
Acknowledgements
This work was supported in part by the National Cancer Institute (K08CA140946, to Y.C.), (5F32CA130372, to P.C.), (CA47179, to C.R.A. and R.G.M.), (CA148260, to R.G.M.), US National Institute of Mental Health (R21MH087840, to D.Z.), National Cancer Institute–American Society of Clinical Oncology Cancer Foundation Clinical Investigator Team Leadership Supplemental Award (to R.G.M.), American Society of Clinical Oncology Young Investigator Award (to P.C.), the Doris Duke (to C.L.S.), Charles H Revson (to Y.C.) and Charles A. Dana (to Y.C.) foundations, The Rockefeller University Fund (to C.D.A.), American Cancer Society Mentored Research Scholar Grant CCE-106841 (to C.R.A.), P01CA47179 (to C.R.A. and R.G.M.), the Life Raft Group (to C.R.A.), the GIST Cancer Research Fund (to C.R.A.), the Shuman Family Fund for GIST Research (to P.C., C.R.A. and R.G.M.), Cycle for Survival (to R.G.M.), Startup Funds from the Albert Einstein College of Medicine (to D.Z.), National Institutes of Health (GM40922, to C.D.A.) and the Starr Cancer Consortium (to P.C., Y.C., C.L.S. and C.D.A.). We thank the International Genomics Consortium for generating ExpO data. We thank G. Wang, P. Iaquinta, and H. Hieronymus for discussions, and especially T. M. Jessell and J. N. Betley for providing and breeding Etv1−/− mice.
Author information
Authors and Affiliations
Contributions
Author contributions P.C., Y.C., C.D.A. and C.L.S. designed the experiments. R.G.M. and C.R.A. provided advice about experimental design. P.C. and Y.C. performed most of the experiments, including data mining, data analysis, tissue culture experiments, tissue processing, immunofluorescence microscopy, colony formation assays and ChIP-Seq experiments. J.W. and T.S. performed xenograft and some qRT-PCR experiments. L.Z. and C.R.A. provided human tumour samples and performed fluorescence in situ hybridization and immunohistochemistry on them. S.D. performed the Solexa sequencing and genomic alignment, and X.G. and D.Z. analysed ChIP-Seq data. J.A.F. provided key experimental reagents. P.C., Y.C., C.D.A. and C.L.S. wrote the manuscript. All authors discussed results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-17 with legends. (PDF 14059 kb)
Supplementary Table 1
This table shows a complete gene lists of three GIST-signature gene sets and ICC-MY and ICC-DMP-signature gene sets. Probe ID, gene symbol, average Z-score of expression, and Z-score difference are shown. (XLS 81 kb)
Supplementary Table 2
This table shows gene sets with FDR < 0.25 for overrepresented among downregulated by ETV1 knockdown by GSEA - gene set name, size, enrichment score, normalized enrichment score, p-value, and FDR are shown. (XLS 47 kb)
Supplementary Movie 1
Whole Mount Reconstruction of Kit IF in Etv1+/+ Large Intestine - the movie focuses from the ICC-MY layer through the ICC-IM layer, ending with the ICC-SMP layer. Image is 176x132 μm. (MOV 284 kb)
Supplementary Movie 2
Whole Mount Reconstruction of Kit IF in Etv1-/- Large Intestine ? the movie focuses from the ICC-MY layer through the ICC-IM layer, ending with the ICC-SMP layer. Image is 176x132 μm. (MOV 81 kb)
Supplementary Movie 3
Whole Mount Reconstruction of Kit IF in Etv1+/+ Small Intestine ? the movie focuses from the ICC-MY layer into the ICC-DMP layer. Image is 59x44 μm. (MOV 81 kb)
Supplementary Movie 4
Whole Mount Reconstruction of Kit IF in Etv1-/- Small Intestine - the movie focuses from the ICC-MY layer into the ICC-DMP layer. Image is 59x44 μm. (MOV 25 kb)
Supplementary Movie 5
Whole Mount Reconstruction of Kit IF in Etv1+/+ Cecum - the movie focuses from the ICC-MY layer into the ICC-IM layer. Image is 176x132 μm. (MOV 124 kb)
Supplementary Movie 6
Whole Mount Reconstruction of Kit IF in Etv1-/- Cecum. Movie focuses from the ICC-MY layer into the ICC-IM layer. Image is 176x132 μm. (MOV 26 kb)
Supplementary Movie 7
Whole Mount Reconstruction of Kit IF in Etv1+/+ Gastric Fundus. Movie focuses from the outer longitudinal intramuscular ICC layer into the inner circular intramuscular ICC layer. Image is 176x132 μm. (MOV 104 kb)
Supplementary Movie 8
Whole Mount Reconstruction of Kit IF in Etv1-/- Gastric Fundus. Movie focuses from the outer longitudinal intramuscular ICC layer into the inner circular intramuscular ICC layer. Image is 176x132 μm. (MOV 45 kb)
Rights and permissions
About this article
Cite this article
Chi, P., Chen, Y., Zhang, L. et al. ETV1 is a lineage survival factor that cooperates with KIT in gastrointestinal stromal tumours. Nature 467, 849–853 (2010). https://doi.org/10.1038/nature09409
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09409
This article is cited by
-
KIT mutations and expression: current knowledge and new insights for overcoming IM resistance in GIST
Cell Communication and Signaling (2024)
-
Jejunal gastrointestinal stromal tumor that developed in a patient with neurofibromatosis type 1: a case report
Diagnostic Pathology (2023)
-
The microphthalmia-associated transcription factor is involved in gastrointestinal stromal tumor growth
Cancer Gene Therapy (2023)
-
Deubiquitylation of Rab35 by USP32 promotes the transmission of imatinib resistance by enhancing exosome secretion in gastrointestinal stromal tumours
Oncogene (2023)
-
HGF-mediated elevation of ETV1 facilitates hepatocellular carcinoma metastasis through upregulating PTK2 and c-MET
Journal of Experimental & Clinical Cancer Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.