Abstract
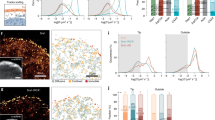
Crawling locomotion of eukaryotic cells is achieved by a process dependent on the actin cytoskeleton1: protrusion of the leading edge requires assembly of a network of actin filaments2, which must be disassembled at the cell rear for sustained motility. Although ADF/cofilin proteins have been shown to contribute to actin disassembly3, it is not clear how activity of these locally acting proteins could be coordinated over the distance scale of the whole cell. Here we show that non-muscle myosin II has a direct role in actin network disassembly in crawling cells. In fish keratocytes undergoing motility, myosin II is concentrated in regions at the rear with high rates of network disassembly. Activation of myosin II by ATP in detergent-extracted cytoskeletons results in rear-localized disassembly of the actin network. Inhibition of myosin II activity and stabilization of actin filaments synergistically impede cell motility, suggesting the existence of two disassembly pathways, one of which requires myosin II activity. Our results establish the importance of myosin II as an enzyme for actin network disassembly; we propose that gradual formation and reorganization of an actomyosin network provides an intrinsic destruction timer, enabling long-range coordination of actin network treadmilling in motile cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Abercrombie, M. The Croonian lecture, 1978: the crawling movement of metazoan cells. Proc. R. Soc. Lond. B 207, 129–147 (1980)
Theriot, J. A. & Mitchison, T. J. Actin microfilament dynamics in locomoting cells. Nature 352, 126–131 (1991)
Pollard, T. D. & Borisy, G. G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003)
Euteneuer, U. & Schliwa, M. Persistent, directional motility of cells and cytoplasmic fragments in the absence of microtubules. Nature 310, 58–61 (1984)
Small, J. V., Herzog, M., Häner, M. & Abei, U. Visualization of actin filaments in keratocyte lamellipodia: negative staining compared with freeze-drying. J. Struct. Biol. 113, 135–141 (1994)
Svitkina, T. M., Verkhovsky, A. B., McQuade, K. M. & Borisy, G. G. Analysis of the actin-myosin II system in fish epidermal keratocytes: mechanism of cell body translocation. J. Cell Biol. 139, 397–415 (1997)
Ji, L. & Danuser, G. Tracking quasi-stationary flow of weak fluorescent signals by adaptive multi-frame correlation. J. Microsc. 220, 150–167 (2005)
Wilson, C. A. & Theriot, J. A. A correlation-based approach to calculate rotation and translation of moving cells. IEEE Trans. Image Process. 15, 1939–1951 (2006)
Vallotton, P., Gupton, S. L., Waterman-Storer, C. M. & Danuser, G. Simultaneous mapping of filamentous actin flow and turnover in migrating cells by quantitative fluorescent speckle microscopy. Proc. Natl Acad. Sci. USA 101, 9660–9665 (2004)
Guha, M., Zhou, M. & Wang, Y.-L. Cortical actin turnover during cytokinesis requires myosin II. Curr. Biol. 15, 732–736 (2005)
Murthy, K. & Wadsworth, P. Myosin-II-dependent localization and dynamics of F-actin during cytokinesis. Curr. Biol. 15, 724–731 (2005)
Sai, X. & Ladher, R. K. FGF signaling regulates cytoskeletal remodeling during epithelial morphogenesis. Curr. Biol. 18, 976–981 (2008)
Medeiros, N. A., Burnette, D. T. & Forscher, P. Myosin II functions in actin-bundle turnover in neuronal growth cones. Nature Cell Biol. 8, 215–226 (2006)
Haviv, L., Gillo, D., Backouche, F. & Bernheim-Groswasser, A. A cytoskeletal demolition worker: myosin II acts as an actin depolymerization agent. J. Mol. Biol. 375, 325–330 (2008)
Straight, A. F. et al. Dissecting temporal and spatial control of cytokinesis with a myosin II inhibitor. Science 299, 1743–1747 (2003)
Bubb, M. R., Senderowicz, A. M., Sausville, E. A., Duncan, K. L. & Korn, E. D. Jasplakinolide, a cytotoxic natural product, induces actin polymerization and competitively inhibits the binding of phalloidin to F-actin. J. Biol. Chem. 269, 14869–14871 (1994)
Bretscher, A. & Weber, K. Villin is a major protein of the microvillus cytoskeleton which binds both G and F actin in a calcium-dependent manner. Cell 20, 839–847 (1980)
Glenney, J. R., Kaulfus, P. & Weber, K. F actin assembly modulated by villin: Ca++-dependent nucleation and capping of the barbed end. Cell 24, 471–480 (1981)
Ponti, A., Machacek, M., Gupton, S. L., Waterman-Storer, C. M. & Danuser, G. Two distinct actin networks drive the protrusion of migrating cells. Science 305, 1782–1786 (2004)
Kron, S. J. & Spudich, J. A. Fluorescent actin filaments move on myosin fixed to a glass surface. Proc. Natl Acad. Sci. USA 83, 6272–6276 (1986)
Kovács, M., Thirumurugan, K., Knight, P. J. & Sellers, J. R. Load-dependent mechanism of nonmuscle myosin 2. Proc. Natl Acad. Sci. USA 104, 9994–9999 (2007)
Yumura, S., Mori, H. & Fukui, Y. Localization of actin and myosin for the study of ameboid movement in Dictyostelium using improved immunofluorescence. J. Cell Biol. 99, 894–899 (1984)
Keller, H. U. & Niggli, V. Colchicine-induced stimulation of PMN motility related to cytoskeletal changes in actin, α-actinin, and myosin. Cell Motil. Cytoskeleton 25, 10–18 (1993)
Vicente-Manzanares, M., Zareno, J., Whitmore, L., Choi, C. K. & Horwitz, A. F. Regulation of protrusion, adhesion dynamics, and polarity by myosins IIA and IIB in migrating cells. J. Cell Biol. 176, 573–580 (2007)
Yam, P. T. et al. Actin–myosin network reorganization breaks symmetry at the cell rear to spontaneously initiate polarized cell motility. J. Cell Biol. 178, 1207–1221 (2007)
Lacayo, C. I. et al. Emergence of large-scale cell morphology and movement from local actin filament growth dynamics. PLoS Biol. 5, e233 (2007)
Sakamoto, T., Limouze, J., Combs, C. A., Straight, A. F. & Sellers, J. R. Blebbistatin, a myosin II inhibitor, is photoinactivated by blue light. Biochemistry 44, 584–588 (2005)
Kolega, J. Phototoxicity and photoinactivation of blebbistatin in UV and visible light. Biochem. Biophys. Res. Commun. 320, 1020–1025 (2004)
Doyle, A. D. & Lee, J. Cyclic changes in keratocyte speed and traction stress arise from Ca2+-dependent regulation of cell adhesiveness. J. Cell Sci. 118, 369–379 (2005)
Acknowledgements
We thank M. J. Footer for the gift of purified GST–villin, A. Mogilner and S. Sivaramakrishnan for discussions, and Z. Pincus, N. Dye and T. Y.-C. Tsai for reading the manuscript. C.A.W. and J.A.T. were supported by National Institutes of Health grant R01AI067712 (J.A.T.). M.A.T. and E.L.B. were supported by National Institutes of Health grant T32GM007276. G.M.A. was supported by the Stanford Medical Scientist Training Program. P.T.Y. was supported by a Howard Hughes Medical Institute Predoctoral Fellowship, Stanford Graduate Fellowship and a Skye International Foundation Scholarship. K.T.A. was supported by a National Science Foundation Graduate Research Fellowship. K.K. was a Damon Runyon Fellow supported by the Damon Runyon Cancer Research Foundation (DRG-#1854-05). K.T.A., L.J. and G.D. were supported by National Institutes of Health grants U54GM64346 and U01GM67230 (G.D.). J.A.T. was supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.A.W., M.A.T. and J.A.T. conceived and designed the experiments. C.A.W. and P.T.Y. performed FSM on untreated motile keratocytes. C.A.W. performed the pharmacological manipulation experiments, FSM observation and the analysis. L.J. and G.D. developed the flow tracking algorithm specific to the needs of this analysis. C.A.W. and P.T.Y. developed methods and software to integrate the flow tracking algorithm with these experiments and analysis. K.T.A., C.A.W. and G.D. developed algorithms for the F-actin turnover analysis. E.L.B. imaged myosin II localization. G.M.A., K.K. and E.L.B. collected the cell speed data and observed fixed cells under the different treatments; G.M.A. and C.A.W. analysed these data. M.A.T. performed experiments on detergent-extracted cytoskeletons and analysed the results. M.A.T., C.A.W. and J.A.T. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3, Supplementary Figures 1-6 with legends, captions for Supplementary Movies 1-4 and References. (PDF 5572 kb)
Supplementary Movie 1
This movie shows that myosin II inhibition alters actin network flow (see Supplementary Information file for full caption). (MOV 20825 kb)
Supplementary Movie 2
This movie shows that inward traction force generation requires myosin II activity (see Supplementary Information file for full caption). (MOV 8661 kb)
Supplementary Movie 3
This movie shows that jasplakinolide halts actin dynamics of cells in which myosin II is inhibited (see Supplementary Information file for full caption). (MOV 15729 kb)
Supplementary Movie 4
This movie shows that actin network disassembly in the rear of detergent-extracted keratocyte cytoskeletons is ATP-dependent and blebbistatin-sensitive (see Supplementary Information file for full caption). (MOV 424 kb)
Rights and permissions
About this article
Cite this article
Wilson, C., Tsuchida, M., Allen, G. et al. Myosin II contributes to cell-scale actin network treadmilling through network disassembly. Nature 465, 373–377 (2010). https://doi.org/10.1038/nature08994
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08994
This article is cited by
-
Role of cell rearrangement and related signaling pathways in the dynamic process of tip cell selection
Cell Communication and Signaling (2024)
-
Cell state dependent effects of Bmal1 on melanoma immunity and tumorigenicity
Nature Communications (2024)
-
Salmonella manipulates macrophage migration via SteC-mediated myosin light chain activation to penetrate the gut-vascular barrier
The EMBO Journal (2024)
-
Switch of cell migration modes orchestrated by changes of three-dimensional lamellipodium structure and intracellular diffusion
Nature Communications (2023)
-
A coarse-grained approach to model the dynamics of the actomyosin cortex
BMC Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.