Abstract
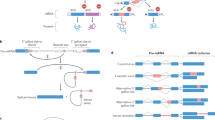
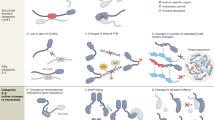
The collection of components required to carry out the intricate processes involved in generating and maintaining a living, breathing and, sometimes, thinking organism is staggeringly complex. Where do all of the parts come from? Early estimates stated that about 100,000 genes would be required to make up a mammal; however, the actual number is less than one-quarter of that, barely four times the number of genes in budding yeast. It is now clear that the 'missing' information is in large part provided by alternative splicing, the process by which multiple different functional messenger RNAs, and therefore proteins, can be synthesized from a single gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Alt, F. W. et al. Synthesis of secreted and membrane-bound immunoglobulin μ heavy chains is directed by mRNAs that differ at their 3′ ends. Cell 20, 293–301 (1980).
Early, P. et al. Two mRNAs can be produced from a single immunoglobulin μ gene by alternative RNA processing pathways. Cell 20, 313–319 (1980).
Rosenfeld, M. G. et al. Calcitonin mRNA polymorphism: peptide switching associated with alternative RNA splicing events. Proc. Natl Acad. Sci. USA 79, 1717–1721 (1982).
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nature Genet. 40, 1413–1415 (2008).
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008). References 4 and 5 provide detailed views of the human transcriptome as determined by using deep-sequencing data. The authors conclude that the pre-mRNAs from all multi-exon genes are alternatively spliced.
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000).
Navaratnam, D. S., Bell, T. J., Tu, T. D., Cohen, E. L. & Oberholtzer, J. C. Differential distribution of Ca2+-activated K+ channel splice variants among hair cells along the tonotopic axis of the chick cochlea. Neuron 19, 1077–1085 (1997).
Rosenblatt, K. P., Sun, Z. P., Heller, S. & Hudspeth, A. J. Distribution of Ca2+-activated K+ channel isoforms along the tonotopic gradient of the chicken's cochlea. Neuron 19, 1061–1075 (1997).
Dorn, R., Reuter, G. & Loewendorf, A. Transgene analysis proves mRNA trans-splicing at the complex mod(mdg4) locus in Drosophila. Proc. Natl Acad. Sci. USA 98, 9724–9729 (2001).
Labrador, M. et al. Protein encoding by both DNA strands. Nature 409, 1000 (2001). References 9 and 10 show that trans-splicing of two separate pre-mRNAs can generate a single protein-coding mRNA.
Sánchez, L. Sex-determining mechanisms in insects. Int. J. Dev. Biol. 52, 837–856 (2008).
Makeyev, E. V., Zhang, J., Carrasco, M. A. & Maniatis, T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 27, 435–448 (2007).
Boutz, P. L. et al. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 21, 1636–1652 (2007).
Xie, J. & Black, D. L. A CaMK IV responsive RNA element mediates depolarization-induced alternative splicing of ion channels. Nature 410, 936–939 (2001).
Lynch, K. W. Regulation of alternative splicing by signal transduction pathways. Adv. Exp. Med. Biol. 623, 161–174 (2007).
Shin, C. & Manley, J. L. Cell signalling and the control of pre-mRNA splicing. Nature Rev. Mol. Cell Biol. 5, 727–738 (2004).
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nature Rev. Mol. Cell Biol. 6, 386–398 (2005).
Gabut, M., Chaudhry, S. & Blencowe, B. J. The splicing regulatory machinery. Cell 133, 192 (2008).
Buckanovich, R. J., Posner, J. B. & Darnell, R. B. Nova, the paraneoplastic Ri antigen, is homologous to an RNA-binding protein and is specifically expressed in the developing motor system. Neuron 11, 657–672 (1993).
Polydorides, A. D., Okano, H. J., Yang, Y. Y., Stefani, G. & Darnell, R. B. A brain-enriched polypyrimidine tract-binding protein antagonizes the ability of Nova to regulate neuron-specific alternative splicing. Proc. Natl Acad. Sci. USA 97, 6350–6355 (2000).
Markovtsov, V. et al. Cooperative assembly of an hnRNP complex induced by a tissue-specific homolog of polypyrimidine tract binding protein. Mol. Cell. Biol. 20, 7463–7479 (2000).
Underwood, J. G., Boutz, P. L., Dougherty, J. D., Stoilov, P. & Black, D. L. Homologues of the Caenorhabditis elegans Fox-1 protein are neuronal splicing regulators in mammals. Mol. Cell. Biol. 25, 10005–10016 (2005).
Jin, Y. et al. A vertebrate RNA-binding protein Fox-1 regulates tissue-specific splicing via the pentanucleotide GCAUG. EMBO J. 22, 905–912 (2003).
Warzecha, C. C., Sato, T. K., Nabet, B., Hogenesch, J. B. & Carstens, R. P. ESRP1 and ESRP2 are epithelial cell-type-specific regulators of FGFR2 splicing. Mol. Cell 33, 591–601 (2009).
Calarco, J. A. et al. Regulation of vertebrate nervous system-specific alternative splicing and development by an SR-related protein. Cell 138, 898–910 (2009).
Lin, S. & Fu, X. D. SR proteins and related factors in alternative splicing. Adv. Exp. Med. Biol. 623, 107–122 (2007).
Martinez-Contreras, R. et al. hnRNP proteins and splicing control. Adv. Exp. Med. Biol. 623, 123–147 (2007).
Fu, X. D. Towards a splicing code. Cell 119, 736–738 (2004).
Zhang, X. H., Arias, M. A., Ke, S. & Chasin, L. A. Splicing of designer exons reveals unexpected complexity in pre-mRNA splicing. RNA 15, 367–376 (2009).
Blanchette, M. et al. Genome-wide analysis of alternative pre-mRNA splicing and RNA-binding specificities of the Drosophila hnRNP A/B family members. Mol. Cell 33, 438–449 (2009).
Valcárcel, J., Singh, R., Zamore, P. D. & Green, M. R. The protein Sex-lethal antagonizes the splicing factor U2AF to regulate alternative splicing of transformer pre-mRNA. Nature 362, 171–175 (1993).
Zuo, P. & Maniatis, T. The splicing factor U2AF35 mediates critical protein–protein interactions in constitutive and enhancer-dependent splicing. Genes Dev. 10, 1356–1368 (1996).
House, A. E. & Lynch, K. W. An exonic splicing silencer represses spliceosome assembly after ATP-dependent exon recognition. Nature Struct. Mol. Biol. 13, 937–944 (2006).
Giles, K. E. & Beemon, K. L. Retroviral splicing suppressor sequesters a 3′ splice site in a 50S aberrant splicing complex. Mol. Cell. Biol. 25, 4397–4405 (2005).
Siebel, C. W., Fresco, L. D. & Rio, D. C. The mechanism of somatic inhibition of Drosophila P-element pre-mRNA splicing: multiprotein complexes at an exon pseudo-5′ splice site control U1 snRNP binding. Genes Dev. 6, 1386–1401 (1992).
Sharma, S., Kohlstaedt, L. A., Damianov, A., Rio, D. C. & Black, D. L. Polypyrimidine tract binding protein controls the transition from exon definition to an intron defined spliceosome. Nature Struct. Mol. Biol. 15, 183–191 (2008).
Martinez-Contreras, R. et al. Intronic binding sites for hnRNP A/B and hnRNP F/H proteins stimulate pre-mRNA splicing. PLoS Biol. 4, e21 (2006).
Smith, D. J., Query, C. C. & Konarska, M. M. “Nought may endure but mutability”: spliceosome dynamics and the regulation of splicing. Mol. Cell 30, 657–666 (2008).
Kornblihtt, A. R. Coupling transcription and alternative splicing. Adv. Exp. Med. Biol. 623, 175–189 (2007).
Park, J. W., Parisky, K., Celotto, A. M., Reenan, R. A. & Graveley, B. R. Identification of alternative splicing regulators by RNA interference in Drosophila. Proc. Natl Acad. Sci. USA 101, 15974–15979 (2004).
Pleiss, J. A., Whitworth, G. B., Bergkessel, M. & Guthrie, C. Transcript specificity in yeast pre-mRNA splicing revealed by mutations in core spliceosomal components. PLoS Biol. 5, e90 (2007).
Fox-Walsh, K. L. et al. The architecture of pre-mRNAs affects mechanisms of splice-site pairing. Proc. Natl Acad. Sci. USA 102, 16176–16181 (2005).
Yu, Y. et al. Dynamic regulation of alternative splicing by silencers that modulate 5′ splice site competition. Cell 135, 1224–1236 (2008).
Dye, M. J., Gromak, N. & Proudfoot, N. J. Exon tethering in transcription by RNA polymerase II. Mol. Cell 21, 849–859 (2006).
Alló, M. et al. Control of alternative splicing through siRNA-mediated transcriptional gene silencing. Nature Struct. Mol. Biol. 16, 717–724 (2009).
Kolasinska-Zwierz, P. et al. Differential chromatin marking of introns and expressed exons by H3K36me3. Nature Genet. 41, 376–381 (2009).
Schwartz, S., Meshorer, E. & Ast, G. Chromatin organization marks exon–intron architecture. Nature Struct. Mol. Biol. 16, 990–995 (2009).
Tilgner, H. et al. Nucleosome positioning as a determinant of exon recognition. Nature Struct. Mol. Biol. 16, 996–1001 (2009).
Neves, G., Zucker, J., Daly, M. & Chess, A. Stochastic yet biased expression of multiple Dscam splice variants by individual cells. Nature Genet. 36, 240–246 (2004).
Graveley, B. R. Mutually exclusive splicing of the insect Dscam pre-mRNA directed by competing intronic RNA secondary structures. Cell 123, 65–73 (2005).
Anastassiou, D., Liu, H. & Varadan, V. Variable window binding for mutually exclusive alternative splicing. Genome Biol. 7, R2 (2006).
Olson, S. et al. A regulator of Dscam mutually exclusive splicing fidelity. Nature Struct. Mol. Biol. 14, 1134–1140 (2007).
Ule, J. et al. An RNA map predicting Nova-dependent splicing regulation. Nature 444, 580–586 (2006). This paper shows that the neural-specific splicing regulator NOVA functions as either an activator or a repressor depending on the context.
Modrek, B. & Lee, C. J. Alternative splicing in the human, mouse and rat genomes is associated with an increased frequency of exon creation and/or loss. Nature Genet. 34, 177–180 (2003).
Sorek, R. & Ast, G. Intronic sequences flanking alternatively spliced exons are conserved between human and mouse. Genome Res. 13, 1631–1637 (2003).
Sugnet, C. W., Kent, W. J., Ares, M. J. & Haussler, D. Transcriptome and genome conservation of alternative splicing events in humans and mice. Pac. Symp. Biocomput. 2004, 66–77 (2004).
Ohler, U., Shomron, N. & Burge, C. B. Recognition of unknown conserved alternatively spliced exons. PLoS Comput. Biol. 1, 113–122 (2005). This paper shows that previously unannotated alternatively spliced exons can be identified accurately by using DNA sequence alone.
Fairbrother, W. G., Yeh, R. F., Sharp, P. A. & Burge, C. B. Predictive identification of exonic splicing enhancers in human genes. Science 297, 1007–1013 (2002). This paper describes one of the first genome-wide predictions of splicing regulatory sequences.
Goren, A. et al. Comparative analysis identifies exonic splicing regulatory sequences the complex definition of enhancers and silencers. Mol. Cell 22, 769–781 (2006).
Shepard, P. J. & Hertel, K. J. Conserved RNA secondary structures promote alternative splicing. RNA 14, 1463–1469 (2008).
Hasselmann, M. et al. Evidence for the evolutionary nascence of a novel sex determination pathway in honeybees. Nature 454, 519–522 (2008).
Beye, M., Hasselmann, M., Fondrk, M. K., Page, R. E. & Omholt, S. W. The gene csd is the primary signal for sexual development in the honeybee and encodes an SR-type protein. Cell 114, 419–429 (2003).
Cartegni, L., Chew, S. L. & Krainer, A. R. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nature Rev. Genet. 3, 285–298 (2002).
Xing, Y. & Lee, C. Alternative splicing and RNA selection pressure — evolutionary consequences for eukaryotic genomes. Nature Rev. Genet. 7, 499–509 (2006).
Zhang, X. H. & Chasin, L. A. Comparison of multiple vertebrate genomes reveals the birth and evolution of human exons. Proc. Natl Acad. Sci. USA 103, 13427–13432 (2006).
Calarco, J. A. et al. Global analysis of alternative splicing differences between humans and chimpanzees. Genes Dev. 21, 2963–2975 (2007).
Graveley, B. R. The haplo-spliceo-transcriptome: common variations in alternative splicing in the human population. Trends Genet. 24, 5–7 (2008).
C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Clamp, M. et al. Distinguishing protein-coding and noncoding genes in the human genome. Proc. Natl Acad. Sci. USA 104, 19428–19433 (2007).
Hillier, L. W. et al. Massively parallel sequencing of the polyadenylated transcriptome of C. elegans. Genome Res. 19, 657–666 (2009).
Stolc, V. et al. A gene expression map for the euchromatic genome of Drosophila melanogaster. Science 306, 655–660 (2004).
Celniker, S. E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009).
Nern, A. et al. An isoform-specific allele of Drosophila N-cadherin disrupts a late step of R7 targeting. Proc. Natl Acad. Sci. USA 102, 12944–12949 (2005).
Wang, Z. & Burge, C. B. Splicing regulation: from a parts list of regulatory elements to an integrated splicing code. RNA 14, 802–813 (2008).
Wojtowicz, W. M. et al. A vast repertoire of Dscam binding specificities arises from modular interactions of variable Ig domains. Cell 130, 1134–1145 (2007).
Demir, E. & Dickson, B. J. fruitless splicing specifies male courtship behavior in Drosophila. Cell 121, 785–794 (2005).
Kwon, S. Y., Xiao, H., Wu, C. & Badenhorst, P. Alternative splicing of NURF301 generates distinct NURF chromatin remodeling complexes with altered modified histone binding specificities. PLoS Genet. 5, e1000574 (2009).
Goodman, S. J., Branda, C. S., Robinson, M. K., Burdine, R. D. & Stern, M. J. Alternative splicing affecting a novel domain in the C. elegans EGL-15 FGF receptor confers functional specificity. Development 130, 3757–3766 (2003).
Muriel, J. M., Dong, C., Hutter, H. & Vogel, B. E. Fibulin-1C and Fibulin-1D splice variants have distinct functions and assemble in a hemicentin-dependent manner. Development 132, 4223–4234 (2005).
Ono, K., Yamashiro, S. & Ono, S. Essential role of ADF/cofilin for assembly of contractile actin networks in the C. elegans somatic gonad. J. Cell Sci. 121, 2662–2670 (2008).
Boucard, A. A., Chubykin, A. A., Comoletti, D., Taylor, P. & Südhof, T. C. A splice code for trans-synaptic cell adhesion mediated by binding of neuroligin 1 to α- and β-neurexins. Neuron 48, 229–236 (2005).
Chen, H. et al. The N-terminal domain of Slack determines the formation and trafficking of Slick/Slack heteromeric sodium-activated potassium channels. J. Neurosci. 29, 5654–5665 (2009).
Sorensen, J. B. et al. Differential control of the releasable vesicle pools by SNAP-25 splice variants and SNAP-23. Cell 114, 75–86 (2003).
Acknowledgements
We apologize to the many authors whose publications are not cited directly because of space limitations. Research in our laboratories is supported by grants from the National Institutes of Health (T.W.N. and B.R.G.), the Raymond and Beverly Sackler Fund for the Arts and Sciences (B.R.G.) and the State of Connecticut's Stem Cell Research Fund (B.R.G.).
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available at http://www.nature.com/reprints. Correspondence should be addressed to T.W.N. (twn@case.edu) or B.R.G. (graveley@neuron.uchc.edu).
Rights and permissions
About this article
Cite this article
Nilsen, T., Graveley, B. Expansion of the eukaryotic proteome by alternative splicing. Nature 463, 457–463 (2010). https://doi.org/10.1038/nature08909
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08909
This article is cited by
-
SF3B4 downregulation restrains lung adenocarcinoma tumorigenesis via 5′ alternative splicing of KAT2A
Scientific Reports (2024)
-
Transcriptional regulation and alternative splicing reveal the molecular strategies of Bombay duck Harpadon nehereus to hypoxia
Fisheries Science (2024)
-
Neural EGFL-like 1, a craniosynostosis-related osteochondrogenic molecule, strikingly associates with neurodevelopmental pathologies
Cell & Bioscience (2023)
-
Splicing complexity as a pivotal feature of alternative exons in mammalian species
BMC Genomics (2023)
-
Mapping intron retention events contributing to complex traits using splice quantitative trait locus
Plant Methods (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.